Abstract
Many antibiotics inhibit bacterial growth by binding to the ribosome and interfering with protein biosynthesis. Macrolides represent one of the most successful classes of ribosome-targeting antibiotics. The main clinically relevant mechanism of resistance to macrolides is dimethylation of the 23S rRNA nucleotide A2058, located in the drug-binding site, a reaction catalyzed by Erm-type rRNA methyltransferases. Here, we present the crystal structure of the Erm-dimethylated 70S ribosome at 2.4 Å resolution, together with the structures of unmethylated 70S ribosome functional complexes alone or in combination with macrolides. Altogether, our structural data do not support previous models and, instead, suggest a principally new explanation of how A2058 dimethylation confers resistance to macrolides. Moreover, high-resolution structures of two macrolide antibiotics bound to the unmodified ribosome reveal a previously unknown role of the desosamine moiety in drug binding, laying a foundation for the rational knowledge-based design of macrolides that can overcome Erm-mediated resistance.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Coordinates and structure factors were deposited in the RCSB Protein Data Bank with accession codes 6XHV for the A2058-dimethylated Tth 70S ribosome in complex with mRNA, aminoacylated A site Phe-NH-tRNAPhe, aminoacylated P site fMet-NH-tRNAiMet and deacylated E site tRNAPhe; 6XHW for the A2058-unmethylated Tth 70S ribosome in complex with mRNA, aminoacylated A site Phe-NH-tRNAPhe, aminoacylated P site fMet-NH-tRNAiMet and deacylated E site tRNAPhe; 6XHX for the Tth 70S ribosome in complex with ERY and PY; 6XHY for the Tth 70S ribosome in complex with TEL, mRNA, aminoacylated A site Phe-NH-tRNAPhe, aminoacylated P site fMet-NH-tRNAiMet and deacylated E site tRNAPhe.
All previously published structures that were used in this work for structural comparisons were retrieved from the RCSB Protein Data Bank (PDB entries 1VY4, 6ND6, 4V7U, 4V7V, 4V7X, 4V7Y, 4V7Z, 1YI2, 1YHQ, 1YIJ, 4U26).
No sequence data were generated in this study. Analyzed protein sequences are presented with their corresponding accession numbers in the phylogenetic tree (Supplementary Fig. 1) for retrieval from the NCBI protein database. Source data are provided with this paper.
References
Wilson, D. N. Ribosome-targeting antibiotics and mechanisms of bacterial resistance. Nat. Rev. Microbiol. 12, 35–48 (2014).
Fernandes, P., Martens, E., Bertrand, D. & Pereira, D. The solithromycin journey—it is all in the chemistry. Bioorg. Med. Chem. 24, 6420–6428 (2016).
Dinos, G. P. The macrolide antibiotic renaissance. Br. J. Pharmacol. 174, 2967–2983 (2017).
Vazquez-Laslop, N. & Mankin, A. S. How macrolide antibiotics work. Trends Biochem. Sci. 43, 668–684 (2018).
Nunez-Samudio, V. & Chesneau, O. Functional interplay between the ATP binding cassette Msr(D) protein and the membrane facilitator superfamily Mef(E) transporter for macrolide resistance in Escherichia coli. Res. Microbiol. 164, 226–235 (2013).
Fyfe, C., Grossman, T. H., Kerstein, K. & Sutcliffe, J. Resistance to macrolide antibiotics in public health pathogens. Cold Spring Harb. Perspect. Med. 6, a025395 (2016).
Morar, M., Pengelly, K., Koteva, K. & Wright, G. D. Mechanism and diversity of the erythromycin esterase family of enzymes. Biochemistry 51, 1740–1751 (2012).
Shakya, T. & Wright, G. D. Nucleotide selectivity of antibiotic kinases. Antimicrob. Agents Chemother. 54, 1909–1913 (2010).
Poehlsgaard, J. & Douthwaite, S. Macrolide antibiotic interaction and resistance on the bacterial ribosome. Curr. Opin. Investig. Drugs 4, 140–148 (2003).
