Abstract
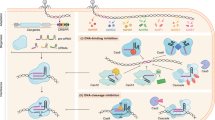
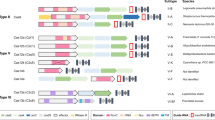
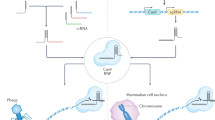
Many bacterial and archaeal organisms use clustered regularly interspaced short palindromic repeats–CRISPR associated (CRISPR–Cas) systems to defend themselves from mobile genetic elements. These CRISPR–Cas systems are classified into six types based on their composition and mechanism. CRISPR–Cas enzymes are widely used for genome editing and offer immense therapeutic opportunity to treat genetic diseases. To realize their full potential, it is important to control the timing, duration, efficiency and specificity of CRISPR–Cas enzyme activities. In this Review we discuss the mechanisms of natural CRISPR–Cas regulatory biomolecules and engineering strategies that enhance or inhibit CRISPR–Cas immunity by altering enzyme function. We also discuss the potential applications of these CRISPR regulators and highlight unanswered questions about their evolution and purpose in nature.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Jansen, R., Embden, J. D., Gaastra, W. & Schouls, L. M. Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 43, 1565–1575 (2002). This article reports the use of the acronym “CRISPR”.
Mojica, F. J. M., Díez-Villaseñor, C., García-Martínez, J. & Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 60, 174–182 (2005). This report shows that spacers within CRISPR arrays serve as a memory of past infections.
Makarova, K. S. et al. Evolutionary classification of CRISPR-Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol. 18, 67–83 (2020).
Vale, P. F. et al. Costs of CRISPR-Cas-mediated resistance in Streptococcus thermophilus. Proc. Biol. Sci. 282, 20151270 (2015).
Westra, E. R. et al. H-NS-mediated repression of CRISPR-based immunity in Escherichia coli K12 can be relieved by the transcription activator LeuO. Mol. Microbiol. 77, 1380–1393 (2010).
Medina-Aparicio, L. et al. The CRISPR/Cas immune system is an operon regulated by LeuO, H-NS, and leucine-responsive regulatory protein in Salmonella enterica serovar Typhi. J. Bacteriol. 193, 2396–2407 (2011).
Liu, T. et al. Coupling transcriptional activation of CRISPR-Cas system and DNA repair genes by Csa3a in Sulfolobus islandicus. Nucleic Acids Res. 45, 8978–8992 (2017).
He, F., Vestergaard, G., Peng, W., She, Q. & Peng, X. CRISPR-Cas type I-A Cascade complex couples viral infection surveillance to host transcriptional regulation in the dependence of Csa3b. Nucleic Acids Res. 45, 1902–1913 (2017).
Patterson, A. G., Chang, J. T., Taylor, C. & Fineran, P. C. Regulation of the Type I-F CRISPR-Cas system by CRP-cAMP and GalM controls spacer acquisition and interference. Nucleic Acids Res. 43, 6038–6048 (2015).
Perez-Rodriguez, R. et al. Envelope stress is a trigger of CRISPR RNA-mediated DNA silencing in Escherichia coli. Mol. Microbiol. 79, 584–599 (2011).
Patterson, A. G. et al. Quorum sensing controls adaptive immunity through the regulation of multiple CRISPR-Cas systems. Mol. Cell 64, 1102–1108 (2016).
Høyland-Kroghsbo, N. M. et al. Quorum sensing controls the Pseudomonas aeruginosa CRISPR-Cas adaptive immune system. Proc. Natl. Acad. Sci. USA 114, 131–135 (2017).
Borges, A. L. et al. Bacterial alginate regulators and phage homologs repress CRISPR-Cas immunity. Nat. Microbiol. 5, 679–687 (2020).
Høyland-Kroghsbo, N. M., Muñoz, K. A. & Bassler, B. L. Temperature, by controlling growth rate, regulates CRISPR-Cas activity in Pseudomonas aeruginosa. MBio 9, e02184–18 (2018).
