Abstract
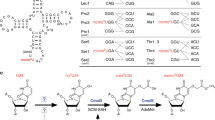
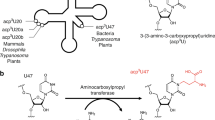
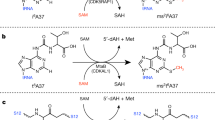
Modification of tRNA anticodons plays a critical role in ensuring accurate translation. N4-acetylcytidine (ac4C) is present at the anticodon first position (position 34) of bacterial elongator tRNAMet. Herein, we identified Bacillus subtilis ylbM (renamed tmcAL) as a novel gene responsible for ac4C34 formation. Unlike general acetyltransferases that use acetyl-CoA, TmcAL activates an acetate ion to form acetyladenylate and then catalyzes ac4C34 formation through a mechanism similar to tRNA aminoacylation. The crystal structure of TmcAL with an ATP analog reveals the molecular basis of ac4C34 formation. The ΔtmcAL strain displayed a cold-sensitive phenotype and a strong genetic interaction with tilS that encodes the enzyme responsible for synthesizing lysidine (L) at position 34 of tRNAIle to facilitate AUA decoding. Mistranslation of the AUA codon as Met in the ΔtmcAL strain upon tilS repression suggests that ac4C34 modification of tRNAMet and L34 modification of tRNAIle act cooperatively to prevent misdecoding of the AUA codon.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Suzuki, T. in Fine-tuning of RNA Functions by Modification and Editing (ed. Grosjean, H.) 24–69 (Springer-Verlag, NY, 2005).
Curran, J. F. Modification and Editing of RNA. (eds. Grosjean, H. & Benne, R.) 493–516 (ASM press, Washington, D.C., 1998).
Bjork, G. in tRNA: Structure, Biosynthesis, and Function (eds. Söll, D. & Rajbhandary, U.L.) 165–205 (ASM Press, Washington, D.C., 1995).
Suzuki, T. & Numata, T. Convergent evolution of AUA decoding in bacteria and archaea. RNA Biol. 11, 1586–1596 (2014).
Muramatsu, T. et al. Codon and amino-acid specificities of a transfer RNA are both converted by a single post-transcriptional modification. Nature 336, 179–181 (1988).
Soma, A. et al. An RNA-modifying enzyme that governs both the codon and amino acid specificities of isoleucine tRNA. Mol. Cell 12, 689–698 (2003).
Ikeuchi, Y. et al. molecular mechanism of lysidine synthesis that determines tRNA identity and codon recognition. Mol. Cell 19, 235–246 (2005).
Oashi, Z. et al. Characterization of C+located in the first position of the anticodon of Escherichia coli tRNA Met as N 4 -acetylcytidine. Biochim. Biophys. Acta 262, 209–213 (1972).
Machnicka, M. A. et al. MODOMICS: a database of RNA modification pathways--2013 update. Nucleic Acids Res. 41, D262–D267 (2013).
Kawai, G., Hashizume, T., Miyazawa, T., McCloskey, J. A. & Yokoyama, S. Conformational characteristics of 4-acetylcytidine found in tRNA. Nucleic Acids Symp. Ser. 21, 61–62 (1989).
Stern, L. & Schulman, L. H. The role of the minor base N 4-acetylcytidine in the function of the Escherichia coli noninitiator methionine transfer RNA. J. Biol. Chem. 253, 6132–6139 (1978).
Ikeuchi, Y., Kitahara, K. & Suzuki, T. The RNA acetyltransferase driven by ATP hydrolysis synthesizes N 4-acetylcytidine of tRNA anticodon. EMBO J. 27, 2194–2203 (2008).
Chimnaronk, S. et al. RNA helicase module in an acetyltransferase that modifies a specific tRNA anticodon. EMBO J. 28, 1362–1373 (2009).
Ito, S. et al. A single acetylation of 18S rRNA is essential for biogenesis of the small ribosomal subunit in Saccharomyces cerevisiae. J. Biol. Chem. 289, 26201–26212 (2014).
Ito, S. et al. Human NAT10 is an ATP-dependent RNA acetyltransferase responsible for N4 -acetylcytidine formation in 18S ribosomal RNA (rRNA). J. Biol. Chem. 289, 35724–35730 (2014).
Sharma, S. et al. Yeast Kre33 and human NAT10 are conserved 18S rRNA cytosine acetyltransferases that modify tRNAs assisted by the adaptor Tan1/THUMPD1. Nucleic Acids Res. 43, 2242–2258 (2015).
Miyauchi, K., Ohara, T. & Suzuki, T. Automated parallel isolation of multiple species of non-coding RNAs by the reciprocal circulating chromatography method. Nucleic Acids Res. 35, e24 (2007).
