Abstract
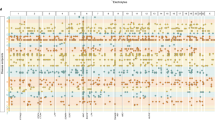
Autoimmune and inflammatory diseases are polygenic disorders of the immune system. Many genomic loci harbor risk alleles for several diseases, but the limited resolution of genetic mapping prevents determining whether the same allele is responsible, indicating a shared underlying mechanism. Here, using a collection of 129,058 cases and controls across 6 diseases, we show that ~40% of overlapping associations are due to the same allele. We improve fine-mapping resolution for shared alleles twofold by combining cases and controls across diseases, allowing us to identify more expression quantitative trait loci driven by the shared alleles. The patterns indicate widespread sharing of pathogenic mechanisms but not a single global autoimmune mechanism. Our approach can be applied to any set of traits and is particularly valuable as sample collections become depleted.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
This paper analyzes existing, publicly available data. The accession numbers for these datasets are listed in Supplementary Table 2.
Code availability
Code used in this analysis is available on GitHub at https://github.com/cotsapaslab/CrossDiseaseImmunochip (ref. 80) and archived on Zenodo at https://doi.org/10.5281/zenodo.8371032 (ref. 81).
References
Rosenblum, M. D., Remedios, K. A. & Abbas, A. K. Mechanisms of human autoimmunity. J. Clin. Invest. 125, 2228–2233 (2015).
International Multiple Sclerosis Genetics Consortium. Multiple sclerosis genomic map implicates peripheral immune cells and microglia in susceptibility. Science 365, eaav7188 (2019).
de Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Gusev, A. et al. Partitioning heritability of regulatory and cell-type-specific variants across 11 common diseases. Am. J. Hum. Genet. 95, 535–552 (2014).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Hemminki, K., Li, X., Sundquist, K. & Sundquist, J. Shared familial aggregation of susceptibility to autoimmune diseases. Arthritis Rheum. 60, 2845–2847 (2009).
Eaton, W. W., Rose, N. R., Kalaydjian, A., Pedersen, M. G. & Mortensen, P. B. Epidemiology of autoimmune diseases in Denmark. J. Autoimmun. 29, 1–9 (2007).
Kuo, C.-F. et al. Familial aggregation of systemic lupus erythematosus and coaggregation of autoimmune diseases in affected families. JAMA Intern. Med. 175, 1518–1526 (2015).
Cotsapas, C. et al. Pervasive sharing of genetic effects in autoimmune disease. PLoS Genet. 7, e1002254 (2011).
Ellinghaus, D. et al. Analysis of five chronic inflammatory diseases identifies 27 new associations and highlights disease-specific patterns at shared loci. Nat. Genet. 48, 510–518 (2016).
Sirota, M., Schaub, M. A., Batzoglou, S., Robinson, W. H. & Butte, A. J. Autoimmune disease classification by inverse association with SNP alleles. PLoS Genet. 5, e1000792 (2009).
Shirai, Y. et al. Multi-trait and cross-population genome-wide association studies across autoimmune and allergic diseases identify shared and distinct genetic component. Ann. Rheum. Dis. 81, 1301–1312 (2022).
Pouget, J. G. et al. Cross-disorder analysis of schizophrenia and 19 immune-mediated diseases identifies shared genetic risk. Hum. Mol. Genet. 28, 3498–3513 (2019).
Fortune, M. D. et al. Statistical colocalization of genetic risk variants for related autoimmune diseases in the context of common controls. Nat. Genet. 47, 839–846 (2015).
Parkes, M., Cortes, A., van Heel, D. A. & Brown, M. A. Genetic insights into common pathways and complex relationships among immune-mediated diseases. Nat. Rev. Genet. 14, 661–673 (2013).
Burren, O. S. et al. Genetic feature engineering enables characterisation of shared risk factors in immune-mediated diseases. Genome Med. 12, 106 (2020).
van de Bunt, M. et al. Evaluating the performance of fine-mapping strategies at common variant GWAS loci. PLoS Genet. 11, e1005535 (2015).
Turley, P. et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat. Genet. 50, 229–237 (2018).
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Zhernakova, A., Withoff, S. & Wijmenga, C. Clinical implications of shared genetics and pathogenesis in autoimmune diseases. Nat. Rev. Endocrinol. 9, 646–659 (2013).
Matzaraki, V., Kumar, V., Wijmenga, C. & Zhernakova, A. The MHC locus and genetic susceptibility to autoimmune and infectious diseases. Genome Biol. 18, 76 (2017).
Cortes, A. & Brown, M. A. Promise and pitfalls of the ImmunoChip. Arthritis Res. Ther. 13, 101 (2011).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Benner, C. et al. FINEMAP: efficient variable selection using summary data from genome-wide association studies. Bioinformatics 32, 1493–1501 (2016).
Mohanan, V. et al. C1orf106 is a colitis risk gene that regulates stability of epithelial adherens junctions. Science 359, 1161–1166 (2018).