Ahator, S. D., Jianhe, W. & Zhang, L.-H. The ECF sigma factor PvdS regulates the type I-F CRISPR-Cas system in Pseudomonas aeruginosa. Preprint at bioRxiv https://doi.org/10.1101/2020.01.31.929752 (2020).
Koonin, E. V. & Makarova, K. S. Discovery of oligonucleotide signaling mediated by CRISPR-associated polymerases solves two puzzles but leaves an enigma. ACS Chem. Biol. 13, 309–312 (2018).
Lin, P. et al. High-throughput screen reveals sRNAs regulating crRNA biogenesis by targeting CRISPR leader to repress Rho termination. Nat. Commun. 10, 3728 (2019).
Workman, R. E. et al. A natural single-guide RNA repurposes Cas9 to autoregulate CRISPR-Cas expression. Preprint at bioRxiv https://doi.org/10.1101/2020.05.21.102756 (2020).
Bondy-Denomy, J. et al. A unified resource for tracking anti-CRISPR names. CRISPR J. 1, 304–305 (2018).
Chowdhury, S. et al. Structure reveals mechanisms of viral suppressors that intercept a CRISPR RNA-guided surveillance complex. Cell 169, 47–57.e11 (2017).
Rollins, M. F. et al. Structure reveals a mechanism of CRISPR-RNA-guided nuclease recruitment and anti-CRISPR viral mimicry. Mol. Cell 74, 132–142.e5 (2019).
Wang, X. et al. Structural basis of Cas3 inhibition by the bacteriophage protein AcrF3. Nat. Struct. Mol. Biol. 23, 868–870 (2016).
Harrington, L. B. et al. A broad-spectrum inhibitor of CRISPR-Cas9. Cell 170, 1224–1233.e15 (2017).
Thavalingam, A. et al. Inhibition of CRISPR-Cas9 ribonucleoprotein complex assembly by anti-CRISPR AcrIIC2. Nat. Commun. 10, 2806 (2019).
Liu, L., Yin, M., Wang, M. & Wang, Y. Phage AcrIIA2 DNA mimicry: structural basis of the CRISPR and anti-CRISPR arms race. Mol. Cell 73, 611–620.e3 (2019).
Shin, J. et al. Disabling Cas9 by an anti-CRISPR DNA mimic. Sci. Adv. 3, e1701620 (2017).
Meeske, A. J. et al. A phage-encoded anti-CRISPR enables complete evasion of type VI-A CRISPR-Cas immunity. Science 369, 54–59 (2020).
Fuchsbauer, O. et al. Cas9 allosteric inhibition by the anti-CRISPR protein AcrIIA6. Mol. Cell 76, 922–937.e7 (2019).
Knott, G. J. et al. Structural basis for AcrVA4 inhibition of specific CRISPR-Cas12a. eLife 8, e49110 (2019).
Zhu, Y. et al. Diverse mechanisms of CRISPR-Cas9 inhibition by type IIC anti-CRISPR proteins. Mol. Cell 74, 296–309.e7 (2019).
Knott, G. J. et al. Broad-spectrum enzymatic inhibition of CRISPR-Cas12a. Nat. Struct. Mol. Biol. 26, 315–321 (2019). This is one of the first reports showing that Acrs can possess enzymatic activity.
Dong, L. et al. An anti-CRISPR protein disables type V Cas12a by acetylation. Nat. Struct. Mol. Biol. 26, 308–314 (2019). This is one of the first reports showing that Acrs can possess enzymatic activity.
Athukoralage, J. S. et al. An anti-CRISPR viral ring nuclease subverts type III CRISPR immunity. Nature 577, 572–575 (2020). This paper reports the discovery of an Acr against type III CRISPR systems.
Garcia, B. et al. Anti-CRISPR AcrIIA5 potently inhibits all Cas9 homologs used for genome editing. Cell Rep. 29, 1739–1746.e5 (2019).