Sutcliffe, J. & Leclercq, R. Mechanisms of resistance to macrolides, lincosamides, and ketolides. In Macrolide Antibiotics (eds. Schönfeld, W. & Kirst, H. A.) 281–318 (Birkhäuser Verlag, 2002).
Roberts, M. C. et al. Nomenclature for macrolide and macrolide–lincosamide–streptogramin B resistance determinants. Antimicrob. Agents Chemother. 43, 2823–2830 (1999).
Skinner, R., Cundliffe, E. & Schmidt, F. J. Site of action of a ribosomal RNA methylase responsible for resistance to erythromycin and other antibiotics. J. Biol. Chem. 258, 12702–12706 (1983).
Uchiyama, H. & Weisblum, B. N-methyl transferase of Streptomyces erythraeus that confers resistance to the macrolide–lincosamide–streptogramin B antibiotics: amino acid sequence and its homology to cognate R-factor enzymes from pathogenic bacilli and cocci. Gene 38, 103–110 (1985).
Arthur, M., Brisson-Noel, A. & Courvalin, P. Origin and evolution of genes specifying resistance to macrolide, lincosamide and streptogramin antibiotics: data and hypotheses. J. Antimicrob. Chemother. 20, 783–802 (1987).
Lai, C. J. & Weisblum, B. Altered methylation of ribosomal RNA in an erythromycin-resistant strain of Staphylococcus aureus. Proc. Natl Acad. Sci. USA 68, 856–860 (1971).
Goldman, R. C. & Kadam, S. K. Binding of novel macrolide structures to macrolides–lincosamides–streptogramin B-resistant ribosomes inhibits protein synthesis and bacterial growth. Antimicrob. Agents Chemother. 33, 1058–1066 (1989).
Vester, B. & Douthwaite, S. Macrolide resistance conferred by base substitutions in 23S rRNA. Antimicrob. Agents Chemother. 45, 1–12 (2001).
Farrell, D. J., Mendes, R. E. & Jones, R. N. Antimicrobial activity of solithromycin against serotyped macrolide-resistant Streptococcus pneumoniae isolates collected from U.S. medical centers in 2012. Antimicrob. Agents Chemother. 59, 2432–2434 (2015).
Liu, M. & Douthwaite, S. Activity of the ketolide telithromycin is refractory to Erm monomethylation of bacterial rRNA. Antimicrob. Agents Chemother. 46, 1629–1633 (2002).
Liu, M. & Douthwaite, S. Resistance to the macrolide antibiotic tylosin is conferred by single methylations at 23S rRNA nucleotides G748 and A2058 acting in synergy. Proc. Natl Acad. Sci. USA 99, 14658–14663 (2002).
Hansen, J. L. et al. The structures of four macrolide antibiotics bound to the large ribosomal subunit. Mol. Cell 10, 117–128 (2002).
Tu, D., Blaha, G., Moore, P. B. & Steitz, T. A. Structures of MLSBK antibiotics bound to mutated large ribosomal subunits provide a structural explanation for resistance. Cell 121, 257–270 (2005).
Bulkley, D., Innis, C. A., Blaha, G. & Steitz, T. A. Revisiting the structures of several antibiotics bound to the bacterial ribosome. Proc. Natl Acad. Sci. USA 107, 17158–17163 (2010).
Dunkle, J. A., Xiong, L., Mankin, A. S. & Cate, J. H. Structures of the Escherichia coli ribosome with antibiotics bound near the peptidyl transferase center explain spectra of drug action. Proc. Natl Acad. Sci. USA 107, 17152–17157 (2010).
LeTourneau, N., Vimal, P., Klepacki, D., Mankin, A. & Melman, A. Synthesis and antibacterial activity of desosamine-modified macrolide derivatives. Bioorg. Med. Chem. Lett. 22, 4575–4578 (2012).
Small, M. C., Lopes, P., Andrade, R. B. & Mackerell, A. D. Jr. Impact of ribosomal modification on the binding of the antibiotic telithromycin using a combined Grand Canonical Monte Carlo/molecular dynamics simulation approach. PLoS Comput. Biol. 9, e1003113 (2013).
Weisblum, B. Erythromycin resistance by ribosome modification. Antimicrob. Agents Chemother. 39, 577–585 (1995).
