Abstract
Cancer genetics has uncovered many tumor-suppressor and oncogenic pathways, but few alterations have revealed mechanisms involved in tumor spreading. Here, we examined the role of the third most significant chromosomal deletion in human melanoma that inactivates the adherens junction gene NECTIN1 in 55% of cases. We found that NECTIN1 loss stimulates melanoma cell migration in vitro and spreading in vivo in both zebrafish and human tumors specifically in response to decreased IGF1 signaling. In human melanoma biopsy specimens, adherens junctions were seen exclusively in areas with low IGF1 levels, but not in NECTIN1-deficient tumors. Our study establishes NECTIN1 as a major determinant of melanoma dissemination and uncovers a genetic control of the response to microenvironmental signals.
Similar content being viewed by others
Main
A clinical challenge in solid tumors remains the formation of metastases1. In melanoma, tumor dissemination is the leading cause of mortality2. Although few studies have implicated genetic lesions in tumor dissemination3, early work in pancreatic cancer detected metastatic clones within parental primary tumors and suggested the absence of a genetic signature of metastasis4. More recently, the genomic comparison of metastases and matched primary tumors failed to identify genetic drivers of tumor spreading5,6,7, pointing to a role for non-cancer cell-autonomous mechanisms in metastasis8. The relative contributions of genetic alterations and signals from the tumor microenvironment to metastatic traits remain controversial9.
Early events implicated in local tumor invasion or detachment of cells from the primary tumor, such as alterations of cell–cell adhesion10, have been extensively studied in epithelial tumors. For example E-cadherin loss has been reported to promote metastasis in pancreatic cancer11, and epithelial-to-mesenchymal transition (EMT) serves as an early driver of metastasis12,13. However, the role of cell–cell adhesion remains elusive in non-epithelial cancers like melanoma. Melanocytes themselves emerge from the neural crest during embryonic development and thus possess more mesenchymal than epithelial features14. Loss of E-cadherin expression has been shown to favor melanoma progression by disrupting interactions between melanocytes and neighboring keratinocytes in coculture experiments15, and differences in cadherin patterns, notably a switch from E-cadherin to N-cadherin, have been noted in melanoma metastases compared to primary tumors16. Yet, functional evidence explaining the contribution of cell–cell adhesion changes to the metastatic dissemination of non-epithelial tumors is lacking.
Here, we identified deletions of the gene NECTIN1 as a frequent driver of melanoma dissemination. NECTIN1 is an adhesion molecule that can engage in homotypic or heterotypic interactions between two cells17,18 to establish early adherens junctions19. We found that NECTIN1 loss changes the response of melanoma cells to microenvironmental insulin-like growth factor 1 (IGF1) signaling by controlling a switch from cell–cell adhesion to cell–matrix adhesion. This mechanism integrates the status of adherens junctions and local growth factor concentration to direct the decision of cancer cells to stay in the niche or to leave.
Results
NECTIN1 is frequently deleted in human melanoma
By analyzing copy-number alterations20,21 in The Cancer Genome Atlas (TCGA) cohort of 363 human melanomas, we found that the third most significant focal deletion at chr11q23.3 only contained the gene NECTIN1 (Fig. 1a and Extended Data Fig. 1a,b). Although biallelic loss of NECTIN1 was detected in 4.4% of cases (16 cases), half of melanomas exhibited shallow deletions (186 cases) totaling an overall frequency of deletion of 55% (Fig. 1b). Most deletions affected the entire locus, with shallow deletions likely taking out one NECTIN1 allele and resulting in heterozygous loss (Extended Data Fig. 1c,d). NECTIN1 loss did not correlate with any alterations in major oncogenes (BRAF and NRAS) or tumor-suppressor genes (NF1, CDKN2A, TP53 and PTEN) (Extended Data Fig. 1e). NECTIN1 mRNA expression correlated with linear copy number (Extended Data Fig. 2a), with human melanoma samples with deep or shallow deletions of NECTIN1 expressing significantly less NECTIN1 mRNA than diploid samples (Extended Data Fig. 2b). Immunohistochemistry for NECTIN1 in human melanoma revealed that NECTIN1 protein levels were significantly lower in metastases compared to primary tumors (Fig. 1c,d and Extended Data Fig. 2c). Metastases were overrepresented among samples with low NECTIN1 staining whereas samples with high NECTIN1 were enriched for primary melanomas (Extended Data Fig. 2d). In human melanoma, NECTIN1 status did not correlate with the expression of E-cadherin, another adherens junction component whose loss has been linked to metastatic progression in multiple cancers (Extended Data Fig. 2e). Coimmunofluorescence in 20 human melanoma tissue sections also failed to reveal any correlation between NECTIN1 and E-cadherin protein levels (Extended Data Fig. 2f,g). Importantly, NECTIN1 deletions were associated with lower overall survival of patients with cutaneous melanoma (Fig. 1e). Together, these data nominate NECTIN1 as a frequently altered gene in melanoma with a potential role in metastasis.
a, List of the five most significant chromosomal deletions found in TCGA dataset of 363 human cutaneous melanomas. False discovery rate q-values were calculated by GISTIC. b, Linear copy number of the NECTIN1 locus in human melanoma (363 samples represented on the y axis). Blue denotes deletions, red, amplifications. The gray bar on the scale represents the focal deletion found by GISTIC analysis. c, Images of human melanoma sections stained for NECTIN1 by immunohistochemistry. Scale bar, 20 μm. d, Quantification of overall NECTIN1 staining intensity in 180 primary melanomas and 73 melanoma metastases (mean ± standard deviation (s.d.). Two-tailed t-test. e, Overall survival curves of patients with melanoma stratified according to NECTIN1 copy number status (> −0.25: no deletion, n = 190; < −0.25: deletion, n = 168 (log-rank test).
NECTIN1 loss promotes melanoma spreading in vivo
We inactivated nectin1 in primary zebrafish melanomas with a tissue-specific CRISPR technique22,23 (Fig. 2a and Methods). NECTIN1 has two orthologs in zebrafish, nectin1a and nectin1b, of which only nectin1b is substantially expressed in primary zebrafish melanomas (Extended Data Fig. 3a). To prevent compensation between the two paralogs, we targeted both concomitantly. Sequencing of 65 primary zebrafish melanomas demonstrated high targeting efficiency at both loci (hereafter together referred to as nectin1) (Fig. 2b–d and Supplementary Note).
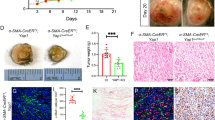
a, Schematic representation of the generation of genetically engineered primary melanomas in adult zebrafish (Methods). b, Proportion of nectin1a and nectin1b mutant alleles in 65 primary zebrafish melanomas expressing a CRISPR vector targeting nectin1a and nectin1b. Bar represents median; paired two-tailed t-test. c, Box-and-whisker plot representing the number of nectin1a and nectin1b mutant alleles per primary zebrafish melanoma described in panel b (min-max, 1st and 4th quartiles, median, n = 65; paired two-tailed t-test). d, Type and proportion of nectin1a and nectin1b mutant alleles in 10 representative primary zebrafish melanomas. D, deletion, I, insertion; numbers denote affected base pairs. The last section in each bar represents all other minor alleles. e, Tumor-free survival curves of Tg(mitf:BRAFV600E);tp53−/− zebrafish injected with vectors targeting either nectin1 or a control gene (pooled data of three independent experiments; log-rank test). f, Representative images of sections of control- or nectin1-knockout (KO) primary zebrafish melanomas, stained for phospho-histone3 (p-H3) by immunohistochemistry (IHC). Scale bar, 20 μm. g, Quantification of overall p-H3 staining intensity in 8 nectin1-wildype and 10 nectin1-knockout primary zebrafish melanomas (mean ± s.d.; two-tailed t-test). NS, not significant. h, Pictures of organs presenting with disseminated tumor cells (arrowheads) in hematoxylin/eosin-stained sections of 16-week-old zebrafish bearing nectin1-knockout melanoma. Scale bar, 20 μm. i, Number of 16-week-old zebrafish with disseminated melanoma cells in the indicated organs. j, Quantification of organ involvement in 16-week-old zebrafish bearing control (n = 11) or nectin1-knockout (n = 10) melanoma (mean ± s.d.). Two-tailed t-test. k, Schematic representation of the melanoma spreading assay in adult zebrafish (top) and representative images of casper zebrafish 7 or 21 days after transplantation with control- or nectin1-knockout melanoma cells (bottom). Insets show ×2 magnification views. Arrows indicate patches of disseminated melanoma cells. Scale bar, 1 cm. l, Quantification of the proportion of secondary recipients of control- (n = 10) or nectin1-knockout (n = 9) tumors showing tumor spreading (mean ± s.d.; two-tailed t-test). m, Cell count of control- and nectin1-knockout primary zebrafish melanoma lines in the presence or absence of zebrafish (zf) culture media. Data are presented as mean ± s.d. of three independent experiments.