Mahendra, C. et al. Broad-spectrum anti-CRISPR proteins facilitate horizontal gene transfer. Nat. Microbiol. 5, 620–629 (2020).
Wei, Y., Terns, R. M. & Terns, M. P. Cas9 function and host genome sampling in Type II-A CRISPR-Cas adaptation. Genes Dev. 29, 356–361 (2015).
Malone, L. M. et al. A jumbo phage that forms a nucleus-like structure evades CRISPR-Cas DNA targeting but is vulnerable to type III RNA-based immunity. Nat. Microbiol. 5, 48–55 (2020).
Mendoza, S. D. et al. A bacteriophage nucleus-like compartment shields DNA from CRISPR nucleases. Nature 577, 244–248 (2020).
Chaikeeratisak, V. et al. Assembly of a nucleus-like structure during viral replication in bacteria. Science 355, 194–197 (2017). This report shows that bacteriophages can form nucleus-like structures during infection in bacteria.
Al-Shayeb, B. et al. Clades of huge phages from across Earth’s ecosystems. Nature 578, 425–431 (2020).
Shmakov, S. A., Makarova, K. S., Wolf, Y. I., Severinov, K. V. & Koonin, E. V. Systematic prediction of genes functionally linked to CRISPR-Cas systems by gene neighborhood analysis. Proc. Natl. Acad. Sci. USA 115, E5307–E5316 (2018).
Samai, P. et al. Co-transcriptional DNA and RNA cleavage during type III CRISPR-Cas immunity. Cell 161, 1164–1174 (2015).
Liu, T. Y., Liu, J.-J., Aditham, A. J., Nogales, E. & Doudna, J. A. Target preference of Type III-A CRISPR-Cas complexes at the transcription bubble. Nat. Commun. 10, 3001 (2019).
Elmore, J. R. et al. Bipartite recognition of target RNAs activates DNA cleavage by the Type III-B CRISPR-Cas system. Genes Dev. 30, 447–459 (2016).
Kazlauskiene, M., Tamulaitis, G., Kostiuk, G., Venclovas, Č. & Siksnys, V. Spatiotemporal control of type III-A CRISPR-Cas immunity: coupling DNA degradation with the target RNA recognition. Mol. Cell 62, 295–306 (2016).
Han, W. et al. A Type III-B Cmr effector complex catalyzes the synthesis of cyclic oligoadenylate second messengers by cooperative substrate binding. Nucleic Acids Res. 46, 10319–10330 (2018).
Mogila, I. et al. Genetic dissection of the type III-A CRISPR-Cas system Csm complex reveals roles of individual subunits. Cell Rep. 26, 2753–2765.e4 (2019).
McMahon, S. A. et al. Structure and mechanism of a Type III CRISPR defence DNA nuclease activated by cyclic oligoadenylate. Nat. Commun. 11, 500 (2020).
Kazlauskiene, M., Kostiuk, G., Venclovas, Č., Tamulaitis, G. & Siksnys, V. A cyclic oligonucleotide signaling pathway in type III CRISPR-Cas systems. Science 357, 605–609 (2017).
Niewoehner, O. et al. Type III CRISPR-Cas systems produce cyclic oligoadenylate second messengers. Nature 548, 543–548 (2017).
Niewoehner, O. & Jinek, M. Structural basis for the endoribonuclease activity of the type III-A CRISPR-associated protein Csm6. RNA 22, 318–329 (2016).
Garcia-Doval, C. et al. Activation and self-inactivation mechanisms of the cyclic oligoadenylate-dependent CRISPR ribonuclease Csm6. Nat. Commun. 11, 1596 (2020).
Molina, R. et al. Structure of Csx1-cOA4 complex reveals the basis of RNA decay in Type III-B CRISPR-Cas. Nat. Commun. 10, 4302 (2019).
Grüschow, S., Athukoralage, J. S., Graham, S., Hoogeboom, T. & White, M. F. Cyclic oligoadenylate signalling mediates Mycobacterium tuberculosis CRISPR defence. Nucleic Acids Res. 47, 9259–9270 (2019).
