Abstract
Glucocorticoids display remarkable anti-inflammatory activity, but their use is limited by on-target adverse effects including insulin resistance and skeletal muscle atrophy. We used a chemical systems biology approach, ligand class analysis, to examine ligands designed to modulate glucocorticoid receptor activity through distinct structural mechanisms. These ligands displayed diverse activity profiles, providing the variance required to identify target genes and coregulator interactions that were highly predictive of their effects on myocyte glucose disposal and protein balance. Their anti-inflammatory effects were linked to glucose disposal but not muscle atrophy. This approach also predicted selective modulation in vivo, identifying compounds that were muscle-sparing or anabolic for protein balance and mitochondrial potential. Ligand class analysis defined the mechanistic links between the ligand–receptor interface and ligand-driven physiological outcomes, a general approach that can be applied to any ligand-regulated allosteric signaling system.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw and processed nascent RNA sequencing dataset generated in this study has been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (GEO), with the accession code GSE149453. Other messenger RNA sequencing data cited in this work was downloaded from the GEO with accession code GSE12463654. Sample R scripts used are available at https://github.com/jnwachuk/ML_in_GR_signaling_networks. Compound profiling assay data, results of correlation and machine learning analyses are in Supplemental_Data1.xls.
References
Watson, M. L. et al. A cell-autonomous role for the glucocorticoid receptor in skeletal muscle atrophy induced by systemic glucocorticoid exposure. Am. J. Physiol. Endocrinol. Metab. 302, E1210–E1220 (2012).
Bodine, S. C. & Furlow, J. D. Glucocorticoids and skeletal muscle. Adv. Exp. Med. Biol. 872, 145–176 (2015).
Nettles, K. W. & Greene, G. L. Ligand control of coregulator recruitment to nuclear receptors. Annu. Rev. Physiol. 67, 309–333 (2005).
Kuo, T. et al. Genome-wide analysis of glucocorticoid receptor-binding sites in myotubes identifies gene networks modulating insulin signaling. Proc. Natl Acad. Sci. USA 109, 11160–11165 (2012).
Chen, S. L., Dowhan, D. H., Hosking, B. M. & Muscat, G. E. The steroid receptor coactivator, GRIP-1, is necessary for MEF-2C-dependent gene expression and skeletal muscle differentiation. Genes Dev. 14, 1209–1228 (2000).
Tobimatsu, K. et al. Overexpression of the transcriptional coregulator Cited2 protects against glucocorticoid-induced atrophy of C2C12 myotubes. Biochem. Biophys. Res. Commun. 378, 399–403 (2009).
Amat, R., Solanes, G., Giralt, M. & Villarroya, F. SIRT1 is involved in glucocorticoid-mediated control of uncoupling protein-3 gene transcription. J. Biol. Chem. 282, 34066–34076 (2007).
Nwachukwu, J. C. et al. Systems structural biology analysis of ligand effects on ERα predicts cellular response to environmental estrogens and anti-hormone therapies. Cell Chem. Biol. 24, 35–45 (2017).
Nwachukwu, J. C. et al. Predictive features of ligand-specific signaling through the estrogen receptor. Mol. Syst. Biol. 12, 864 (2016).
Srinivasan, S. et al. Ligand-binding dynamics rewire cellular signaling via estrogen receptor-α. Nat. Chem. Biol. 9, 326–332 (2013).
Jin, Z. et al. Synthesis of novel steroidal agonists, partial agonists, and antagonists for the glucocorticoid receptor. Bioorg. Med. Chem. Lett. 27, 347–353 (2017).
Hu, X. et al. The antagonists but not partial agonists of glucocorticoid receptor ligands show substantial side effect dissociation. Endocrinology 152, 3123–3134 (2011).
Stock, T., Fleishaker, D., Wang, X., Mukherjee, A. & Mebus, C. Improved disease activity with fosdagrocorat (PF-04171327), a partial agonist of the glucocorticoid receptor, in patients with rheumatoid arthritis: a phase 2 randomized study. Int. J. Rheum. Dis. 20, 960–970 (2017).
Nwachukwu, J. C. et al. Resveratrol modulates the inflammatory response via an estrogen receptor-signal integration network. eLife 3, e02057 (2014).