Nectin1 inactivation in primary zebrafish melanoma modestly accelerated tumor onset (Fig. 2e). It did not significantly affect melanoma cell proliferation as assessed by immunohistochemistry for phospho-histone3 (Fig. 2f,g). By examining hematoxylin/eosin-stained longitudinal sections of 16-week-old adult zebrafish bearing primary melanomas, we assessed local tumor invasion and dissemination to distant organs including gills, thymus, kidney, spleen, liver, and intestine (Fig. 2h). Although no differences were detected in local tumor invasion into adjacent muscle, we observed significantly higher organ involvement with nectin1-knockout tumors than with control tumors (Fig. 2i,j). To evaluate the metastatic propensity of genetically-modified zebrafish melanoma cells in vivo, we used a subcutaneous transplantation assay24, where pigmented primary zebrafish melanoma cells can migrate away from the injection site along the sides of transparent adult recipients, which we call ‘spreading’ (Fig. 2k). This method enables direct visualization of cancer cell behavior in a physiologically relevant environment25. Primary melanoma cells expressing a control CRISPR vector only rarely spread in secondary recipients (Fig. 2k,l). In contrast, nectin1 inactivation significantly increased tumor cell spreading, as evidenced by the proportion of recipients showing patches of melanoma cells that had disseminated past the midline (Fig. 2k,l, Extended Data Fig. 3b and Supplementary Note). These results demonstrate that nectin1 loss promotes melanoma dissemination in vivo.
NECTIN1 loss alters cell adhesion upon serum depletion
The spreading capacity of zebrafish melanoma cells was associated with altered mechanical properties of primary tumors: nectin1-deficient primary zebrafish melanomas were significantly softer than their wild-type counterparts, as measured by atomic force microscopy (Extended Data Fig. 3c). To further investigate the features of nectin1-deficient tumors, we established cell lines from primary melanomas following a protocol involving the progressive replacement of the initial zebrafish-specific rich media by regular culture media25. Nectin1-knockout cell lines failed to passage outside of zebrafish media, as evidenced by the sharp drop in the proportion of nectin1 CRISPR mutant alleles compared to wild-type alleles over time (Fig. 2m and Extended Data Fig. 3d). This was due at least in part to a failure to adhere to the culture dish or to each other, since cells harvested from culture supernatant could reattach and grow in zebrafish media (Extended Data Fig. 3e). Moreover, nectin1-deficient melanoma cells displayed altered spheroid formation capacities in low-attachment conditions in the absence of zebrafish media, compared to nectin1-wild-type cells (Extended Data Fig. 3f). These observations suggest a link between cell adhesion and environmental factors in nectin1-deficient melanoma cells.
NECTIN1 inactivation induces human melanoma cell migration
We inactivated NECTIN1 by short hairpin RNA (shRNA) or CRISPR in the A375 human melanoma cell line (Extended Data Fig. 3g–j and Methods). Consistent with our observations in zebrafish cells, NECTIN1-deficient cells appeared more spread out than their wild-type counterparts in low-attachment conditions in the absence, but not in the presence of serum, which translated into the formation of significantly less circular colonies, suggesting either weaker cell–cell interactions or increased cell adhesion to culture surfaces (Fig. 3a,b and Extended Data Fig. 4a–d). This indicated that serum withdrawal elicited a change in melanoma cell adhesion. In the absence of serum, NECTIN1-deficient colonies were significantly more dispersed in collagen-rich matrix, implying that NECTIN1 loss increases the spreading capacity of melanoma cells in a 3D environment (Fig. 3c,d). NECTIN1-deficient cells migrated significantly more than their wild-type counterparts in a modified transwell assay where cells were plated without serum for 12 h before the establishment of the chemotactic gradient (Fig. 3e and Extended Data Fig. 4e). Supporting a critical role of serum starvation in this phenotype, the difference in migration between NECTIN1-deficient and proficient A375 cells was enhanced by longer starvation (Fig. 3e). Inducible expression of a NECTIN1 construct mutated in the shRNA target sequence demonstrated partial rescue of this migration phenotype (Extended Data Figs. 3g and 4f). By adding Matrigel to transwells, similar differences were measured between NECTIN1-wild-type and deficient cells, indicating that NECTIN1 loss increases both 2D migration and 3D invasion of melanoma cells (Fig. 3f). To evaluate whether the effects of NECTIN1 inactivation were specific to A375 cells, we downregulated NECTIN1 in five additional human melanoma cell lines spanning the spectrum of melanocytic differentiation (from AXLhigh/MITFlow to AXLlow/MITFhigh) and belonging to the major phenotypic subtypes of melanoma: undifferentiated, neural crest-like and melanocytic26 (Extended Data Fig. 4g). These lines exhibited various levels of NECTIN1 expression and different basal migratory capacities (Extended Data Fig. 4h,i). NECTIN1 inactivation significantly increased melanoma cell migration following serum withdrawal in all cell lines (Fig. 3g and Extended Data Fig. 4i), indicating that the function of NECTIN1 does not depend on melanoma differentiation state or basal migratory capacity.
a, Images of the structures formed by 10,000 A375 human melanoma cells stably expressing a control shRNA (shCTRL) or an shRNA directed against NECTIN1 (shNECTIN1) after 8 days in low-attachment conditions in the presence or absence of serum (FBS). Scale bar, 200 μm. Data are representative of nine independent experiments. b, Circularity of the colonies formed by A375 human melanoma cells in low-attachment conditions as shown in panel a (Methods). Data are presented as mean ± s.d. of nine independent colonies per condition (paired two-tailed t-test). c, Images of the colonies formed by 1,000 of the cells described in panel a after 7 days in collagen-rich matrix in the absence of serum (FBS) (top). Scale bar, 50 μm. Data are representative of three independent experiments. Analysis of the images shown on top using ImageJ (bottom). d, Spread of the colonies formed by A375 human melanoma cells in collagen-rich matrix as shown in panel c (Methods). Data are presented as mean ± s.d. (n = 41 (shCTRL) and n = 45 (shNECTIN1)) of three independent experiments (two-tailed t-test). a.u., arbitrary units. e, Migration of A375 human melanoma cells stably expressing an shRNA directed against NECTIN1 (shNECTIN1) relative to cells expressing a control shRNA (shCTRL) in a transwell assay after 12 or 24 h of serum starvation. Data are presented as mean ± s.d. of four independent experiments. Cells were allowed to migrate for 6 h (paired two-tailed t-test). f, Invasion through Matrigel of A375 human melanoma cells after 12 h of serum starvation as described in panel e. Cells were allowed to migrate for 10 h. Data are presented as mean ± s.d. of four independent experiments (paired two-tailed t-test). g, Migration of six human melanoma cell lines after 12 h of serum starvation as described in panel e. Cells were allowed to migrate for different times depending on the cell line (Extended Data Fig. 4i and Methods). Data are presented as mean ± s.d. of four independent experiments (paired two-tailed t-test).