Lau, R. K. et al. Structure and mechanism of a cyclic trinucleotide-activated bacterial endonuclease mediating bacteriophage immunity. Mol. Cell 77, 723–733.e6 (2020).
Chou-Zheng, L. & Hatoum-Aslan, A. A type III-A CRISPR-Cas system employs degradosome nucleases to ensure robust immunity. eLife 8, e45393 (2019).
Smargon, A. A. et al. Cas13b is a type VI-B CRISPR-associated RNA-guided RNase differentially regulated by accessory proteins Csx27 and Csx28. Mol. Cell 65, 618–630.e7 (2017).
Yan, W. X. et al. Cas13d is a compact RNA-targeting type VI CRISPR effector positively modulated by a WYL-domain-containing accessory protein. Mol. Cell 70, 327–339.e5 (2018).
Zhang, H., Dong, C., Li, L., Wasney, G. A. & Min, J. Structural insights into the modulatory role of the accessory protein WYL1 in the Type VI-D CRISPR-Cas system. Nucleic Acids Res. 47, 5420–5428 (2019).
Stanley, S. Y. et al. Anti-CRISPR-associated proteins are crucial repressors of anti-CRISPR transcription. Cell 178, 1452–1464.e13 (2019).
Birkholz, N., Fagerlund, R. D., Smith, L. M., Jackson, S. A. & Fineran, P. C. The autoregulator Aca2 mediates anti-CRISPR repression. Nucleic Acids Res. 47, 9658–9665 (2019).
Watters, K. E. et al. Potent CRISPR-Cas9 inhibitors from Staphylococcus genomes. Proc. Natl. Acad. Sci. USA 117, 6531–6539 (2020).
Osuna, B. A. et al. Critical anti-CRISPR locus repression by a bi-functional Cas9 inhibitor. Cell Host Microbe 28, 23–30.e5 (2020).
O’Connell, M. R. Molecular mechanisms of RNA targeting by Cas13-containing type VI CRISPR-Cas systems. J. Mol. Biol. 431, 66–87 (2019).
Meeske, A. J., Nakandakari-Higa, S. & Marraffini, L. A. Cas13-induced cellular dormancy prevents the rise of CRISPR-resistant bacteriophage. Nature 570, 241–245 (2019).
Rostøl, J. T. & Marraffini, L. A. Non-specific degradation of transcripts promotes plasmid clearance during type III-A CRISPR-Cas immunity. Nat. Microbiol. 4, 656–662 (2019).
Jia, N., Jones, R., Yang, G., Ouerfelli, O. & Patel, D. J. CRISPR-Cas III-A Csm6 CARF domain is a ring nuclease triggering stepwise cA4 cleavage with ApA>p formation terminating RNase activity. Mol. Cell 75, 944–956.e6 (2019).
Athukoralage, J. S. et al. The dynamic interplay of host and viral enzymes in type III CRISPR-mediated cyclic nucleotide signalling. eLife 9, e55852 (2020).
Athukoralage, J. S., Rouillon, C., Graham, S., Grüschow, S. & White, M. F. Ring nucleases deactivate type III CRISPR ribonucleases by degrading cyclic oligoadenylate. Nature 562, 277–280 (2018).
Athukoralage, J. S. et al. Tetramerisation of the CRISPR ring nuclease Crn3/Csx3 facilitates cyclic oligoadenylate cleavage. eLife 9, e57627 (2020).
Samolygo, A., Athukoralage, J. S., Graham, S. & White, M. F. Fuse to defuse: a self-limiting ribonuclease-ring nuclease fusion for type III CRISPR defence. Nucleic Acids Res. 48, 6149–6156 (2020).
Lin, P. et al. CRISPR-Cas13 inhibitors block RNA editing in bacteria and mammalian cells. Mol. Cell 78, 850–861.e5 (2020).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Baeumler, T. A., Ahmed, A. A. & Fulga, T. A. Engineering synthetic signaling pathways with programmable dCas9-based chimeric receptors. Cell Rep. 20, 2639–2653 (2017).