Aarts, J. M. et al. Robust array-based coregulator binding assay predicting ERα-agonist potency and generating binding profiles reflecting ligand structure. Chem. Res. Toxicol. 26, 336–346 (2013).
Geiss, G. K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat. Biotechnol. 26, 317–325 (2008).
Stitt, T. N. et al. The IGF-1/PI3K/Akt pathway prevents expression of muscle atrophy-induced ubiquitin ligases by inhibiting FOXO transcription factors. Mol. Cell 14, 395–403 (2004).
Goodyear, L. J. & Kahn, B. B. Exercise, glucose transport, and insulin sensitivity. Annu. Rev. Med. 49, 235–261 (1998).
Magomedova, L. & Cummins, C. L. Glucocorticoids and metabolic control. Handb. Exp. Pharmacol. 233, 73–93 (2016).
Kuo, T., Harris, C. A. & Wang, J. C. Metabolic functions of glucocorticoid receptor in skeletal muscle. Mol. Cell. Endocrinol. 380, 79–88 (2013).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Kursa, M. B. Robustness of random forest-based gene selection methods. BMC Bioinformatics 15, 8 (2014).
Luciani, D. S. et al. Bcl-2 and Bcl-xL suppress glucose signaling in pancreatic β-cells. Diabetes 62, 170–182 (2013).
Pei, H. et al. FKBP51 affects cancer cell response to chemotherapy by negatively regulating Akt. Cancer Cell 16, 259–266 (2009).
Balsevich, G. et al. Stress-responsive FKBP51 regulates AKT2-AS160 signaling and metabolic function. Nat. Commun. 8, 1725 (2017).
Bowerman, S. & Wereszczynski, J. Detecting allosteric networks using molecular dynamics simulation. Methods Enzymol. 578, 429–447 (2016).
Block, T. et al. Mifepristone plasma level and glucocorticoid receptor antagonism associated with response in patients with psychotic depression. J. Clin. Psychopharmacol. 37, 505–511 (2017).
Arora, V. K. et al. Glucocorticoid receptor confers resistance to antiandrogens by bypassing androgen receptor blockade. Cell 155, 1309–1322 (2013).
Yu, B. et al. Epigenetic landscapes reveal transcription factors that regulate CD8+ T cell differentiation. Nat. Immunol. 18, 573–582 (2017).
Coppo, M., Chinenov, Y., Sacta, M. A. & Rogatsky, I. The transcriptional coregulator GRIP1 controls macrophage polarization and metabolic homeostasis. Nat. Commun. 7, 12254 (2016).
Geltink, R. I. K., Kyle, R. L. & Pearce, E. L. Unraveling the complex interplay between T cell metabolism and function. Annu. Rev. Immunol. 36, 461–488 (2018).
Van den Bossche, J., O’Neill, L. A. & Menon, D. Macrophage immunometabolism: where are we (going)? Trends Immunol. 38, 395–406 (2017).
Hudson, W. H. et al. Cryptic glucocorticoid receptor-binding sites pervade genomic NF-κB response elements. Nat. Commun. 9, 1337 (2018).
Chinenov, Y. et al. Role of transcriptional coregulator GRIP1 in the anti-inflammatory actions of glucocorticoids. Proc. Natl Acad. Sci. USA 109, 11776–11781 (2012).
Stallcup, M. R. & Poulard, C. Gene-specific actions of transcriptional coregulators facilitate physiological plasticity: evidence for a physiological coregulator code. Trends Biochem. Sci. 45, 497–510 (2020).
Stashi, E., York, B. & O’Malley, B. W. Steroid receptor coactivators: servants and masters for control of systems metabolism. Trends Endocrinol. Metab. 25, 337–347 (2014).
Voss, T. C. et al. Dynamic exchange at regulatory elements during chromatin remodeling underlies assisted loading mechanism. Cell 146, 544–554 (2011).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Mei, S. et al. Cistrome Data Browser: a data portal for ChIP–Seq and chromatin accessibility data in human and mouse. Nucleic Acids Res. 45, D658–D662 (2017).
