Abstract
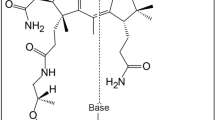
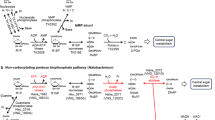
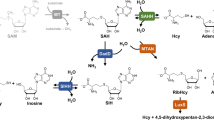
Archaeosine (G+), 7-formamidino-7-deazaguanosine, is an archaea-specific modified nucleoside found at the 15th position of tRNAs. In Euryarchaeota, 7-cyano-7-deazaguanine (preQ0)-containing tRNA (q0N-tRNA), synthesized by archaeal tRNA-guanine transglycosylase (ArcTGT), has been believed to be converted to G+-containing tRNA (G+-tRNA) by the paralog of ArcTGT, ArcS. However, we found that several euryarchaeal ArcSs have lysine transfer activity to q0N-tRNA to form q0kN-tRNA, which has a preQ0 lysine adduct as a base. Through comparative genomics and biochemical experiments, we found that ArcS forms a robust complex with a radical S-adenosylmethionine (SAM) enzyme named RaSEA. The ArcS–RaSEA complex anaerobically converted q0N-tRNA to G+-tRNA in the presence of SAM and lysine via q0kN-tRNA. We propose that ArcS and RaSEA should be considered an archaeosine synthase α-subunit (lysine transferase) and β-subunit (q0kN-tRNA lyase), respectively.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data supporting findings in this study are available from the corresponding author for reasonable requests.
References
Boccaletto, P. et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 46, D303–D307 (2018).
Hori, H. et al. Transfer RNA modification enzymes from thermophiles and their modified nucleosides in tRNA. Microorganisms 6, E110 (2018).
Kilpatrick, M. W. & Walker, R. T. The nucleotide sequence of the tRNAM Met from the archaebacterium Thermoplasma acidophilum. Nucleic Acids Res. 9, 4387–4390 (1981).
Gupta, R. Halobacterium volcanii tRNAs. Identification of 41 tRNAs covering all amino acids, and the sequences of 33 class I tRNAs. J. Biol. Chem. 259, 9461–9471 (1984).
Gregson, J. M. et al. Structure of the archaeal transfer RNA nucleoside G*-15 (2-amino-4,7-dihydro-4-oxo-7-β-d-ribofuranosyl-1H-pyrrolo[2,3-d]pyrimidine-5-carboximidamide (archaeosine)). J. Biol. Chem. 268, 10076–10086 (1993).
Noon, K. R. et al. Influence of temperature on tRNA modification in archaea: Methanococcoides burtonii (optimum growth temperature [T opt], 23 °C) and Stetteria hydrogenophila (T opt, 95 °C). J. Bacteriol. 185, 5483–5490 (2003).
Tomikawa, C. et al. Distinct tRNA modifications in the thermo-acidophilic archaeon, Thermoplasma acidophilum. FEBS Lett. 587, 3575–3580 (2013).
Oliva, R., Tramontano, A. & Cavallo, L. Mg2+ binding and archaeosine modification stabilize the G15 C48 Levitt base pair in tRNAs. RNA 13, 1427–1436 (2007).
Watanabe, M. et al. Biosynthesis of archaeosine, a novel derivative of 7-deazaguanosine specific to archaeal tRNA, proceeds via a pathway involving base replacement on the tRNA polynucleotide chain. J. Biol. Chem. 272, 20146–20151 (1997).
Bai, Y., Fox, D. T., Lacy, J. A., Van Lanen, S. G. & Iwata-Reuyl, D. Hypermodification of tRNA in thermophilic archaea. Cloning, overexpression, and characterization of tRNA-guanine transglycosylase from Methanococcus jannaschii. J. Biol. Chem. 275, 28731–28738 (2000).
Watanabe, M., Nameki, N., Matsuo-Takasaki, M., Nishimura, S. & Okada, N. tRNA recognition of tRNA-guanine transglycosylase from a hyperthermophilic archaeon, Pyrococcus horikoshii. J. Biol. Chem. 276, 2387–2394 (2001).
Ishitani, R. et al. Crystal structure of archaeosine tRNA-guanine transglycosylase. J. Mol. Biol. 318, 665–677 (2002).
Ishitani, R. et al. Alternative tertiary structure of tRNA for recognition by a posttranscriptional modification enzyme. Cell 113, 383–394 (2003).
Sabina, J. & Söll, D. The RNA-binding PUA domain of archaeal tRNA-guanine transglycosylase is not required for archaeosine formation. J. Biol. Chem. 281, 6993–7001 (2006).
Phillips, G. et al. Discovery and characterization of an amidinotransferase involved in the modification of archaeal tRNA. J. Biol. Chem. 285, 12706–12713 (2010).
Phillips, G. et al. Diversity of archaeosine synthesis in Crenarchaeota. ACS Chem. Biol. 7, 300–305 (2012).
Kawamura, T. et al. Multisite-specific archaeosine tRNA-guanine transglycosylase (ArcTGT) from Thermoplasma acidophilum, a thermo-acidophilic archaeon. Nucleic Acids Res. 44, 1894–1908 (2016).