Pokkunuri, I. & Champney, W. S. Characteristics of a 50S ribosomal subunit precursor particle as a substrate for ErmE methyltransferase activity and erythromycin binding in Staphylococcus aureus. RNA Biol. 4, 147–153 (2007).
Polikanov, Y. S., Melnikov, S. V., Soll, D. & Steitz, T. A. Structural insights into the role of rRNA modifications in protein synthesis and ribosome assembly. Nat. Struct. Mol. Biol. 22, 342–344 (2015).
Almutairi, M. M. et al. Resistance to ketolide antibiotics by coordinated expression of rRNA methyltransferases in a bacterial producer of natural ketolides. Proc. Natl Acad. Sci. USA 112, 12956–12961 (2015).
Carr, J. F., Danziger, M. E., Huang, A. L., Dahlberg, A. E. & Gregory, S. T. Engineering the genome of Thermus thermophilus using a counterselectable marker. J. Bacteriol. 197, 1135–1144 (2015).
Bailey, M., Chettiath, T. & Mankin, A. S. Induction of erm(C) expression by noninducing antibiotics. Antimicrob. Agents Chemother. 52, 866–874 (2008).
Polikanov, Y. S., Steitz, T. A. & Innis, C. A. A proton wire to couple aminoacyl-tRNA accommodation and peptide-bond formation on the ribosome. Nat. Struct. Mol. Biol. 21, 787–793 (2014).
Almutairi, M. M. et al. Co-produced natural ketolides methymycin and pikromycin inhibit bacterial growth by preventing synthesis of a limited number of proteins. Nucleic Acids Res. 45, 9573–9582 (2017).
Svetlov, M. S. et al. High-resolution crystal structures of ribosome-bound chloramphenicol and erythromycin provide the ultimate basis for their competition. RNA 25, 600–606 (2019).
Khabibullina, N. F. et al. Structure of dirithromycin bound to the bacterial ribosome suggests new ways for rational improvement of macrolides. Antimicrob. Agents Chemother. 63, e02266-18 (2019).
Zhang, Y., El Kouni, M. H. & Ealick, S. E. Substrate analogs induce an intermediate conformational change in Toxoplasma gondii adenosine kinase. Acta Crystallogr. D Biol. Crystallogr. 63, 126–134 (2007).
Pestka, S. & Lemahieu, R. A. Effect of erythromycin analogues on binding of [14C]erythromycin to Escherichia coli ribosomes. Antimicrob. Agents Chemother. 6, 479–488 (1974).
Gupta, P., Sothiselvam, S., Vazquez-Laslop, N. & Mankin, A. S. Deregulation of translation due to post-transcriptional modification of rRNA explains why erm genes are inducible. Nat. Commun. 4, 1984 (2013).
Douthwaite, S. & Aagaard, C. Erythromycin binding is reduced in ribosomes with conformational alterations in the 23S rRNA peptidyl transferase loop. J. Mol. Biol. 232, 725–731 (1993).
Pfister, P. et al. The structural basis of macrolide–ribosome binding assessed using mutagenesis of 23S rRNA positions 2058 and 2059. J. Mol. Biol. 342, 1569–1581 (2004).
Calcutt, M. J. & Cundliffe, E. Cloning of a lincosamide resistance determinant from Streptomyces caelestis, the producer of celesticetin, and characterization of the resistance mechanism. J. Bacteriol. 172, 4710–4714 (1990).
Roost, C. et al. Structure and thermodynamics of N6-methyladenosine in RNA: a spring-loaded base modification. J. Am. Chem. Soc. 137, 2107–2115 (2015).
Contreras, A. & Vazquez, D. Cooperative and antagonistic interactions of peptidyl-tRNA and antibiotics with bacterial ribosomes. Eur. J. Biochem. 74, 539–547 (1977).
Pestka, S. Binding of [14C]erythromycin to Escherichia coli ribosomes. Antimicrob. Agents Chemother. 6, 474–478 (1974).
Noeske, J. et al. Synergy of streptogramin antibiotics occurs independently of their effects on translation. Antimicrob. Agents Chemother. 58, 5269–5279 (2014).