Cell migration is due to failure to form adherens junctions
We next examined cell–cell contacts between melanoma cells. Immunofluorescence in A375 cells with various cell–cell junction markers revealed that serum depletion triggers the robust formation of adherens junctions between NECTIN1-wild-type cells, as assessed by α-catenin or N-cadherin staining (Fig. 4a and Extended Data Fig. 5a,b). In contrast, NECTIN1-deficient cells failed to form adherens junctions, in accordance with the previously reported requirement of NECTIN1 for their establishment27 (Fig. 4a,b and Extended Data Fig. 5b). Failure to form adherens junctions was associated with increased stress fiber formation, cytoskeleton reorganization and changes in cell shape, as evidenced by F-actin and α-actinin staining and consistent with the migratory behavior of these cells (Fig. 4a,c and Extended Data Fig. 5b,c). We hypothesized that the inability of NECTIN1-deficient cells to establish strong cell–cell contacts under stress conditions might cause their migration. Indeed, limiting cell–cell interactions between NECTIN1-wild-type melanoma cells in transwell assays, either by seeding them at low density or by subjecting them to the chemotactic gradient before they could form a cell layer, increased their migratory capacity to a level similar to that of NECTIN1-knockout cells (Fig. 4d and Extended Data Fig. 5d), implying that the formation of adherens junctions inhibits migration. To evaluate the potential contribution of cell proliferation, we measured the growth of melanoma cells upon NECTIN1 manipulation. NECTIN1-deficient cells displayed a lower proliferation rate in 2D culture (Fig. 4e and Extended Data Fig. 5e), as well as reduced colony formation capacity in semi-solid media (Fig. 4f,g and Extended Data Fig. 5f) compared to NECTIN1-wild-type cells, consistent with a previously noted balance between proliferation and migration in melanoma28. Together, these results show that NECTIN1 loss prevents adherens junction formation between melanoma cells and promotes their migration specifically under serum depletion.
a, Immunofluorescence analysis of α-catenin (green) and F-actin (red) in A375 human melanoma cells stably expressing a control shRNA (shCTRL) or an shRNA directed against NECTIN1 (shNECTIN1) cultured in the presence or absence of serum (FBS) for 12 h. Scale bar, 10 μm. Representative images of five independent experiments are shown. DAPI, 4,6-diamidino-2-phenylindole (blue), b, Quantification of adherens junction (AJ) formation as measured by α-catenin signal intensity on immunofluorescence images as shown in panel a. Data represent mean ± s.d. (n = 4 fields per condition; paired two-tailed t-test). c, Quantification of cell surface area based on immunofluorescence images as shown in panel a. Data represent mean ± s.d. (n = 21, 38, 18, 27 cells for shCTRL+FBS, shCTRL-FBS, shNECTIN1+FBS, and shNECTIN1-FBS, respectively; paired two-tailed t-test). d, Migration of three NECTIN1-knockout (KO) A375 human melanoma cell lines relative to three NECTIN1-wild-type (WT) A375 human melanoma cell lines in a transwell assay after 12 h of serum starvation. Cells were either allowed to settle for 12 h (pre-seeding) or not before being put in the presence of the chemotactic gradient. Cells were allowed to migrate for 6 h. Data represent mean ± s.d. of three independent experiments (paired two-tailed t-test). e, Proliferation of the cells described in panel a. Data represent mean ± s.d. of six independent experiments (paired two-tailed t-test). f, Representative images of colonies formed in Matrigel by the cells described in panel a after 14 days in culture. Scale bar, 1 mm. g, Number of colonies quantified from the experiments described in panel f. Data represent mean ± s.d. of six independent experiments (paired two-tailed t-test).
NECTIN1 loss activates integrin-dependent matrix adhesion
To define the changes in cell state associated with NECTIN1 inactivation, we compared the transcriptional profiles of NECTIN1-knockout and NECTIN1-wild-type A375 cells after serum depletion. Gene Ontology analysis of the 156 differentially regulated genes revealed the prevalence of biological process terms related to cell adhesion and migration (Extended Data Fig. 6a). A more detailed gene set enrichment analysis of the same comparison uncovered specific adhesion signatures including integrin αvβ3 and focal adhesion pathways (Extended Data Fig. 6b,c and Supplementary Table 1). That gene expression mirrors the cellular phenotypes described above suggests that the migration of NECTIN1-deficient melanoma cells and the formation of adherens junctions between NECTIN1-wild-type melanoma cells correspond to specific transcriptional programs.
We examined the cell-surface makeup of NECTIN1-deficient and proficient A375 cells upon serum depletion by purifying cell-surface proteins and analyzing them by quantitative mass spectrometry. Integrins (in particular integrins α6β4 and α5β1) dominated the most significantly enriched proteins in NECTIN1-deficient cells, whereas N-cadherin (CDH2), the main cadherin expressed in A375 cells, was among the most downregulated proteins at the cell surface, consistent with the absence of adherens junctions (Fig. 5a). Accordingly, adhesion of A375 cells to several extracellular matrix (ECM) proteins, including fibronectin, laminin and, to a lesser extent, vitronectin, but not collagen-I, was significantly increased in the absence of NECTIN1 (Fig. 5b–e). Laminin is the major ligand of integrin α6β4, whereas vitronectin binds to integrins αvβ3 and αvβ5, and fibronectin to integrins α5β1 and αvβ3, among others29. We systematically downregulated each beta integrin by short interfering RNA (siRNA) in NECTIN1-deficient cells and measured the impact of these genetic manipulations on cell migration in the modified transwell assay (Fig. 5f). Knocking down integrins β3, β4 and β5 substantially reduced the migration of NECTIN1-deficient cells, whereas downregulating integrins β1 and β2 had a milder effect (Fig. 5g). Moreover, an antibody blocking integrin α6β4, as well as small-molecule inhibitors of integrins αvβ3 and αvβ5, significantly decreased the migration of A375 cells lacking NECTIN1 (Fig. 5h,i), directly implicating these integrins in the motility of adherens junction-incompetent melanoma cells. These data indicate that NECTIN1-deficient melanoma cells switch to an integrin-dependent cell–matrix adhesion program in response to serum deprivation.
a, Volcano plot representing the significance and relative abundance of cell-surface proteins in A375 human melanoma cells in the presence or absence of NECTIN1. A P value (two-tailed t-test) threshold of 0.05 was chosen (red: P < 0.05, black: P > 0.05). Some of the most differentially expressed proteins are indicated. b–e, Adhesion of A375 human melanoma cells stably expressing an shRNA directed against NECTIN1 (shNECTIN1) relative to cells expressing a control shRNA (shCTRL) to collagen-I-, fibronectin-, laminin- or vitronectin-coated surfaces at various timepoints after seeding. Data represent mean ± s.d. of four independent experiments (paired two-tailed t-test). f, Western blot analysis of ITGB levels in the cells described in panel b transfected with a control siRNA (C) or siRNAs targeting ITGB1 (1), ITGB2 (2), ITGB3 (3), ITGB4 (4) or ITGB5 (5). Data are representative of four independent experiments. g, Migration of the cells described in panel f relative to cells stably expressing a control shRNA (shCTRL) and transfected with a control siRNA in a transwell assay after 12 h of serum starvation. Cells were allowed to migrate for 6 h. Data represent mean ± s.d. of four independent experiments (paired two-tailed t-test; P values are shown for the comparisons siITGB versus siCTRL (shNECTIN1)). h, Migration of A375 human melanoma cells stably expressing an shRNA directed against NECTIN1 (shNECTIN1) relative to cells expressing a control shRNA (shCTRL) in a transwell assay upon treatment with an integrin α6β4 blocking antibody (ITG Ab, GoH3, 40 μg ml−1). Cells were allowed to migrate for 6 h. Data represent mean ± s.d. of four independent experiments (paired two-tailed t-test). IgG, immunoglobulin G. i, Migration of A375 human melanoma cells stably expressing an shRNA directed against NECTIN1 (shNECTIN1) relative to cells expressing a control shRNA (shCTRL) in a transwell assay upon treatment with two integrin αvβ3 and αvβ5 inhibitors (ITGi#1: SB273005; ITGi#2: echistatin). Cells were allowed to migrate for 6 h. Data represent mean ± s.d. of four independent experiments (paired two-tailed t-test).
Similar migration differences were observed between NECTIN1-deficient and proficient cells in the presence or absence of different ECM proteins (Fig. 3e,f and Extended Data Fig. 6d), suggesting that, in transwell assays, matrix is dispensable for the migration of melanoma cells lacking NECTIN1. This apparent paradox could be explained by the synthesis of several ECM proteins by melanoma cells themselves. Indeed, RNA-sequencing data indicated that A375 human melanoma cells express high levels of fibronectin (FN1) and several laminin genes (Extended Data Fig. 6e). We also observed strong fibronectin and laminin staining of these cells cultured in transwell conditions by immunofluorescence (Extended Data Fig. 6f). Using antibodies previously shown to recognize zebrafish laminin and fibronectin, we further detected extensive laminin and fibronectin matrices around melanoma cells in the transplantation area of recipient fish, suggesting that these matrices could also play a role in the in vivo spreading assay (Extended Data Fig. 6g,h). Our functional data thus support the idea that, to migrate and disseminate, NECTIN1-deficient cells at least in part rely on interactions with the ECM that they may have deposited themselves.