Perli, S. D., Cui, C. H. & Lu, T. K. Continuous genetic recording with self-targeting CRISPR-Cas in human cells. Science 353, aag0511 (2016). Pioneering work demonstrating the use of Cas9 as a genetic recorder of molecular events in mammalian cells and mice is detailed.
Kempton, H. R., Goudy, L. E., Love, K. S. & Qi, L. S. Multiple input sensing and signal integration using a split Cas12a system. Mol. Cell 78, 184–191.e3 (2020).
Marino, N. D., Pinilla-Redondo, R., Csörgő, B. & Bondy-Denomy, J. Anti-CRISPR protein applications: natural brakes for CRISPR-Cas technologies. Nat. Methods 17, 471–479 (2020).
Hoffmann, M. D. et al. Cell-specific CRISPR-Cas9 activation by microRNA-dependent expression of anti-CRISPR proteins. Nucleic Acids Res. 47, e75 (2019).
Hirosawa, M., Fujita, Y. & Saito, H. Cell-type-specific CRISPR activation with microRNA-responsive AcrllA4 switch. ACS Synth. Biol. 8, 1575–1582 (2019).
Lee, J. et al. Tissue-restricted genome editing in vivo specified by microRNA-repressible anti-CRISPR proteins. RNA 25, 1421–1431 (2019).
Basgall, E. M. et al. Gene drive inhibition by the anti-CRISPR proteins AcrIIA2 and AcrIIA4 in Saccharomyces cerevisiae. Microbiology 164, 464–474 (2018).
Maji, B. et al. A high-throughput platform to identify small-molecule inhibitors of CRISPR-Cas9. Cell 177, 1067–1079.e19 (2019).
Polstein, L. R. & Gersbach, C. A. A light-inducible CRISPR-Cas9 system for control of endogenous gene activation. Nat. Chem. Biol. 11, 198–200 (2015).
Oakes, B. L. et al. Profiling of engineering hotspots identifies an allosteric CRISPR-Cas9 switch. Nat. Biotechnol. 34, 646–651 (2016).
Oakes, B. L. et al. CRISPR-Cas9 circular permutants as programmable scaffolds for genome modification. Cell 176, 254–267.e16 (2019).
Manna, D. et al. A singular system with precise dosing and spatiotemporal control of CRISPR-Cas9. Angew. Chem. Int. Ed. Engl. 58, 6285–6289 (2019).
Kleinjan, D. A., Wardrope, C., Nga Sou, S. & Rosser, S. J. Drug-tunable multidimensional synthetic gene control using inducible degron-tagged dCas9 effectors. Nat. Commun. 8, 1191 (2017).
Maji, B. et al. Multidimensional chemical control of CRISPR-Cas9. Nat. Chem. Biol. 13, 9–11 (2017).
Iwasaki, R. S., Ozdilek, B. A., Garst, A. D., Choudhury, A. & Batey, R. T. Small molecule regulated sgRNAs enable control of genome editing in E. coli by Cas9. Nat. Commun. 11, 1394 (2020).
Kundert, K. et al. Controlling CRISPR-Cas9 with ligand-activated and ligand-deactivated sgRNAs. Nat. Commun. 10, 2127 (2019).
Tang, W., Hu, J. H. & Liu, D. R. Aptazyme-embedded guide RNAs enable ligand-responsive genome editing and transcriptional activation. Nat. Commun. 8, 15939 (2017).
Ferry, Q. R. V., Lyutova, R. & Fulga, T. A. Rational design of inducible CRISPR guide RNAs for de novo assembly of transcriptional programs. Nat. Commun. 8, 14633 (2017).
Siu, K.-H. & Chen, W. Riboregulated toehold-gated gRNA for programmable CRISPR-Cas9 function. Nat. Chem. Biol. 15, 217–220 (2019).
Oesinghaus, L. & Simmel, F. C. Switching the activity of Cas12a using guide RNA strand displacement circuits. Nat. Commun. 10, 2092 (2019).