Rivas, M. A. et al. Deep resequencing of GWAS loci identifies independent rare variants associated with inflammatory bowel disease. Nat. Genet. 43, 1066–1073 (2011).
Farh, K. K.-H. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
Trynka, G. et al. Chromatin marks identify critical cell types for fine mapping complex trait variants. Nat. Genet. 45, 124–130 (2013).
Chun, S. et al. Limited statistical evidence for shared genetic effects of eQTLs and autoimmune-disease-associated loci in three major immune-cell types. Nat. Genet. 49, 600–605 (2017).
Chen, L. et al. Genetic drivers of epigenetic and transcriptional variation in human immune cells. Cell 167, 1398–1414.e24 (2016).
Pappalardo, J. L. et al. Transcriptomic and clonal characterization of T cells in the human central nervous system. Sci. Immunol. 5, eabb8786 (2020).
Kumar, S., Ambrosini, G. & Bucher, P. SNP2TFBS—a database of regulatory SNPs affecting predicted transcription factor binding site affinity. Nucleic Acids Res. 45, D139–D144 (2017).
Kundu, K. et al. Genetic associations at regulatory phenotypes improve fine-mapping of causal variants for 12 immune-mediated diseases. Nat. Genet. 54, 251–262 (2022).
Foley, C. N. et al. A fast and efficient colocalization algorithm for identifying shared genetic risk factors across multiple traits. Nat. Commun. 12, 764 (2021).
Hernández, N. et al. The flashfm approach for fine-mapping multiple quantitative traits. Nat. Commun. 12, 6147 (2021).
Umans, B. D., Battle, A. & Gilad, Y. Where are the disease-associated eQTLs? Trends Genet. 37, 109–124 (2021).
The Lenercept Multiple Sclerosis Study Group & The University of British Columbia MS/MRI Analysis Group. TNF neutralization in MS: results of a randomized, placebo-controlled multicenter study. Neurology 53, 457–465 (1999).
Gregory, A. P. et al. TNF receptor 1 genetic risk mirrors outcome of anti-TNF therapy in multiple sclerosis. Nature 488, 508–511 (2012).
Buniello, A. et al. The NHGRI-EBI GWAS catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012 (2019).
Paternoster, L. et al. Multi-ancestry genome-wide association study of 21,000 cases and 95,000 controls identifies new risk loci for atopic dermatitis. Nat. Genet. 47, 1449–1456 (2015).
Ferreira, M. A. et al. Shared genetic origin of asthma, hay fever and eczema elucidates allergic disease biology. Nat. Genet. 49, 1752–1757 (2017).
Han, Y. et al. Genome-wide analysis highlights contribution of immune system pathways to the genetic architecture of asthma. Nat. Commun. 11, 1776 (2020).
Dubois, P. C. A. et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 42, 295–302 (2010).
Lyons, P. A. et al. Genome-wide association study of eosinophilic granulomatosis with polyangiitis reveals genomic loci stratified by ANCA status. Nat. Commun. 10, 5120 (2019).
Bronson, P. G. et al. Common variants at PVT1, ATG13–AMBRA1, AHI1 and CLEC16A are associated with selective IgA deficiency. Nat. Genet. 48, 1425–1429 (2016).
Cousminer, D. L. et al. First genome-wide association study of latent autoimmune diabetes in adults reveals novel insights linking immune and metabolic diabetes. Diabetes Care 41, 2396–2403 (2018).
Cordell, H. J. et al. International genome-wide meta-analysis identifies new primary biliary cirrhosis risk loci and targetable pathogenic pathways. Nat. Commun. 6, 8019 (2015).
Ji, S.-G. et al. Genome-wide association study of primary sclerosing cholangitis identifies new risk loci and quantifies the genetic relationship with inflammatory bowel disease. Nat. Genet. 49, 269–273 (2017).
Aterido, A. et al. Genetic variation at the glycosaminoglycan metabolism pathway contributes to the risk of psoriatic arthritis but not psoriasis. Ann. Rheum. Dis. 78, 355–364 (2019).
Bentham, J. et al. Genetic association analyses implicate aberrant regulation of innate and adaptive immunity genes in the pathogenesis of systemic lupus erythematosus. Nat. Genet. 47, 1457–1464 (2015).
López-Isac, E. et al. GWAS for systemic sclerosis identifies multiple risk loci and highlights fibrotic and vasculopathy pathways. Nat. Commun. 10, 4955 (2019).
Jin, Y. et al. Genome-wide association studies of autoimmune vitiligo identify 23 new risk loci and highlight key pathways and regulatory variants. Nat. Genet. 48, 1418–1424 (2016).
Feng, B.-J. et al. Multiple loci within the major histocompatibility complex confer risk of psoriasis. PLoS Genet. 5, e1000606 (2009).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
International HapMap 3 Consortium et al. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
Trynka, G. et al. Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nat. Genet. 43, 1193–1201 (2011).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
International Multiple Sclerosis Genetics Consortium (IMSGC) et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat. Genet. 45, 1353–1360 (2013).