NECTIN1-deficient cells migrate via the FAK/SRC pathway
We next measured the phosphorylation level of a panel of kinases upon serum depletion using a phospho-kinase array (Extended Data Fig. 7a). Focal adhesion kinase (FAK, also known as PTK2) was the most differentially activated kinase between NECTIN1-deficient and proficient A375 melanoma cells (Extended Data Fig. 7b). To identify the pathway mediating migration of NECTIN1-deficient cells under serum starvation, we performed a small-scale chemical screen with inhibitors of migration-related pathways using the modified transwell assay. The only molecules that notably suppressed the migration of NECTIN1-deficient cells were FAK and SRC inhibitors (Fig. 6a and Extended Data Fig. 7c). These findings were validated using a second set of inhibitors (Extended Data Fig. 6d), as well as genetic downregulation of FAK and SRC by siRNAs (Extended Data Fig. 7e,f). The repressive effect of FAK and SRC inhibitors was also seen on the invasion of NECTIN1-deficient cells in semi-solid media (Extended Data Fig. 7g). FAK blockade suppressed the migration of four out of five additional human melanoma cell lines upon NECTIN1 downregulation (Fig. 6b), demonstrating that most melanomas lacking NECTIN1 depend on FAK signaling to migrate. Accordingly, we verified the differential activation of FAK and SRC by phosphorylation between NECTIN1-deficient and proficient A375 cells by Western Blotting (Fig. 6c). Using pharmacological inhibition of integrins αvβ3 and αvβ5, FAK and SRC, we also confirmed the sequential activation of the three members of the pathway in A375 human melanoma cells (Extended Data Fig. 7h). In line with these observations, nectin1-knockout primary zebrafish melanoma cells exhibited much higher levels of phospho-fak than nectin1-wild-type cells in the zebrafish tumor spreading assay, as evidenced by immunohistochemistry (Fig. 6d,e and Extended Data Fig. 8a,b). 36% (10/28) of transplanted tumors displayed a staining pattern in the shape of a gradient, with stronger signal towards the invasive edge than directly under the skin (Extended Data Fig. 8c). We targeted the two zebrafish orthologs of FAK, fak1a and fak1b (hereafter collectively referred to as fak), in addition to nectin1 in zebrafish melanocytes to generate fak/nectin1-knockout melanomas in vivo (Extended Data Fig. 8d and Supplementary Note). Fak inactivation did not affect melanoma onset in zebrafish (Extended Data Fig. 8e) or change proliferation rates in primary tumors, as assessed by phospho-Histone3 immunohistochemistry (Extended Data Fig. 8f). However, examination of hematoxylin/eosin-stained tissue sections revealed a lower organ involvement upon fak knockout in adult zebrafish bearing nectin1-knockout primary melanomas (Fig. 6f,g), supporting a critical role for FAK in distant seeding by NECTIN1-deficient melanoma cells. Genetic inactivation of fak in primary zebrafish melanomas lacking nectin1 significantly reduced cell dissemination in the zebrafish tumor spreading assay (Fig. 6h, i), confirming the importance of FAK signaling. Surprisingly, western blot analysis of NECTIN1 and phospho-FAK revealed a positive correlation between FAK activity and NECTIN1 levels in our panel of 6 human melanoma cell lines (Extended Data Fig. 8g,h). However, in three out of four cell lines with relatively high NECTIN1 expression, pharmacological inhibition of FAK induced a marked decrease in NECTIN1 levels (Extended Data Fig. 8i), suggesting feedback regulation of NECTIN1 expression by FAK activity. Note that this regulatory loop is likely hindered in cases of NECTIN1 genetic inactivation. Our results show that upon serum deprivation, NECTIN1-deficient cells activate an integrin/FAK/SRC axis fostering cell motility and spreading.
a, Migration of A375 human melanoma cells stably expressing an shRNA directed against NECTIN1 (shNECTIN1) relative to cells expressing a control shRNA (shCTRL) in a transwell assay upon treatment with a FAK inhibitor (FAKi, PF562271) or a SRC inhibitor (SRCi, dasatinib). Cells were allowed to migrate for 6 h. Data represent mean ± s.d. of three independent experiments. Paired two-tailed t-test. b, Migration of six human melanoma cell lines stably expressing an shRNA directed against NECTIN1 (shNECTIN1) relative to cells expressing a control shRNA (shCTRL) in a transwell assay upon treatment with two independent FAK inhibitors (FAKi#1, PF562271; FAKi#2, GSK2256098). Cells were allowed to migrate for different times depending on the cell line (Extended Data Fig. 4i and Methods). Data represent mean ± s.d. of four independent experiments (paired two-tailed t-test). c, Western blot analysis of FAK and SRC phosphorylation levels in the cells described in panel a in the absence of serum. C, shCTRL; Sh, shNECTIN1. Representative example of five independent experiments. Signal intensity was normalized to GAPDH and presented as a ratio to the vehicle-treated shCTRL condition. Data represent mean ± s.d. (paired two-tailed t-test). d, Representative images of sections of adult casper recipients transplanted with control- or nectin1-knockout primary zebrafish melanomas, stained for phospho-fak by immunohistochemistry. Scale bar, 50 μm. e, Quantification of phospho-fak (p-fak) staining intensity in casper recipients transplanted with nectin1-wild-type (n = 19) or nectin1-knockout (n = 28) zebrafish melanomas. Data represent mean ± s.d. (two-tailed t-test). f, Number of 16-week-old zebrafish with disseminated melanoma cells in the indicated organs. g, Quantification of organ involvement in 16-week-old zebrafish bearing nectin1/ctrl-knockout (n = 9) or nectin1/fak-knockout (n = 9) melanoma. Data represent mean ± s.d. (two-tailed t-test). h, Representative images of adult casper zebrafish transplanted with nectin1-knockout primary zebrafish melanoma cells expressing a control gRNA (ctrl-knockout) or a gRNA targeting fak (fak-knockout). Insets show ×2 magnification views. Arrowheads indicate patches of disseminated melanoma cells. Scale bar, 1 cm. i, Quantification of the proportion of secondary recipients of nectin1/ctrl-knockout (ctrl-KO, n = 7) or nectin1/fak-knockout (fak-KO, n = 6) tumors showing tumor spreading. Data represent mean ± s.d. (two-tailed t-test).
A drop in IGF1 elicits NECTIN1 loss-of-function phenotypes
We sought to identify the serum components responsible for the difference in cell behavior between NECTIN1-proficient and deficient melanoma cells. Culture media supplemented with dextran-treated charcoal-stripped serum, which contains nutrients but is depleted in hormones and growth factors, triggered adherens junction formation between NECTIN1-wild-type, but not NECTIN1-deficient, A375 human melanoma cells to the same extent as serum-free media, as assessed by immunofluorescence (Extended Data Fig. 9a), pointing to the lack of growth factors or hormones as the likely cause of this difference. Similar to serum withdrawal, dextran-treated charcoal-stripped serum also activated FAK/SRC signaling in cells lacking NECTIN1 (Extended Data Fig. 9b). We examined the transcriptional response of A375 cells to serum depletion by RNA sequencing. This response was dominated by proliferation arrest and responses to external stimuli, as indicated by Gene Ontology analysis (Extended Data Fig. 9c and Supplementary Table 2). Gene set enrichment analysis revealed a number of growth factor and hormone signaling pathways underlying this response to external stimuli (Extended Data Fig. 9d and Supplementary Table 3), which allowed us to establish a list of 12 candidate secreted factors (Extended Data Fig. 9e) that we tested for their ability to prevent adherens junction formation upon serum withdrawal as detected by immunofluorescence for the adherens junction marker α-catenin. Only IGF1 and, to a lesser extent, insulin scored in this assay (Fig. 7a and Extended Data Fig. 9f). The expression of a constitutively active form of the IGF1 receptor (IGF1R*)30 recapitulated this effect (Fig. 7b and Extended Data Fig. 9g). Conversely, small-molecule inhibitors of IGF1R triggered adherens junction formation between A375 cells even in the presence of serum (Fig. 7c), demonstrating that the absence of IGF1 signaling is both necessary and sufficient to elicit the behavioral changes observed in melanoma cells. Moreover, forced IGF1 signaling partially reversed the migratory propensity of NECTIN1-deficient cells in the modified transwell assay in 6 different human melanoma cell lines (Fig. 7d), similar to direct treatment with IGF1 itself (Fig. 7e). These results were confirmed in vivo by expressing IGF1R* in nectin1-deficient zebrafish melanomas, which accelerated tumor onset compared to a GFP-negative control (Extended Data Fig. 10a), in line with the generally protumorigenic role of aberrant IGF1 signaling, although we did not detect significant differences in tumor cell proliferation by phospho-Histone3 staining of zebrafish melanoma sections (Extended Data Fig. 10b). Analysis of tissue sections of adult zebrafish bearing nectin1-knockout tumors expressing either GFP or IGF1R* did not reveal any differences in organ involvement (Extended Data Fig. 10c), likely due to IGF1R*-expressing tumors forming earlier than GFP-expressing ones and having more time to invade adjacent tissues and disseminate in the fish. However, expression of IGF1R* abolished the dissemination of nectin1-knockout zebrafish melanoma cells upon subcutaneous transplantation into adult recipients (Fig. 7f,g). These data define a functional interaction between IGF1 signaling and adherens junctions in the regulation of melanoma cell behavior, with a decrease in IGF1 eliciting the formation of cell–cell contacts between NECTIN1-wild-type adherens junctions-competent cells or the migration of NECTIN1-deficient adherens junctions-impaired cells.