Seiple, I. B. et al. A platform for the discovery of new macrolide antibiotics. Nature 533, 338–345 (2016).
Li, Q. et al. Synthetic group A streptogramin antibiotics that overcome Vat resistance. Nature 586, 145–150 (2020).
Hansen, L. H., Mauvais, P. & Douthwaite, S. The macrolide–ketolide antibiotic binding site is formed by structures in domains II and V of 23S ribosomal RNA. Mol. Microbiol. 31, 623–631 (1999).
Scheinfeld, N. Telithromycin: a brief review of a new ketolide antibiotic. J. Drugs Dermatol. 3, 409–413 (2004).
McArthur, A. G. et al. The Comprehensive Antibiotic Resistance Database. Antimicrob. Agents Chemother. 57, 3348–3357 (2013).
Katoh, K., Kuma, K., Toh, H. & Miyata, T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–518 (2005).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Miller, M. A. et al. A RESTful API for access to phylogenetic tools via the CIPRES Science Gateway. Evol. Bioinform. Online 11, 43–48 (2015).
Koyama, Y., Hoshino, T., Tomizuka, N. & Furukawa, K. Genetic transformation of the extreme thermophile Thermus thermophilus and of other Thermus spp. J. Bacteriol. 166, 338–340 (1986).
Gregory, S. T., Carr, J. F., Rodriguez-Correa, D. & Dahlberg, A. E. Mutational analysis of 16S and 23S rRNA genes of Thermus thermophilus. J. Bacteriol. 187, 4804–4812 (2005).
Gamper, H. & Hou, Y. M. tRNA 3′-amino-tailing for stable amino acid attachment. RNA 24, 1878–1885 (2018).
Schuttelkopf, A. W. & van Aalten, D. M. PRODRG: a tool for high-throughput crystallography of protein–ligand complexes. Acta Crystallogr. D Biol. Crystallogr. 60, 1355–1363 (2004).
Acknowledgements
We thank N. Vazquez-Laslop for valuable discussions and critical feedback and all members of the A.S.M. and Y.S.P. laboratories for valuable suggestions. We thank the staff at NE-CAT beamlines 24-ID-C and 24-ID-E for help with data collection and freezing of the crystals, especially M. Capel, F. Murphy, I. Kourinov, A. Lynch, S. Banerjee, D. Neau, J. Schuermann, N. Sukumar, J. Withrow, K. Perry and C. Salbego. This work is based on research conducted at the Northeastern Collaborative Access Team beamlines, which are funded by the National Institute of General Medical Sciences from the National Institutes of Health (P30-GM124165 to NE-CAT). The Eiger 16M detector on the 24-ID-E beamline is funded by an NIH-ORIP HEI grant (S10-OD021527 to NE-CAT). This research used resources from the Advanced Photon Source, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under contract no. DE-AC02-06CH11357. This work was supported by Illinois State startup funds (to Y.S.P.), the National Institutes of Health (R21-AI137584 to A.S.M. and Y.S.P.; R01-GM132302 to Y.S.P.; R35-GM127134 to A.S.M.; R01-GM094157 to S.T.G.), the USDA National Institute for Food and Agriculture (Hatch project 1016013 to S.T.G.) and the Swedish Research Council (Vetenskapsrådet) (2015-04746 and 2019-01085 to G.C.A.).
Author information
Authors and Affiliations
Contributions
M.S.S., with help from S.T.G., constructed the Tth HB27 strain expressing Erm-like methyltransferase; M.S.S. performed the assessment of A2058 methylation; M.S.S. and E.A.S. grew Tth cells and purified A2058-methylated and unmethylated 70S ribosomes; E.A.S. and E.V.A. prepared hydrolysis-resistant aminoacylated tRNA molecules; M.S.S., E.A.S., E.V.A. and Y.S.P. designed and performed X-ray crystallography experiments; A.S.M. and Y.S.P. supervised the experiments. G.C.A. performed phylogenetic analysis and predicted thermostable erm genes. All authors interpreted the results. M.S.S., S.T.G., A.S.M. and Y.S.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks Stephen Douthwaite and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Electron density maps of N6-dimethylated and unmethylated adenine residues in the current structure of the A2058-methylated Thermus thermophilus 70S ribosome.