Eyre, S. et al. High-density genetic mapping identifies new susceptibility loci for rheumatoid arthritis. Nat. Genet. 44, 1336–1340 (2012).
Langefeld, C. D. et al. Transancestral mapping and genetic load in systemic lupus erythematosus. Nat. Commun. 8, 16021 (2017).
Zhao, J. et al. A missense variant in NCF1 is associated with susceptibility to multiple autoimmune diseases. Nat. Genet. 49, 433–437 (2017).
Onengut-Gumuscu, S. et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat. Genet. 47, 381–386 (2015).
Rasmussen, A. et al. The lupus family registry and repository. Rheumatology 50, 47–59 (2011).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, 7 (2015).
Scrucca, L., Fop, M., Murphy, T. B. & Raftery, A. E. mclust 5: clustering, classification and density estimation using Gaussian finite mixture models. R J. 8, 289–317 (2016).
Abraham, G. & Inouye, M. Fast principal component analysis of large-scale genome-wide data. PLoS ONE 9, e93766 (2014).
Delaneau, O., Zagury, J.-F. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods 10, 5–6 (2013).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Viechtbauer, W. Conducting meta-analyses in R with the metafor package. J. Stat. Softw. 36, 1–48 (2010).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2009).
Stegle, O., Parts, L., Durbin, R. & Winn, J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLoS Comput. Biol. 6, e1000770 (2010).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
cotsapaslab / CrossDiseaseImmunochip. GitHub https://github.com/cotsapaslab/CrossDiseaseImmunochip (2021).
Lincold, M. R. matthewlincoln/xd-release: release. Zenodo https://doi.org/10.5281/zenodo.8371032 (2023).
Acknowledgements
We thank the EAGLE eczema consortium for providing GWAS summary statistics. This research utilizes resources provided by the T1DGC, a collaborative clinical study sponsored by the National Institute of Diabetes and Digestive and Kidney Diseases, National Institute of Allergy and Infectious Diseases, National Human Genome Research Institute, National Institute of Child Health and Human Development and JDRF and supported by U01 DK062418. We thank the RACI consortium for access to RA data and the International IBD Genetics Consortium for access to IBD data. Deidentified data were provided from a total of 4,617 samples (2,563 SLE cases and 2,054 population controls) in the Lupus Family Registry and Repository collection at the Oklahoma Medical Research Foundation. The SLE Genentech samples were originally genotyped and analyzed as part of a large SLEGEN Consortium ImmunoChip study. The Alliance for Lupus Research (now Lupus Research Alliance) provided funds for the SLE ImmunoChip study. M.R.L. is supported by a Career Transition Fellowship from the Consortium of MS Centers and the National MS Society (TA-2206-39622) and an Early Career Award in MS from the Waugh Family Foundation. C.G. received a research fellowship from the Deutsche Forschungsgemeinschaft (German Research Foundation) for this project. She further received funding from the Hans und Klementia Langmatz-Stifung and the Hertie Network of Excellence in Clinical Neuroscience, not related to this study. C.W. was supported by an ERC advanced grant (FP/2007-2013/ERC grant 2012-322698), the Spinoza prize grant (NWO SPI 92-266), a grant from Stiftelsen K. G. Jebsen and the Netherlands Organ-on-Chip Initiative—an NWO Gravitation project (024.003.001) funded by the Ministry of Education, Culture and Science of the government of the Netherlands. S.W. was supported by the Netherlands Organ-on-Chip Initiative, an NWO Gravitation project (024.003.001) funded by the Ministry of Education, Culture and Science of the government of the Netherlands. I.H.J. is supported by a Rosalind Franklin Fellowship from the University of Groningen and an NWO VIDI grant (no. 016.171.047).
Author information
Authors and Affiliations
Consortia
Contributions
M.R.L., N.C., P.-P.A., C.G. and M.M. curated and analyzed data. D.v.H., C.W., S.W., I.H.J., L.P., International Multiple Sclerosis Genetics Consortium, S.S.R., R.R.G., P.M.G., C.D.L., T.J.V. and D.A.H. provided data. M.R.L. and C.C. wrote and edited the paper, with input from all co-authors. S.C. and S.R.S. designed and implemented analytical methods. C.C. conceptualized and oversaw the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Carl Anderson, Anne Barton and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–18 and Note.
Supplementary Table
Supplementary Tables 1–4.
Supplementary Data 1
Accounting of study participants.
Supplementary Data 2
Accounting of SNPs.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lincoln, M.R., Connally, N., Axisa, PP. et al. Genetic mapping across autoimmune diseases reveals shared associations and mechanisms. Nat Genet 56, 838–845 (2024). https://doi.org/10.1038/s41588-024-01732-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-024-01732-8