a, Immunofluorescence analysis of α-catenin (green) and F-actin (red) in A375 human melanoma cells in the presence or absence of serum (FBS) (blue, DAPI), complemented or not with 100 ng ml−1 IGF1 for 12 h. Scale bar, 10 μm. Data are representative of four independent experiments. b, Immunofluorescence analysis of α-catenin (green) and F-actin (red) in A375 human melanoma cells expressing or not a constitutively active form of the IGF1 receptor (CD8-IGF1R, IGF1R*), and cultured in the absence of serum (FBS) for 12 h (blue, DAPI). Scale bar, 10 μm. Data are representative of three independent experiments. c, Immunofluorescence analysis of α-catenin (green) and F-actin (red) in A375 human melanoma cells cultured in the presence of serum (FBS) and treated with DMSO (vehicle) or with two different IGF1R inhibitors (IGF1Ri#1: Linsitinib, 1 μM; IGF1Ri#2: GSK1838705A, 1 μM) for 12 h (blue, DAPI). Scale bar, 10 μm. Data are representative of four independent experiments. d, Migration of six human melanoma cell lines conditionally expressing GFP or a constitutively active form of IGF1R (CD8-IGF1R, IGF1R*), and stably expressing an shRNA directed against NECTIN1 (shNECTIN1) relative to cells expressing a control shRNA (shCTRL) in a transwell assay. Cells were allowed to migrate for different times depending on the cell line (Extended Data Fig. 4i and Methods). Data represent mean ± s.d. of four independent experiments (paired two-tailed t-test). e, Migration of A375 human melanoma cells stably expressing either a control shRNA (shCTRL) or an shRNA directed against NECTIN1 (shNECTIN1) in a transwell assay after 12 h of serum starvation in the presence or absence of 100 ng ml−1 IGF1. Cells were allowed to migrate for 6 h. Data represent mean ± s.d. of three independent experiments (paired two-tailed t-test). f, Representative images of adult casper zebrafish transplanted with nectin1-knockout primary zebrafish melanoma cells expressing GFP or a constitutively active form of IGF1R (CD8-IGF1R, IGF1R*), 21 days after injection. Insets show ×2 magnification views. Arrowhead indicates patches of disseminated melanoma cells. Scale bar, 1 cm. g, Quantification of the proportion of secondary recipients of nectin1-knockout tumors expressing GFP (n = 6) or nectin1-knockout tumors expressing CD8-IGF1R (IGF1R*) (n = 5) showing tumor spreading. Data represent mean ± s.d. (two-tailed t-test).
Adherens junctions occur in areas of low IGF1 signaling
In humans, low IGF1R expression was associated with reduced progression-free survival in the TCGA cohort of cutaneous melanoma (Extended Data Fig. 10d). We therefore assessed the interaction between IGF1 signaling and adherens junctions in human melanoma in vivo. We performed immunofluorescence in tissue sections of 20 human melanoma biopsies with markers of active IGF1 signaling (phospho-IGF1R) and adherens junctions (α-catenin) and independently evaluated NECTIN1 expression. We noted heterogeneity in IGF1 signaling, some areas of tumors showing weak or no phospho-IGF1R straining whereas others exhibited strong signals, especially near blood vessels. Adherens junctions were only observed in areas that lacked IGF1 signaling (Fig. 8a and Extended Data Fig. 10e). The patterns of phospho-IGF1R and α-catenin were generally mutually exclusive as demonstrated by the anti-correlation between the fluorescence intensity of the two signals (Fig. 8b and Extended Data Fig. 10f), except in the case of endothelial cells lining blood vessels. Adherens junctions almost exclusively occurred in tumors that expressed NECTIN1 (Fig. 8c,d). The only exception was in a melanoma that stained negative for NECTIN1 but expressed E-cadherin, which likely explains the ability to form adherens junctions. These results indicate that adherens junctions are formed by melanoma cells specifically in response to a drop in IGF1 signaling and suggest that the growth factor variations that elicit changes in melanoma cell behavior happen locally in human tumors.
a, Immunofluorescence analysis of α-catenin (green) and phospho-IGF1R (red) on a tissue section of human melanoma (blue, DAPI). Scale bar, 10 μm. Dotted line demarcates areas of high and low IGF1 signaling. Arrows point to adherens junctions. A representative example of nine independent tumors is shown. b, Diagram representing signal intensities of α-catenin (green) and phospho-IGF1R (red) in the section shown in panel a (Pearson’s correlation). c, Immunofluorescence analysis of E-cadherin (green) and NECTIN1 (red) (blue, DAPI) on a tissue section of the same tumor as in panel a. Scale bar, 10 μm. BV, blood vessel. d, Table showing the distribution of 20 tissue sections of human melanoma depending on the presence or absence of adherens junctions (as assessed by α-catenin staining) and NECTIN1 by immunofluorescence. Correlation was measured by chi-squared test. Note that the only NECTIN1-negative tumor exhibiting adherens junctions was positive for E-cadherin. e–g, Model for the role of NECTIN1 in melanoma. e, In the presence of IGF1, melanoma cells actively repress the formation of adherens junctions and proliferate, independent of NECTIN1 status. f, In the absence of IGF1, NECTIN1-wild-type melanoma cells form robust adherens junctions (red) and rely on cell–cell adhesion to stay in the niche. g, In the absence of IGF1, NECTIN1-deficient melanoma cells (that are unable to establish adherens junctions) switch to cell–matrix adhesion and activate an integrin/FAK/SRC axis, leading to their migration and dissemination.
Discussion
We identified NECTIN1 as a major metastasis-suppressor gene in melanoma. The fact that NECTIN1 downregulation by shRNA and complete NECTIN1 ablation by CRISPR elicited similar cellular phenotypes indicates that both homozygous and hemizygous deletions of NECTIN1, together detected in 55% of patients with melanoma, promote metastasis and that NECTIN1 acts as a haploinsufficient metastasis-suppressor gene. In addition to melanoma, NECTIN1 is frequently deleted in bladder, uterine and cervical cancers in TCGA datasets. Whether NECTIN1 controls similar mechanisms in these tumor types remains to be determined.
Our data show that NECTIN1 status determines the response of melanoma cells to environmental stress, specifically a drop in local IGF1 concentration. In the presence of high levels of IGF1, melanoma cells proliferate within the cancer cell niche regardless of NECTIN1 status (Fig. 8e), as demonstrated by previous reports examining the role of IGF1 in early melanoma lesions31. In low IGF1 conditions and in the presence of NECTIN1, melanoma cells form robust cell–cell contacts through adherens junctions, which prevent them from leaving the niche (Fig. 8f). In the absence of NECTIN1, they switch to cell–matrix adhesion and migrate away by activating an integrin/FAK/SRC pathway, leading to tumor dissemination (Fig. 8g). The switch from cell–cell to cell–matrix adhesion may be explained by a previously suggested balance between NECTIN abundance and integrin activity32. Our observations implicating local variations in IGF1 signaling in the regulation of melanoma cell adhesion and behavior are in line with reports indicating that IGF1 signaling decreases cell–cell adhesion and promotes EMT in several epithelial tumors33,34. In melanoma, inhibition of IGF1 has been shown to reduce proliferation and invasion by blocking an EMT-like program35. That low IGF1 increases epithelial features of melanoma cells is consistent with our data. Additionally, our results provide an explanation to the apparent paradox regarding the effect of IGF1 inhibition on migration and invasion by suggesting the effect is governed by the status of cell–cell adhesion, particularly adherens junctions; in cells capable of establishing cell–cell contacts, low IGF1 signaling forces adherens junction formation and prevents migration whereas it triggers the dissemination of cells that are unable to make cell–cell junctions. By identifying a frequent chromosomal deletion that sensitizes cancer cells to local signals in the tumor microenvironment, our findings reconcile the genetic and non-cell-autonomous views of metastasis in melanoma.