Hanewich-Hollatz, M. H., Chen, Z., Hochrein, L. M., Huang, J. & Pierce, N. A. Conditional guide RNAs: programmable conditional regulation of CRISPR/Cas function in bacterial and mammalian cells via dynamic RNA nanotechnology. ACS Cent. Sci. 5, 1241–1249 (2019).
Nielsen, A. A. K. & Voigt, C. A. Multi-input CRISPR/Cas genetic circuits that interface host regulatory networks. Mol. Syst. Biol. 10, 763 (2014). Implementation of Cas9-based genetic circuits capable of performing logic operations in mammalian cells is discussed.
Kiani, S. et al. CRISPR transcriptional repression devices and layered circuits in mammalian cells. Nat. Methods 11, 723–726 (2014).
Nissim, L., Perli, S. D., Fridkin, A., Perez-Pinera, P. & Lu, T. K. Multiplexed and programmable regulation of gene networks with an integrated RNA and CRISPR/Cas toolkit in human cells. Mol. Cell 54, 698–710 (2014).
Nakamura, M. et al. Anti-CRISPR-mediated control of gene editing and synthetic circuits in eukaryotic cells. Nat. Commun. 10, 194 (2019). This paper highlights the utility of Acrs as gene circuit components in eukaryotic cells.
Guo, T. W. et al. Cryo-EM structures reveal mechanism and inhibition of DNA targeting by a CRISPR-Cas surveillance complex. Cell 171, 414–426.e12 (2017).
Acknowledgements
We thank T.Y. Liu for helpful discussions and proofreading the manuscript. This research was developed with funding from the Defense Advanced Research Projects Agency (DARPA) award HR0011-17-2-0043. The views, opinions, and/or findings expressed are those of the author and should not be interpreted as representing the official views or policies of the Department of Defense or the U.S. Government. This material is based upon work supported by the National Science Foundation under award number 1817593. Research reported in this publication was supported by the National Institutes of Health under award number grant award U19AI135990 (Host Pathogen Map Initiative). This research was supported by the Allen Distinguished Investigator Program, through The Paul G. Allen Frontiers Group. B.F.C. is supported by an NIH/NIGMS postdoctoral fellowship (F32 GM131654). G.J.K. is supported by an NHMRC Investigator Grant (ELI, 1175568).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The Regents of the University of California have patents issued and pending for CRISPR technologies on which H.S., B.F.C., G.J.K. and J.A.D. are inventors. J.A.D. is a cofounder of Caribou Biosciences, Editas Medicine, Scribe Therapeutics, Intellia Therapeutics and Mammoth Biosciences. J.A.D. is a scientific advisory board member of Caribou Biosciences, Intellia Therapeutics, eFFECTOR Therapeutics, Scribe Therapeutics, Mammoth Biosciences, Synthego, Algen Biotechnologies, Felix Biosciences and Inari. J.A.D. is a Director at Johnson & Johnson and has research projects sponsored by Biogen, Pfizer, AppleTree Partners and Roche.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shivram, H., Cress, B.F., Knott, G.J. et al. Controlling and enhancing CRISPR systems. Nat Chem Biol 17, 10–19 (2021). https://doi.org/10.1038/s41589-020-00700-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00700-7
This article is cited by
-
A potential paradigm in CRISPR/Cas systems delivery: at the crossroad of microalgal gene editing and algal-mediated nanoparticles
Journal of Nanobiotechnology (2023)
-
Semi-automated optimized method to isolate CRISPR/Cas9 edited human pluripotent stem cell clones
Stem Cell Research & Therapy (2023)
-
Metallic nanoplatforms for COVID-19 diagnostics: versatile applications in the pandemic and post-pandemic era
Journal of Nanobiotechnology (2023)
-
RNA-targeting CRISPR–Cas systems
Nature Reviews Microbiology (2023)
-
Programmable mammalian translational modulators by CRISPR-associated proteins
Nature Communications (2023)