Heikkinen, S., Argmann, C. A., Champy, M.-F. & Auwerx, J. Evaluation of glucose homeostasis. Curr. Protoc. Mol. Biol. Chapter 29, Unit 29B.3 (2007).
Bruno, N. E. et al. Creb coactivators direct anabolic responses and enhance performance of skeletal muscle. EMBO J. 33, 1027–1043 (2014).
Hong, D. H. & Forsberg, N. E. Effects of dexamethasone on protein degradation and protease gene expression in rat L8 myotube cultures. Mol. Cell. Endocrinol. 108, 199–209 (1995).
Walker, D., Htun, H. & Hager, G. L. Using inducible vectors to study intracellular trafficking of GFP-tagged steroid/nuclear receptors in living cells. Methods 19, 386–393 (1999).
Stavreva, D. A. et al. Prevalent glucocorticoid and androgen activity in US water sources. Sci. Rep. 2, 937 (2012).
Eswar, N. et al. Comparative protein structure modeling using Modeller. Curr. Protoc. Bioinformatics Chapter 5, Unit-5.6 (2006).
Pettersen, E. F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Anandakrishnan, R., Aguilar, B. & Onufriev, A. V. H++ 3.0: automating pK prediction and the preparation of biomolecular structures for atomistic molecular modeling and simulations. Nucleic Acids Res. 40, W537–W541 (2012).
Hopkins, C. W., Le Grand, S., Walker, R. C. & Roitberg, A. E. Long-time-step molecular dynamics through hydrogen mass repartitioning. J. Chem. Theory Comput. 11, 1864–1874 (2015).
Roe, D. R. & Cheatham, T. E. 3rd PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Glykos, N. M. Software news and updates. Carma: a molecular dynamics analysis program. J. Comput. Chem. 27, 1765–1768 (2006).
Manjur, A. B. M. K., Lempiäinen, J. K., Malinen, M., Palvimo, J. J. & Niskanen, E. A. IRF2BP2 modulates the crosstalk between glucocorticoid and TNF signaling. J. Steroid Biochem. Mol. Biol. 192, 105382 (2019).
Acknowledgements
N.E.B. was supported by the BallenIsles Men’s Golf Association. K.W.N. was supported by the G.S. Humane Corporation. A W.M. Keck Foundation grant was awarded to E.A.O. D.J.K. was supported by National Institutes of Health (NIH) grant no. DK124870. T.I. is supported by grants from the NIH, the Department of Defense, the National Science Foundation and by start-up funds provided to The Scripps Research Institute from the State of Florida.
Author information
Authors and Affiliations
Contributions
N.E.B. and K.W.N. designed the research. N.E.B., K.W.N., S.V.B., J.C.N., G.L.H., M.D.C. and R.H. designed and developed the methods. N.E.B., S.S., S.V.B., J.C.N., C.C.N., J.N., D.A.S., R.H., D.J.K., X.L., O.E., Z.J. and T.M.K. performed the experiments under the supervision of K.W.N., N.E.B., G.L.H., T.M.K., E.A.O., D.G.P., and T.I. M.D.C. performed the mouse pharmacokinetics study. Z.J. and T.M.K. provided the compounds. K.W.N., N.E.B., J.C.N., X.L., E.A.O., D.J.K. and E.A.O. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
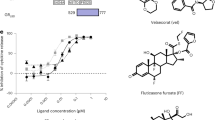
Extended Data Fig. 1 Structure-based design of GR ligands.
a) Chemical structure of the selective GR modulator, PF802. b) Crystal structure of the dexamethasone (Dex) bound GR ligand-binding domain (LBD). Helix 12 is colored red and the NCOA2 coregulator peptide binding in the AF2 binding surface is colored coral. Carbons-3, -11, and -17 are indicated in the chemical structure (pdb entry 1M2Z). c) The bulky dimethylanaline group attached at C11 in RU-486 displaces h12 from the agonist position to disrupt the AF2 surface and generate antagonism (pdb entry 1NHZ). d) Substitutions at C3, as seen with the furoate group in momethasone furoate or the propyne in RU-486 target a small internal pocket to increase affinity (pdb entry 4P6W). e) Substitutions at C3 of the steroidal A-ring enter the solvent channel underneath the AF2 surface, potentially changing the shape of the surface and the ensemble of interacting coregulators (pdb entry 3BQD). f) Model of SR11466 (15) bound to the GR LBD.