The interplay between cell adhesion and external signals may prove especially relevant to non-epithelial tumors like melanoma. The loss of cell–cell contacts, especially of E-cadherin-dependent adherens junctions, occurs in the early phases of the metastatic progression of epithelial tumors36. Melanocytes, however, arise from the neural crest, an embryonic cell layer that delaminates from the neural tube by EMT during vertebrate development and therefore do not usually form adherens junctions14. Melanoma cells appear closer in their behavior to the developing neural crest than to organized epithelia37,38. Interestingly, NECTIN1 inactivating mutations in humans cause cleft lip or cleft palate39 reminiscent of the disruption of craniofacial development in neurocristopathies. Moreover, insulin signaling promotes neural crest induction in vitro40. The consequences of NECTIN1 inactivation appear independent of the differentiation state of melanoma cells and the induction of adherens junctions by IGF1 depletion could be observed in cells that expressed N-cadherin and not E-cadherin, a feature usually associated with a more mesenchymal state41. Our findings thus uncover a mode of regulation of non-epithelial cancer cell migration.
Previous reports have suggested that melanoma cells can oscillate between proliferative and migratory states, a phenomenon known as plasticity28. Our study characterizes genes and mechanisms of the transition to the migratory state, which we found to be initiated by local changes in growth factor availability. The mechanism described here may have evolved to allow cells to survive stress situations, either by establishing contacts with their neighbors to receive supportive signals or by moving away in search of a more favorable environment. This model suggests that melanoma cells successfully metastasize to sites that provide higher levels of IGF1 and that cells that can adapt to variable levels of IGF1 signaling might constitute more efficient metastasizers. The capacity to adapt to local stress may represent a major asset for cancer cells to successfully complete the invasion-metastasis cascade, as previously shown with oxidative stress42. This plasticity of cancer cells can also impinge on their response to treatment as a more migratory state has been associated with drug resistance43.
Our study identifies the genetic determinants of a mechanism whereby cancer cells integrate two types of external signals from their local environment–physical cues sensed through adhesion molecules and chemical cues sensed by growth factor receptors–to govern cell behaviors. This mechanism sheds light on the regulation of cellular decisions in the cancer cell niche that may prove therapeutically actionable.
Methods
The research conducted in the present study complies with all relevant ethical regulations.
Genomic data analysis
NECTIN1 was identified in a frequent focal deletion by GISTIC analysis of the TCGA cohort of 363 patients with melanoma20,21. The overall survival curves of patients presenting or not NECTIN1 deletions were generated by stratifying the survival data of the TCGA cohort of cutaneous melanoma according to NECTIN1 linear copy number with a threshold set at −0.25. The progression-free survival curves of patients with high or low IGF1R expression were generated by stratifying the survival data of the TCGA cohort of cutaneous melanoma according to IGF1R expression z-score and comparing the top and bottom quartiles.
Human melanoma samples
This study was conducted with approval of the institutional review board of Brigham and Women’s Partners Human Research Committee, Harvard Medical School. All pathology specimens were archived samples of the Department of Pathology at Brigham and Women’s Hospital. All samples are deidentified (that is, personally identifiable information and identifiers were not retained and cannot be retrieved). As such, the institutional review board deemed that seeking further informed consent was not necessary.
All human melanomas were formalin fixed, paraffin embedded, sectioned and stained with hematoxylin and eosin for histopathological evaluation. The hematoxylin and eosin-stained sections were independently reviewed and confirmed by a dermatopathologist (C.G.L.). In total, 122 cases were studied retrospectively for NECTIN1 immunohistochemistry (melanocytic nevi (n = 20), primary melanoma (n = 72) and metastatic melanoma (n = 30)), and 20 tissue sections of primary and metastatic melanoma were used for immunofluorescence studies. In addition, two commercial human melanoma tissue-microarrays were purchased from Biomax (ME1004e and ME2082c).
Immunohistochemistry staining
Immunohistochemistry studies were performed on paraffin-embedded sections of formalin-fixed tissue. For staining of human melanoma tissues, tissue was deparaffinized, rehydrated and blocked with 3% peroxide. Heat-induced antigen retrieval was completed using Target Antigen Retrieval solution (Dako) in a pressure cooker for 45 min. Sections were blocked with 10% animal serum for 30 min and incubated with NECTIN1 primary antibody (1:50; HPA026846, Sigma-Aldrich) at 4 °C overnight. Sections were then incubated with goat anti-rabbit horseradish peroxidase (HRP) (1:200, Vector Laboratories) secondary antibody for an hour for visualization and then processed with DAB + Chromagen (Dako) and hematoxylin counterstain. For staining of zebrafish melanoma tissues, immunohistochemistry was performed on the Leica Biosystems Bond III automated staining platform. Phospho-Histone3 (Ser10) antibody (#9701, Cell Signaling Technology) or phospho-FAK (Tyr397) antibody (44–624 G, Invitrogen) were run at 1:200 dilution using the Leica Biosystems Refine Detection Kit with citrate antigen retrieval, after bleaching of the melanin pigment. Immunostains were scored as previously described44 (0 = no staining, 1 = weak and incomplete staining in <10% of targeted cells, 2 = weak/moderate heterogeneous staining in >10% of targeted cells and 3 = strong and complete homogenous staining in >10% of targeted cells). Data analysis was done using GraphPad Prism.
Human melanoma cell lines
293 T, RPMI7951, CJM, A375, SKMEL2, COLO679, and G361 cell lines were purchased from commercial sources (ATCC CRL-3216, ATCC HTB-66, Creative Bioarray CSC-C6421J, ATCC CRL-1619, ATCC HTB-68, Sigma 87061210, and ATCC CRL-1424, respectively) and cultured in DMEM (Life Technologies), except for G361 (McCoy’s media, 10% FBS (Atlanta Biologicals), 1X GlutaMAX (Life Technologies) and 1% penicillin-streptomycin (Life Technologies)). Cells were grown at 37 °C in 5% CO2. All cultures were regularly checked for the presence of mycoplasma. For cell culture experiments implicating genetic or pharmacological modifications, cells were randomly distributed between the different experimental conditions.
Gene inactivation
Genes were inactivated in primary zebrafish melanomas using a technique that we previously developed22,23. This technique is based on CRISPR MiniCoopR, a DNA plasmid containing one or two U6:gRNA cassettes targeting the gene(s) of interest and a zebrafish Cas9 sequence under the control of the melanocyte-specific mitf promoter. The Tol2 transposase technology45 permits insertion of this vector into the zebrafish genome upon microinjection into one-cell stage embryos. Stable expression subsequently results in the CRISPR inactivation of the gene of interest specifically in melanocytes. Use of this technique in a melanoma-prone zebrafish line, such as Tg(mitfa:BRAFV600E), tp53−/−, mitfa−/−, allows to study the role of a gene in genetically engineered primary tumors.
To inactivate NECTIN1 in human melanoma cells, we selected an shRNA consistently reducing endogenous NECTIN1 protein levels by over 50% to mimic the effect of deletions observed in human patients (Extended Data Fig. 3g). We also established NECTIN1 CRISPR knockout lines from individual clones of A375 cells transduced with a lentiviral CRISPR construct targeting early exons of the gene. Knockout was confirmed by the presence of frameshift indels at the target sites by deep-sequencing (Extended Data Fig. 3h) and absence of the protein by western blotting (Extended Data Fig. 3i). Of note, NECTIN1 mRNA level was also reduced in these knockout lines compared to wild-type parental lines, presumably due to non-sense mediated decay (Extended Data Fig. 3j).