Suzuki, T., Ikeuchi, Y., Noma, A., Suzuki, T. & Sakaguchi, Y. Mass spectrometric identification and characterization of RNA-modifying enzymes. Methods Enzymol. 425, 211–229 (2007).
Ohira, T. & Suzuki, T. Precursors of tRNAs are stabilized by methylguanosine cap structures. Nat. Chem. Biol. 12, 648–655 (2016).
Yamada, Y. & Ishikura, H. Nucleotide sequence of non-initiator methionine tRNA from Bacillus subtilis. Nucleic Acids Res. 8, 4517–4520 (1980).
Andachi, Y., Yamao, F., Muto, A. & Osawa, S. Codon recognition patterns as deduced from sequences of the complete set of transfer RNA species in Mycoplasma capricolum. Resemblance to mitochondria. J. Mol. Biol. 209, 37–54 (1989).
Markowitz, V. M. et al. IMG: the Integrated Microbial Genomes database and comparative analysis system. Nucleic Acids Res. 40, D115–D122 (2012).
Markowitz, V. M. et al. IMG 4 version of the integrated microbial genomes comparative analysis system. Nucleic Acids Res. 42, D560–D567 (2014).
Galperin, M. Y., Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Expanded microbial genome coverage and improved protein family annotation in the COG database. Nucleic Acids Res. 43, D261–D269 (2015).
Bork, P., Holm, L., Koonin, E. V. & Sander, C. The cytidylyltransferase superfamily: identification of the nucleotide-binding site and fold prediction. Proteins 22, 259–266 (1995).
Aravind, L., Anantharaman, V. & Koonin, E. V. Monophyly of class I aminoacyl tRNA synthetase, USPA, ETFP, photolyase, and PP-ATPase nucleotide-binding domains: implications for protein evolution in the RNA. Proteins 48, 1–14 (2002).
Izard, T. The crystal structures of phosphopantetheine adenylyltransferase with bound substrates reveal the enzyme’s catalytic mechanism. J. Mol. Biol. 315, 487–495 (2002).
Brick, P., Bhat, T. N. & Blow, D. M. Structure of tyrosyl-tRNA synthetase refined at 2.3A resolution. Interaction of the enzyme with the tyrosyl adenylate intermediate. J. Mol. Biol. 208, 83–98 (1989).
Ullrich, T. C., Blaesse, M. & Huber, R. Crystal structure of ATP sulfurylase from Saccharomyces cerevisiae, a key enzyme in sulfate activation. EMBO J. 20, 316–329 (2001).
Ilyin, V. A. et al. 2.9A crystal structure of ligand-free tryptophanyl-tRNA synthetase: domain movements fragment the adenine nucleotide binding site. Protein Sci. 9, 218–231 (2000).
D’Angelo, I. et al. Structure of nicotinamide mononucleotide adenylyltransferase: a key enzyme in NAD(+) biosynthesis. Structure 8, 993–1004 (2000).
Suzuki, T. & Miyauchi, K. Discovery and characterization of tRNAIle lysidine synthetase (TilS). FEBS Lett. 584, 272–277 (2010).
Kramer, E. B. & Farabaugh, P. J. The frequency of translational misreading errors in E. coli is largely determined by tRNA competition. RNA 13, 87–96 (2007).
Ogasawara, N. Systematic function analysis of Bacillus subtilis genes. Res. Microbiol. 151, 129–134 (2000).
Marchler-Bauer, A. et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 45(D1), D200–D203 (2017).
Schmellenkamp, H. & Eggerer, H. Mechanism of enzymic acetylation of des-acetyl citrate lyase. Proc. Natl Acad. Sci. USA 71, 1987–1991 (1974).
Urbonavicius, J., Skouloubris, S., Myllykallio, H. & Grosjean, H. Identification of a novel gene encoding a flavin-dependent tRNA:m5U methyltransferase in bacteria--evolutionary implications. Nucleic Acids Res. 33, 3955–3964 (2005).
Puri, P. et al. Systematic identification of tRNAome and its dynamics in Lactococcus lactis. Mol. Microbiol. 93, 944–956 (2014).
Moghal, A., Mohler, K. & Ibba, M. Mistranslation of the genetic code. FEBS Lett. 588, 4305–4310 (2014).
Suzuki, T., Ueda, T. & Watanabe, K. The ‘polysemous’ codon--a codon with multiple amino acid assignment caused by dual specificity of tRNA identity. EMBO J. 16, 1122–1134 (1997).
Grosjean, H. & Björk, G. R. Enzymatic conversion of cytidine to lysidine in anticodon of bacterial isoleucyl-tRNA--an alternative way of RNA editing. Trends Biochem. Sci. 29, 165–168 (2004).