a–c, Unbiased Fo-Fc (top row; grey and green mesh) and 2Fo-Fc (bottom row; blue mesh) electron density maps corresponding to the (a) N6-dimethylated A2058 residue, or (b) the unmethylated A2057 residue of the 23S rRNA, or (c) N6-dimethylated A1519 residue of the 16S rRNA (naturally modified by methyltransferase KsgA) in the T. thermophilus 70S ribosome contoured at 3.0σ and 1.0σ, respectively. The refined models are displayed in the corresponding electron density maps. Carbon atoms are colored blue for the methylated A2058, light blue for the unmethylated A2058, and pale yellow for the methylated A1519. Nitrogens are colored dark blue; oxygens are red, phosphorus atoms are green. Grey mesh shows the Fo-Fc map after refinement with the entire modified nucleotide omitted. Green mesh shows the Fo-Fc electron density map after refinement with the nucleotides A2058 and A1519 built as regular unmethylated adenines. We show electron density maps for the A2057 and the A1519 nucleotides as negative and positive control references originating from the same structure.
Extended Data Fig. 2 Aminoacylated A- and P-site tRNAs bound to the A2058-methylated 70S ribosome.
(a, b, c) High-resolution (2.4Å) 2Fo-Fc electron density map (blue mesh) for the ribosome-bound A-site Phe-NH-tRNAPhe (green) and the P-site fMet-NH-tRNAiMet (dark blue). The refined models of tRNAs are displayed in their respective electron density map contoured at 1.2σ. a, The entire bodies of the A- and P-site tRNAs viewed from the back of the 50S subunit, as indicated by the inset. Ribosome subunits are omitted for clarity. b, c, Close-up views of the tRNA CCA-ends carrying phenylalanyl (magenta) and formyl-methionyl (blue) moieties. Nitrogens are colored blue; oxygens are red, sulfur is yellow. In (B), H-bond between the α-amino group and the 2’-OH of the A76 of the P-site tRNA is shown with the black dotted line. This H-bond is pivotal to optimally orient α-amine for an in-line nucleophilic attack onto the carbonyl carbon of the P-site substrate. d, Superimposed models of aminoacylated tRNAs from the current 2.4Å structure of A2058-methylate 70S ribosome with the previous 2.6Å structure of A2058-unmethylated ribosome (PDB entry 1VY433).
Extended Data Fig. 3 Structural comparison of the ribosome-bound macrolides.
a, Superpositioning of the structures of ERY in complex with the 70S ribosomes from T. thermophilus (teal, PDB entry 6ND635; green, PDB entry 4V7X23), E. coli (yellow, PDB entry 4V7U24), or H. marismortui (blue, PDB entry 1YI222). b, Superpositioning of the structures of the 50S ribosomal subunit from H. marismortui in complex with ERY (blue, PDB entry 1YI222), azithromycin (AZI, orange, PDB entry 1YHQ22), and telithromycin (TEL, magenta, PDB entry 1YIJ22). c, Superpositioning of the structures of the 70S ribosome from T. thermophilus in complex with ERY (green, PDB entry 4V7X23), azithromycin (AZI, orange, PDB entry 4V7Y23), and telithromycin (TEL, magenta, PDB entry 4V7Z23). The insets show closed up views of the desosamine sugar. Note a significant variability in the orientation of the dimethylamino groups of the desoamine sugar among ribosome-bound macrolides.