DNA constructs
Two control vectors were used in this study, one containing a non-targeting gRNA and the other targeting arhgap11a, a gene unrelated to nectin1 that does not impact melanoma initiation or progression in our system. gRNAs targeting arhgap11a, nectin1a, nectin1b, fak1a, fak1b (Supplementary Table 4) were introduced into the CRISPR MiniCoopR vector23 by restriction cloning using the BseRI and AarI enzymes. A pLKO shRNA lentiviral vector (TRCN0000373345, Sigma-Aldrich) was used to down-regulate NECTIN1 in human melanoma cell lines.
The LentiCRISPR lentiviral vector46 was used to inactivate NECTIN1 in human melanoma cell lines (Supplementary Table 4).
The CD8-IGF1R ORF was cloned into a middle-entry vector, and assembled by multisite Gateway technology47 with either the MiniCoopR destination vector48 for melanocyte-specific expression in zebrafish or the pHAGE vector for doxycycline-inducible expression in human cells. A GFP construct was used as a negative control. The NECTIN1 ORF was obtained from the Harvard Plasmid repository.
CRISPR sequencing
Genomic DNA from human cell lines or zebrafish tumors was extracted with QuickExtract solution (Epibio). DNA libraries were prepared by PCR amplification of the CRISPR loci using the primers described in Supplementary Table 4.
Sequencing was performed by the MGH DNA core and analyzed using the Basepair online tool (https://www.basepairtech.com).
Zebrafish
Zebrafish (Danio rerio) were handled according to our vertebrate animal protocol that has been approved by Boston Children’s Hospital Animal Care Committee and includes detailed experimental procedures for all in vivo experiments described in this paper. Zebrafish of the casper (mitfa−/−, roy−/−) and Tg(mitfa:BRAFV600E), tp53−/−, mitfa−/− strains48,49 were housed at a density of approximately 10 fish per liter of water at 28.5 °C and bred by pairwise crossing. Embryos were collected for microinjection.
Zebrafish embryo microinjection
Zebrafish of casper (mitfa−/−, roy−/−) and Tg(mitfa:BRAFV600E), tp53−/−, mitfa−/− strains48,49 were bred and embryos were collected for microinjection. The Tol2 transgenesis technology enables the stable integration of expression vectors into the fish genome45. Embryos were injected at the one-cell stage with CRISPR MiniCoopR vectors23 targeting nectin1, fak or a control gene and/or MiniCoopR vectors expressing CD8-IGF1R or GFP. 25 pg DNA constructs and 25 pg Tol2 mRNA were injected into one-cell-stage embryos. After microinjection, embryos were raised in E3 medium at 28.5 °C. Embryos were sorted for melanocyte rescue at 96 h after fertilization and raised to adulthood (15–25 zebrafish per 3.5-liter tank). Tumor formation was monitored between weeks 10 and 22 after injection. Pigmented tumors were collected randomly from adult male and female fish (between 3 and 6 months of age) for transplantation. We did not observe any differences in tumor onset between males and females.
In vivo tumor spreading assay
Transplantation of primary zebrafish melanoma cells into irradiated adult casper recipients was performed as previously described25. Specifically, 3- to 6-month-old zebrafish were irradiated at 30 Gy split over 2 days, and 300,000 melanoma cells in suspension in 3 μl PBS were injected subcutaneously into their dorsal cavity using a 10 μl Hamilton syringe with 32-gauge bevel-tipped needle. Each primary tumor was transplanted into 5–10 secondary recipients that were randomly allocated to each genetic condition. No animals were excluded from data analysis. The proportion of females was lower than that of males (approximately 20%) but identical in all experimental groups. We did not observe any influence of sex in our transplantation experiments. Recipient fish were photographed using a Nikon D3100 camera with an AF-S Micro NIKKOR 60 mm lens every 7 days after transplantation for 4 weeks.
AFM
Atomic force microscopy (AFM) was performed on whole primary melanomas freshly excised from adult zebrafish. Dimension 3100 (Bruker Nano/Veeco) AFM with Nanoscope V controller and nPoint NPXY200Z25 (nPoint) close-loop scanner (200 μm × 200 μm × 25 μm, XYZ) were used in the present study. A standard cantilever holder for operation in liquids was used. The raw indentation data were collected with the vertical ramp size of 15 µm. To minimize viscoelastic effects, the indentation data were recorded at a frequency of 0.75 Hz. The AFM software used was version 7.30 R1Sr3. The data were also pre-processed to align the baseline with SPIP version 6.7.2 (Image Metrology A/S).
The AFM indentation curves were collected using the force-volume mode. 16 × 16 pixels (typically within 80 × 80 µm2 area) images were recorded. Standard V-shaped arrow 200 µm AFM tipless cantilevers NP-O10 (Bruker) were used throughout the study. The cantilever spring constant was measured using the thermal tuning method before gluing the spherical probe. Typical spring constant was ~0.06 N m−1. Silica balls 5 µm in diameter (Bangs Labs) were glued to the cantilevers as previously described50. The radius of the probe was measured by imaging the inverse grid (TGT1 by NT-NGT, Russia). The use of a rather dull AFM probe is important to attain self-consistency of the models51. In each experiment, the AFM cantilever sensitivity was calibrated against a rigid substrate, a small piece of a silicon wafer or coverslip immersed in PBS solution.
In the force-volume mode, one collects both cell sample topography and the indentation data52,53. Furthermore, the sample topography can be corrected for the inevitable deformation of the sample by the action of the AFM probe51. We processed the indentation curves collected over the top points on the sample until we reached the inclination angle of 10–15 degrees. The Young’s modulus of the sample surface was found using the Hertz model built in the SPIP software (Image Metrology A/S). The specific range of fitting was identified by the requirement of independence of the modulus of the indentation depth54.
Primary zebrafish cell cultures
Zebrafish cell lines were established as previously described from BRAFV600E, tp53−/−, nectin1+/+ or nectin1−/− tumors25. Whole tumors were dissociated with liberase, plated in 48-well plates coated with fibronectin and cultured in the presence of zebrafish media (DMEM:F12 supplemented with 15% FBS, 10% zebrafish embryo extract, HEPES, sodium pyruvate, glutamine, insulin, holotransferrin, selenous acid, endothelin 1, chemically defined lipids, non-essential amino acids, penicillin/streptomycin and primocin). After three or four passages (or when cells expanded from a six-well plate to a T25 flask), cells were gradually transitioned to DMEM + 10% FBS. All cultures were regularly checked for the presence of mycoplasma.
Transfection and viral transduction
Lentiviral particles were produced by co-transfection of 293 T cells with pLKO or pHAGE vectors and packaging plasmids pVSV-G and psPAX2 using FuGENE HD (Promega). Viral particles were harvested 48 and 72 h after transfection, concentrated by overnight PEG precipitation, resuspended in PBS and stored at −80 °C. Human melanoma cell lines were overlaid with viral particles diluted in medium supplemented with 5 μg ml−1 polybrene (Sigma-Aldrich) for 12 h at 37 °C. Then, 48 h after transduction, infected cells were selected with 500 μg ml−1 G418 (Gibco) or with 1 μg ml−1 puromycin (Gibco) and maintained under selection by replacing the antibiotics every 48 h.
A375 cells were transfected with pLentiCRISPR vectors using FuGENE HD (Promega). Stable knockout lines were established from single-cell colonies.
siRNA against FAK (SASI_Hs01_00035697), SRC (Hs01_00112907), ITGB1 (SASI_Hs02_00333437), ITGB2 (SASI_Hs01_00245442), ITGB3 (SASI_Hs01_00174219), ITGB4 (SASI_Hs01_00230929), ITGB5 (SASI_Hs02_00333438) were purchased from Millipore and transfected using the N-TER Nanoparticle siRNA Transfection System according to the manufacturer’s instructions.
Adhesion and migration assays
Spheroid formation capacity was evaluated by plating 100–10,000 human melanoma cells in the wells of a Round Bottom Ultra Low Attachment 96-well Microplate (Corning) in the presence or absence of FBS. Spheres were imaged after 4–8 days using a Nikon SMZ18 stereomicroscope or a Nikon C2-si confocal microscope. Circularity was measured using ImageJ and multiplied by 100 for ease of representation. A perfect circle would have a circularity of 100 (Fig. 3b).