Extended Data Fig. 2 Quantitative phenotyping assays for GC action in skeletal muscle.
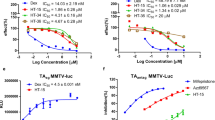
a-e) Myotubes were nutrient-deprived, pre-treated with DMSO, RU-486, or Dex, and treated with insulin as outlined in Methods. a) Effect of insulin on pAKT levels in C2C12 myotubes were compared by In-Cell Western assay (ICW) 48 h after treatment with RU-486 or Dex. b) Quantitation of pAKT in C2C12 myotubes compared by ICW. Bars represent the mean; n = 3, except for DMSO where n = 36 biologically independent samples. c) ICW for surface expression of Glut4 on L6 myotubes. Bars represent the mean; n = 3 biologically independent samples. d) C2C12 myotubes were assayed for protein degradation by release of tritiated phenylalanine. Bars represent the mean; for DMSO, n = 6, and for Dex, n = 4 biologically independent samples. e) ICW for protein synthesis by insulin-induced incorporation of puromycin into C2C12 myotube surface proteins. Bars represent the mean; n = 3 biologically independent samples. f-g) High-content imaging and analysis of C2C12 myoblasts stained with MitoTracker™ dye. Images are representative of 18 images per condition; that is 3 fields x 3 biologically independent samples per condition in each of 2 independent experiments. h) Assay reproducibility from screening 22 compounds on two separate occasions. The Pearson correlation coefficient, r, and its associated p-value is indicated. Each datapoint represents the mean effect of a distinct compound. Also see Methods. i–j) Linear regression demonstrates the predictive power (r2), and associated p-value for the indicated variables. i) Ψm and pAKT predict Synthesis (p = 6 × 10−5 and 8 × 10−5, respectively). j) GR nuclear translocation selectively predicts Glut4 (p = 2 × 10−5) but not pAKT. Each datapoint represents the effects of a distinct ligand.
Extended Data Fig. 3 Relationships among specific genes, peptide interaction assays, and GR-mediated phenotypes.
a-e) Linear regression was performed for the indicated assay pairs, where each point represents a different compound. a) Effects of the ligands on Socs2 expression predicts Glut4 translocation, ψm, and insulin-stimulated protein synthesis. b) Ligand-dependent expression of Bcl2l1, which encodes the mitochondrial anti-apoptotic protein, Bcl-xL does not predict effects of on ψm. c) Fkbp5 expression as a predictor of Glut4 translocation shows an inflection point (arrow). d) The Glut4 data was truncated below the inflection point shown in c). e) GR interaction with an NCOR2 peptide predicts protein synthesis in the C11 subset of ligands. f) The C11- and C3-substituted compounds showed similar variance in the skeletal muscle profiling assays. Blue dashed line, vehicle; red dashed line, Dex. Each datapoint represents the mean effect of a distinct compound, n = 3 biologically independent samples. The error bars represent the mean ± SD of each compound series. For C11, n = 8 distinct compounds; for C3, n = 7 distinct compounds. See also Fig. 2a-e.
Extended Data Fig. 4 Compound structure-activity relationships.
a) Individual compound data for protein degradation, insulin-stimulated protein synthesis, and Glut4 translocation in myotubes, as well as effects on IL-1β-stimulated secretion of IL-6 by A549 cells. Lead compounds are indicated with arrows. Among the 3 compounds with full suppression of IL-6 (13,14,15), only 15 did not inhibit protein synthesis or stimulate protein degradation. 18 showed the greatest anabolic effects, with stimulation of protein synthesis and inhibition of protein degradation. b) Protein degradation in myotubes assayed as described in Extended Data Fig. 2 and Methods. Bars represent the mean, n = 4, except for DMSO where n = 6 biologically independent samples. c) 293 T cells were co-transfected with a GR expression plasmid and MMTV-luciferase reporter. The next day cells were treated with the indicated compounds for 24 h and probed for luciferase activity. For SR16024, was the cells were cotreated with 1 nM Dex. Data are mean ± SEM, n = 3 biologically independent samples. d) Mitochondrial potential of myotubes assayed as described in Extended Data Fig. 2 and Methods. Bars represent the mean; n = 3, except for 15 where n = 2 biologically independent samples.