Nagao, A. et al. Hydroxylation of a conserved tRNA modification establishes non-universal genetic code in echinoderm mitochondria. Nat. Struct. Mol. Biol. 24, 778–782 (2017).
Hirsh, D. Tryptophan tRNA of Escherichia coli. Nature 228, 57 (1970).
Schmeing, T. M., Voorhees, R. M., Kelley, A. C. & Ramakrishnan, V. How mutations in tRNA distant from the anticodon affect the fidelity of decoding. Nat. Struct. Mol. Biol. 18, 432–436 (2011).
Sprinzl, M. & Vassilenko, K. S. Compilation of tRNA sequences and sequences of tRNA genes. Nucleic Acids Res. 33, D139–D140 (2005).
Aluotto, B. B., Wittler, R. G., Williams, C. O. & Faber, J. E. Standardized bacteriologic techniques for the characterization of Mycoplasma species. Int. J. Syst. Bacteriol. 20, 35–58 (1970).
Taniguchi, T. et al. Decoding system for the AUA codon by tRNAIle with the UAU anticodon in Mycoplasma mobile. Nucleic Acids Res. 41, 2621–2631 (2013).
Sato, T., Harada, K. & Kobayashi, Y. Analysis of suppressor mutations of spoIVCA mutations: occurrence of DNA rearrangement in the absence of site-specific DNA recombinase SpoIVCA in Bacillus subtilis. J. Bacteriol. 178, 3380–3383 (1996).
Morimoto, T. et al. Six GTP-binding proteins of the Era/Obg family are essential for cell growth in Bacillus subtilis. Microbiology 148, 3539–3552 (2002).
Ashikaga, S., Nanamiya, H., Ohashi, Y. & Kawamura, F. Natural genetic competence in Bacillus subtilis natto OK2. J. Bacteriol. 182, 2411–2415 (2000).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Terwilliger, T. C. Maximum-likelihood density modification. Acta Crystallogr. D Biol. Crystallogr. 56, 965–972 (2000).
Terwilliger, T. C. Automated main-chain model building by template matching and iterative fragment extension. Acta Crystallogr. D Biol. Crystallogr. 59, 38–44 (2003).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, (213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Sampson, J. R. & Uhlenbeck, O. C. Biochemical and physical characterization of an unmodified yeast phenylalanine transfer RNA transcribed in vitro. Proc. Natl Acad. Sci. USA 85, 1033–1037 (1988).
Coleman, T. M. & Huang, F. RNA-catalyzed thioester synthesis. Chem. Biol. 9, 1227–1236 (2002).
Arai, T. et al. Single methylation of 23S rRNA triggers late steps of 50S ribosomal subunit assembly. Proc. Natl Acad. Sci. USA 112, E4707–E4716 (2015).
Acknowledgements
We thank members of the Suzuki laboratory, especially S. Kimura, for fruitful discussion and many helpful suggestions. We also thank the beamline staff at BL-17A of the Photon Factory for technical assistance during data collection, and M. Miyata in Osaka City University for kindly giving us the culture of M. mobile. This work was supported by the Grants-in-Aid for Scientific Research on Priority Areas from the Ministry of Education, Culture, Sports, Science, and Technology of Japan (26113003, 26220205, 18H05272 to T.S.; 18K05430 to K.M.; and 26113002, 18H03980 to K.T.), by a JSPS Fellowship for Japanese Junior Scientists (to T.T.), and by the Noda Institute for Scientific Research (to A.S.).
Author information
Authors and Affiliations
Contributions
T.T. mainly performed the series of experiments. K.M. assisted biochemical and informatics works. Y.S. performed MS analysis. S.Y. and K.T. performed structural studies. A.S. assisted genetic works. All authors discussed the results. T.T. and T.S. wrote this paper. T.S. designed and supervised all the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–16, Supplementary Tables 1–5
Rights and permissions
About this article
Cite this article
Taniguchi, T., Miyauchi, K., Sakaguchi, Y. et al. Acetate-dependent tRNA acetylation required for decoding fidelity in protein synthesis. Nat Chem Biol 14, 1010–1020 (2018). https://doi.org/10.1038/s41589-018-0119-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0119-z
This article is cited by
-
Dissecting the oncogenic properties of essential RNA-modifying enzymes: a focus on NAT10
Oncogene (2024)
-
Quality control of protein synthesis in the early elongation stage
Nature Communications (2023)
-
Regulation of the epigenome through RNA modifications
Chromosoma (2023)
-
Amniotic fluid mesenchymal stem cells repair mouse corneal cold injury by promoting mRNA N4-acetylcytidine modification and ETV4/JUN/CCND2 signal axis activation
Human Cell (2021)
-
YqfB protein from Escherichia coli: an atypical amidohydrolase active towards N4-acylcytosine derivatives
Scientific Reports (2020)