Mei, X. et al. Crystal structure of the archaeosine synthase QueF-like—insights into amidino transfer and tRNA recognition by the tunnel fold. Proteins 85, 103–116 (2017).
Bon Ramos, A., Bao, L., Turner, B., de Crécy-Lagard, V. & Iwata-Reuyl, D. QueF-like, a non-homologous archaeosine synthase from the Crenarchaeota. Biomolecules 7, E36 (2017).
Wakita, K. et al. Higher-order structure of bovine mitochondrial tRNAPhe lacking the ‘conserved’ GG and TΨCG sequences as inferred by enzymatic and chemical probing. Nucleic Acids Res. 22, 347–353 (1994).
Hasegawa, T. & Yokogawa, T. Escherichia coli proline tRNA: structure and recognition sites for prolyl-tRNA synthetase. Nucleic Acids Symp. Ser. 44, 7–8 (2000).
Awai, T. et al. Aquifex aeolicus tRNA (N 2,N 2-guanine)-dimethyltransferase (Trm1) catalyzes transfer of methyl groups not only to guanine 26 but also to guanine 27 in tRNA. J. Biol. Chem. 284, 20467–20478 (2009).
Tomikawa, C., Yokogawa, T., Kanai, T. & Hori, H. N 7-Methylguanine at position 46 (m7G46) in tRNA from Thermus thermophilus is required for cell viability at high temperatures through a tRNA modification network. Nucleic Acids Res. 38, 942–957 (2010).
Ikeuchi, Y. et al. Agmatine-conjugated cytidine in a tRNA anticodon is essential for AUA decoding in archaea. Nat. Chem. Biol. 6, 277–282 (2010).
Ishida, K. et al. Pseudouridine at position 55 in tRNA controls the contents of other modified nucleotides for low-temperature adaptation in the extreme-thermophilic eubacterium Thermus thermophilus. Nucleic Acids Res. 39, 2304–2318 (2011).
Kawamura, T., Anraku, R., Hasegawa, T., Tomikawa, C. & Hori, H. Transfer RNA methyltransferases from Thermoplasma acidophilum, a thermoacidophilic archaeon. Int. J. Mol. Sci. 16, 91–113 (2014).
Takuma, H. et al. Substrate tRNA recognition mechanism of eubacterial tRNA (m1A58) methyltransferase (TrmI). J. Biol. Chem. 290, 5912–5925 (2015).
Nomura, Y., Ohno, S., Nishikawa, K. & Yokogawa, T. Correlation between the stability of tRNA tertiary structure and the catalytic efficiency of a tRNA-modifying enzyme, archaeal tRNA-guanine transglycosylase. Genes Cells 21, 41–52 (2016).
Yokogawa, T., Kitamura, Y., Nakamura, D., Ohno, S. & Nishikawa, K. Optimization of the hybridization-based method for purification of thermostable tRNAs in the presence of tetraalkylammonium salts. Nucleic Acids Res. 38, e89 (2010).
Nomura, Y. et al. Purification and comparison of native and recombinant tRNA-guanine transglycosylases from Methanosarcina acetivorans. Protein Expr. Purif. 88, 13–19 (2013).
Wishart, D. S. et al. HMDB: a knowledgebase for the human metabolome. Nucleic Acids Res. 37, D603–D610 (2009).
Sofia, H. J., Chen, G., Hetzler, B. G., Reyes-Spindola, J. F. & Miller, N. E. Radical SAM, a novel protein superfamily linking unresolved steps in familiar biosynthetic pathways with radical mechanisms: functional characterization using new analysis and information visualization methods. Nucleic Acids Res. 29, 1097–1106 (2001).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Sawai, H. Preparation of RPC-5 like resin for HPLC (Neosorb LC) and its use for the separation of oligonucleotides and mononucleotides. Nucleic Acids Symp. Ser. 15, 105–108 (1984).
Zhang, Y. et al. Diphthamide biosynthesis requires an organic radical generated by an iron-sulphur enzyme. Nature 465, 891–896 (2010).
Selvadurai, K., Wang, P., Seimetz, J. & Huang, R. H. Archaeal Elp3 catalyzes tRNA wobble uridine modification at C5 via a radical mechanism. Nat. Chem. Biol. 10, 810–812 (2014).
Young, A. P. & Bandarian, V. Pyruvate is the source of the two carbons that are required for formation of the imidazoline ring of 4-demethylwyosine. Biochemistry 50, 10573–10575 (2011).
Pierrel, F., Björk, G. R., Fontecave, M. & Atta, M. Enzymatic modification of tRNAs: MiaB is an iron–sulfur protein. J. Biol. Chem. 277, 13367–13370 (2002).
Arragain, S. et al. Identification of eukaryotic and prokaryotic methylthiotransferase for biosynthesis of 2-methylthio-N 6-threonylcarbamoyladenosine in tRNA. J. Biol. Chem. 285, 28425–28433 (2010).
Toh, S.-M., Xiong, L., Bae, T. & Mankin, A. S. The methyltransferase YfgB/RlmN is responsible for modification of adenosine 2503 in 23S rRNA. RNA 14, 98–106 (2008).