Extended Data Fig. 5 On-target mechanism of action studies.
a) Reporter activity in steroid-deprived 293 T cells co-transfected with an androgen-responsive ARR3-tk-luc reporter and an androgen receptor (AR) expression plasmid or empty vector control, and then treated with the indicated compounds for 24 h. Dose curves for compounds that stimulated AR activity (left) and the indicated compounds (right) are shown. None of the compounds showed activity with the empty vector control. 18 and 19 are isomers differing only in the position of the chlorine on the benzyl substitution. Datapoints are mean ± SEM; n = 3 biologically independent samples. b) Linear regression demonstrating that ligand-specific AR activity profiles do not correlate with protein synthesis. c) Expression of steroid receptor mRNAs in A549 cells. Only Ar which encodes AR, and Nr3c1 which encodes GR were detected by qPCR. Bars represent the mean; n = 3 independent samples. Also see Supplementary Fig. 3. d) Representative qPCR amplification plots for Pgr, Ar, and Nr3c1 in A549 versus MCF7 cells. Pgr, which encodes the progesterone receptor, is not expressed in A549 cells. e) AR antagonists do not reverse the effects of Dex on IL-6 secretion. IL-6 levels in A549 cell media were measured by AlphaLISA after overnight exposure to the indicated conditions. DHT, 5α-dihydrotestosterone; BIC, bicalutamide; ENZ, enzalutamide. For the controls (left), bars represent the mean; n = 6 biologically independent samples. For dose curves, datapoints are mean ± SEM; n = 3 biologically independent samples. f) Luciferase assay showing the effects of 1 nM DHT, 1 µM BIC and 1 µM ENZ on AR activity, demonstrating that the antagonists have cellular activity. Bars represent the mean; n = 3 biologically independent samples.
Extended Data Fig. 6 In vivo compound profiling.
a) Mouse pharmacokinetics studies of the indicated compounds. Data are mean ± SEM; n = 3 biologically independent samples. b-c) Changes in the lean mass and body weights of male C57BL/6 mice treated with (10 mg/kg Dex or SR16024, or 50 mg/kg SR11466) and fasted overnight. Bars represent the mean; n = 5 mice per group (in each of 2 independent experiments). 1-way ANOVA, Sidak’s multiple comparisons test, adjusted p-values are indicated.
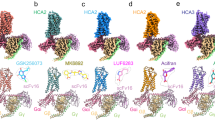
Extended Data Fig. 7 Docking and molecular dynamics simulations.
a) Ribbon diagram of GR LBD bound to the indicated ligands. 7 and 9 were docked with Autodock Vina. b) Differential analysis of correlated motion between Cα atoms from the simulations with the indicated ligands subtracted from Dex. c) Formation of a hydrogen bond with R611 differentially shifts the position of the ligands. d) Formation of the hydrogen bond R611-induced changes in surface structure (red arrows). With Dex, there was a shift in h12 and the C-terminus of h3. With 7, the C-terminus of h11 and N-terminus of h3 were shifted further apart, and away from h12. This destabilization of the h12 interface with h3 and h11 explains why this compound is an antagonist, a mechanism we have called ‘indirect antagonism.’ With 9, there was a rotation of both ends of h3. e) Usage of amino acid residue and edge in the suboptimal pathways between h12 E755 and h5 R614, demonstrating that Dex preferentially utilized R611 instead of W610 as a pathway for correlated motion. Also see Methods.
Supplementary information
Supplementary Information
Supplementary Tables 1–3, Figs. 1–3, Note and references.
Supplementary Data 1
Compound profiling data, correlation results, machine learning results.
Rights and permissions
About this article
Cite this article
Bruno, N.E., Nwachukwu, J.C., Srinivasan, S. et al. Chemical systems biology reveals mechanisms of glucocorticoid receptor signaling. Nat Chem Biol 17, 307–316 (2021). https://doi.org/10.1038/s41589-020-00719-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00719-w
This article is cited by
-
COVID-19 and the pituitary
Pituitary (2021)