Extended Data Fig. 4 Analysis of the orientation of the dimethyl-amino group in previously published structures of ribosome-bound macrolides.
a, b, In the structures of erythromycin (ERY) in complex with H. marismortui 50S subunit (blue, PDB entry 1YI222), or T. thermophilus 70S ribosome (green, PDB entry 4V7X23) the nitrogen lone pair of electrons on the dimethyl-amino group of the desosamine sugar is oriented towards the exocyclic N6-amino group of the A2059 nucleotide of the 23S rRNA (blue), as indicated by the yellow arrow. Note that the formation of a direct H-bond, in this case, is impossible due to the large distance between the potential interacting partners (grey dashed lines). H-bond between 2’-OH group of desosamine sugar and N1 atom of A2058 is shown by the black dashed lines. c, d, In the structures of azithromycin (AZI) in complex with H. marismortui 50S subunit (orange, PDB entry 1YHQ22), or T. thermophilus 70S ribosome (orange, PDB entry 4V7Y23) the nitrogen lone pair of electrons of the dimethyl-amino group on the desosamine is rotated by approximately 90 degrees away from the residue A2059 in the direction of residue A2058, as indicated by the yellow arrow. Note that, in this case, there are no obvious binding partners for this group on the ribosome.
Extended Data Fig. 5 Electron density maps of ribosome-bound erythromycin and telithromycin.
a, c, Chemical structures of NPET-binding macrolide antibiotics erythromycin (a, ERY) and telithromycin (c, TEL). b, d, High-resolution 2Fo-Fc electron density maps (blue mesh) of ERY (b, green) and TEL (d, magenta) in complex with the T. thermophilus 70S ribosome viewed from two different perspectives. The refined models of ERY or TEL are displayed in their respective electron densities contoured at 1.0σ. Nitrogen atoms are colored blue; oxygens are red. Key chemical moieties of each drug are labeled.
Extended Data Fig. 6 In silico modeling of the A2058 mutants.
a, Water-mediated coordination of a macrolide antibiotic (ERY, green) by the wild-type A2058 nucleotide (blue) of the 23S rRNA. Existing H-bonds are shown by the black dashed lines. b, c, d, Strong water-mediated coordination of a macrolide molecule on the ribosome becomes impossible when the adenine nucleotide at position 2058 of the 23S rRNA is mutated to guanine (b), cytosine (c), or uracil (d) nucleotides. Non-existing H-bonds (due to distance restraints) are shown by the grey dashed lines. Note that in the case of A2058G mutation, in addition to the inability of the guanine nucleotide to coordinate a macrolide via a water molecule, the exocyclic N2-amino group of guanine clashes with the C4-methyl group attached to the macrolactone ring of a macrolide molecule.
Extended Data Fig. 7 Alignment of the structures of A2058-methylated 70S ribosome with the ribosome-bound clindamycin or quinupristin.
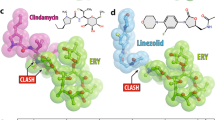
Superposition of the current structure of A2058-methylated 70S ribosome with the structure of ribosome-bound lincosamide antibiotic clindamycin (yellow, PDB entry 4V7V24) (a, b) or streptogramin type B antibiotic quinupristin (teal, PDB entry 4U2646) (c, d). Nucleotides of the Erm-modified and unmodified ribosomes are shown in blue and light blue, respectively. Methylated A2058 residue is shown in blue with the methyl groups highlighted in orange. All structures were aligned based on domain V of the 23S rRNA.
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and Figs. 1 and 2
Source data
Source Data Fig. 1
Unprocessed, uncropped denaturing gel for Fig. 1d,e.
Rights and permissions
About this article
Cite this article
Svetlov, M.S., Syroegin, E.A., Aleksandrova, E.V. et al. Structure of Erm-modified 70S ribosome reveals the mechanism of macrolide resistance. Nat Chem Biol 17, 412–420 (2021). https://doi.org/10.1038/s41589-020-00715-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00715-0
This article is cited by
-
Structural basis of Cfr-mediated antimicrobial resistance and mechanisms to evade it
Nature Chemical Biology (2024)
-
Structural conservation of antibiotic interaction with ribosomes
Nature Structural & Molecular Biology (2023)
-
Structural insights into the mechanism of overcoming Erm-mediated resistance by macrolides acting together with hygromycin-A
Nature Communications (2023)
-
Insights into the ribosome function from the structures of non-arrested ribosome–nascent chain complexes
Nature Chemistry (2023)
-
Loss of allosteric regulation in α-isopropylmalate synthase identified as an antimicrobial resistance mechanism
npj Antimicrobials and Resistance (2023)