Benítez-Páez, A., Villarroya, M. & Armengod, M.-E. The Escherichia coli RlmN methyltransferase is a dual-specificity enzyme that modifies both rRNA and tRNA and controls translational accuracy. RNA 18, 1783–1795 (2012).
Galperin, M. Y., Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Expanded microbial genome coverage and improved protein family annotation in the COG database. Nucleic Acids Res. 43, D261–D269 (2015).
Sato, T., Fukui, T., Atomi, H. & Imanaka, T. Improved and versatile transformation system allowing multiple genetic manipulations of the hyperthermophilic archaeon Thermococcus kodakaraensis. Appl. Environ. Microbiol. 71, 3889–3899 (2005).
Sato, T., Fukui, T., Atomi, H. & Imanaka, T. Targeted gene disruption by homologous recombination in the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J. Bacteriol. 185, 210–220 (2003).
Atomi, H., Fukui, T., Kanai, T., Morikawa, M. & Imanaka, T. Description of Thermococcus kodakaraensis sp. nov., a well studied hyperthermophilic archaeon previously reported as Pyrococcus sp. KOD1. Archaea 1, 263–267 (2004).
Ochman, H., Gerber, A. S. & Hartl, D. L. Genetic applications of an inverse polymerase chain reaction. Genetics 120, 621–623 (1988).
Nagaoka, E., Hidese, R., Imanaka, T. & Fujiwara, S. Importance and determinants of induction of cold-induced DEAD RNA helicase in the hyperthermophilic archaeon Thermococcus kodakarensis. J. Bacteriol. 195, 3442–3450 (2013).
Hirata, A. et al. Archaeal RNA polymerase subunits E and F are not required for transcription in vitro, but a Thermococcus kodakarensis mutant lacking subunit F is temperature-sensitive. Mol. Microbiol. 70, 623–633 (2008).
Taniguchi, N., Nakayama, S., Kawakami, T. & Murakami, H. Patch cloning method for multiple site-directed and saturation mutagenesis. BMC Biotechnol. 13, 91 (2013).
Ikeda-Boku, A. et al. A simple system for expression of proteins containing 3-azidotyrosine at a pre-determined site in Escherichia coli. J. Biochem. 153, 317–326 (2013).
Gasteiger, E et al. in The Proteomics Protocols Handbook (ed. Walker, J.M.) Ch. 52, 571–601 (Humana Press, 2005).
Milligan, J. F., Groebe, D. R., Witherell, G. W. & Uhlenbeck, O. C. Oligoribonucleotide synthesis using T7 RNA polymerase and synthetic DNA templates. Nucleic Acids Res. 15, 8783–8798 (1987).
Nishimura, S., Harada, F., Narushima, U. & Seno, T. Purification of methionine-, valine-, phenylalanine- and tyrosine-specific tRNA from Escherichia coli. Biochim. Biophys. Acta 142, 133–148 (1967).
Ehresmann, B., Imbault, P. & Weil, J. H. Spectrophotometric determination of protein concentration in cell extracts containing tRNA’s and rRNA’s. Anal. Biochem. 54, 454–463 (1973).
Jones, B. N., Pääbo, S. & Stein, S. Amino acid analysis and enzymatic sequence determination of peptides by an improved o-phthaldialdehyde precolumn labeling procedure. J. Liq. Chromatogr. 4, 565–586 (1981).
Acknowledgements
We are grateful to T. Inuzuka, O. Sakurada, S. Obata, Y. Onda, A. Izumoto, T. Okuda, M. Osawa and H. Miyawaki for technical assistance. This work was supported by JSPS KAKENHI grants JP16H04763 (to H.H.), JP17K05929 (to N.O.) and JP18K06088 (to A.H.), and the Koshiyama Science and Technology Foundation (to T.Y.). We thank J. Allen from Edanz Group (https://www.edanzediting.com/ac/) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
T.Y. prepared the preQ0Lys-nucleotide for NMR. Y.N. and A.Y. performed the series of experiments on ArcS. H.O. prepared and analyzed tRNAs and proteins. K.H. and S.N. performed the series of experiments on M. acetivorans and T. kodakarensis ArcS–RaSEA, respectively. N.O. measured NMR of the preQ0Lys-nucleotide and synthesized q0kN under the supervision of K.A. T.K. and A.H. constructed the series of T. kodakarensis strains under the supervision of H.H. S.O. performed MS analysis. All authors discussed the results and commented on the manuscript. T.Y. designed and supervised all the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2, and Figs. 1–21
Supplementary Note
Synthetic procedures
Supplementary Dataset
Gene profiling of an enzyme involved in archaeosine synthesis
Rights and permissions
About this article
Cite this article
Yokogawa, T., Nomura, Y., Yasuda, A. et al. Identification of a radical SAM enzyme involved in the synthesis of archaeosine. Nat Chem Biol 15, 1148–1155 (2019). https://doi.org/10.1038/s41589-019-0390-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0390-7
This article is cited by
-
Solo act revealed to be duet
Nature Chemical Biology (2019)