For the adhesion assay, wells of a 12-well plate were coated with Collagen-I (3440, R&D Systems), fibronectin (F0895, Sigma-Aldrich), laminin (LN521-02, Biolamina), vitronectin (5051, Sigma-Aldrich) at 37 °C for 3 h and then at 4 °C overnight. Human melanoma cells were transduced with a lentiviral vector stably expressing GFP, harvested and resuspended in the absence of FBS. Then, 25,000 cells were seeded in ECM protein-coated well. After 15-, 30- or 45-min incubations, wells were rinsed with PBS and immediately imaged using a Nikon SMZ18 microscope equipped with a DS-Ri2 camera. The number of attached cells was quantified with ImageJ.
The Transwell assay was performed with 6.5 mm Transwell with 8.0 µm Pore Polycarbonate Membrane Insert (Corning) according to the manufacturer’s instructions. For invasion assays, Matrigel (2515251, Corning) containing the cells was added to the transwells. 40,000 cells were seeded in the absence of FBS for 12 h. The membranes were then transferred to a well containing media with FBS. Cells were allowed to migrate for 6 h (A375, CJM), 10 h (RPMI7951, G361) or 24 h (SKMEL2, COLO679), fixed, stained with DAPI and imaged using a Nikon SMZ18 microscope equipped with a DS-Ri2 camera. The number of cells that have migrated through the membrane was quantified with ImageJ.
For colony-forming assays in semi-solid media, 1,000 A375 human melanoma cells were seeded in 250 μl 50% Matrigel (Corning) or 1x Geltrex Collagen-I matrix (R&D Systems) on a layer of 250 μl 100% Matrigel or 10x Geltrex Collagen-I matrix in 24-well plates. Then, 1 ml culture media was added to wells containing Matrigel. Spread was measured with ImageJ as the number of individual particles detected in each colony (Fig. 3d).
Migration inhibitors
The following compounds were tested for toxicity with the CellTiterGlo assay after a 4-day treatment in A375 melanoma cells and used at the maximal nontoxic dose in the transwell assay: Integrin inhibitor (Echistatin, 100 nM, R&D Systems), FAK inhibitor (PF562271 besylate, 500 nM, Cayman Chemicals), SRC inhibitor (Dasatinib, 10 nM, LC Laboratories), RAC1/CDC42 inhibitor (MBQ-167, 50 nM, Selleckchem), Wnt inhibitor (XAV939, 1 μM, Cayman Chemicals), JNK inhibitor (SP600125, 1 μM, LC Laboratories), ROCK inhibitor (RKI-1447, 500 nM, Cayman Chemicals), PKC inhibitor (Staurosporine, 1 nM, LC Laboratories), AMPKa inhibitor (Dorsomorphin, 1 μM, LC Laboratories), integrin inhibitor (SB273005, 100 nM, Selleckchem), FAK inhibitor (GSK2256098, 1 μM, Cayman Chemicals) and SRC inhibitor (Bosutinib, 100 nM, LC Laboratories). Echistatin was resuspended in water. All other compounds were dissolved in DMSO.
In addition, CD49f (Integrin alpha 6) monoclonal antibody (GoH3) (14-0495-85, eBioscience) was used at 40 μg ml−1 to block integrin α6β4.
Screen for growth factors and hormones
The following factors were tested for their ability to prevent the formation of adherens junctions between melanoma cells upon serum withdrawal, as assessed by immunofluorescence for α-catenin: EGF (1 ng ml−1, StemCell Technologies), bFGF (10 ng ml−1, StemCell Technologies), IGF1 (100 ng ml−1, StemCell Technologies), SCF (25 ng ml−1, StemCell Technologies), PDGF-BB (100 ng ml−1, Gibco), TGFβ (10 ng ml−1, StemCell Technologies), insulin (10 μg ml−1, R&D Systems), IFN-α (100 pg mll−1, StemCell Technologies), NTN1 (1 μg ml−1, R&D Systems), retinoic acid (100 ng ml−1, Sigma-Aldrich), β-estradiol (100 pg ml−1, Sigma-Aldrich) and progesterone (20 ng ml−1, Sigma-Aldrich).
Immunofluorescence
Cells were grown on glass coverslips for 2 days, fixed with 4% PFA for 20 min, permeabilized with PBS 0.2% Triton for 30 min at 4 °C, blocked with 1% BSA in PBS, 0.05% Triton for 1 h at room temperature, hybridized with primary antibody in blocking solution for 2 h at room temperature, hybridized with fluorescently-labeled secondary antibody in blocking solution for 2 h at room temperature in the dark, rinsed in PBS and mounted on glass slides using VECTASHIELD Antifade Mounting Medium with DAPI (Vector Laboratories). Primary antibodies: α-E-Catenin (1:200, #3236, Cell Signaling Technology), N-Cadherin (1:200, #13116, Cell Signaling Technology), α-Actinin (1:100, #69758, Cell Signaling Technology), fibronectin (1:400, #26836, Cell Signaling Technology) and laminin (1:100, L9393, Sigma-Aldrich). Secondary antibodies used were Alexa Fluor 488 goat anti-rabbit (1:400, A11008, Life Technologies) and Alexa Fluor 568 goat anti-mouse (1:400, A11004, Life Technologies). F-actin was stained using Alexa Fluor 568 Phalloidin (1:2000, A12380, Life Technologies). Cells were imaged using a Nikon C2-si confocal microscope.
For immunofluorescence on human melanoma tissue or zebrafish tissue, sections were deparaffinized through a xylene/ethanol gradient and rehydrated. Heat-induced antigen retrieval was completed using Antigen Unmasking Solution, Citric Acid Based (H-3300, Vector) in a microwave for 20 min. Sections were blocked with 5% BSA in PBS for 30 min and incubated with α-catenin (1:200, 13-9700, Invitrogen), phospho-IGF1R beta (Tyr1161) (1:200, PA5-37601, Invitrogen), E-cadherin (4A2) (1:200, #14472, Cell Signaling Technology), NECTIN1 (1:200, HPA026846, Sigma-Aldrich), laminin (1:100, L9393, Sigma-Aldrich) and FNDC3A (1:200, PA5-109309, Invitrogen) primary antibodies at 4 °C overnight. Sections were then incubated with Alexa Fluor Plus 488 goat anti-Mouse IgG (1:400, A32723, Life Technologies) and Alexa Fluor 568 goat anti-rabbit IgG (1:400, A11036, Life Technologies) secondary antibodies for 2 h and mounted using VECTASHIELD Antifade Mounting Medium with DAPI (Vector Laboratories).
Protein analysis
For western blot, cells were lyzed with RIPA lysis buffer containing protease inhibitors (cOmplete, Roche) and phosphatase inhibitors (PhosSTOP, Roche). Lysates were incubated for 20 min on ice and spun down for 10 min at 14,000 r.p.m. at 4 °C. Protein concentrations were normalized using the DC protein assay (BioRad). Samples were denatured by adding Laemmli sample buffer (BioRad) with 5% β-mercaptoethanol (Sigma-Aldrich) and boiled at 95 °C for 5 min before loading. Proteins were separated on a 10% mini-PROTEAN TGX (BioRad) precast gel, and transferred onto a PVDF membrane using the iBlot2 system (Thermo Fisher Scientific). Primary antibodies: NECTIN1 (1:1,000, ab66985, Abcam), phospho-FAK (1:1,000, #3283, Cell Signaling Technology), FAK (1:1,000, #71433, Cell Signaling Technology), phospho-SRC (1:1,000, #2101, Cell Signaling Technology), SRC (1:1,000, #2109, Cell Signaling Technology), integrin β1 (1:1,000, #9699, Cell Signaling Technology), integrin β2 (1:1,000, #73663, Cell Signaling Technology), Integrin β3 (1:1,000, #13166, Cell Signaling Technology), integrin β4 (1:1,000, #14803, Cell Signaling Technology), integrin β5 (1:1,000, #3629, Cell Signaling Technology), GAPDH (1:2,000, ab8245, Abcam) and β-ACTIN (1:5,000, A2228, Sigma-Aldrich). Secondary antibodies used were HRP anti-mouse and HRP anti-rabbit (1:2,000, Cell Signaling Technology). Membranes were developed using Pierce ECL Western Blotting Substrate (Thermo Fisher Scientific) or ECL Prime Western Blotting Detection Reagent (Amersham). Activity of intracellular signaling cascades was measured using a Proteome Profiler Human Phospho-Kinase Array Kit (R&D Systems) following the manufacturer’s instructions. Signal from the immunoblot was quantified using ImageJ.
Purification and analysis of cell-surface proteins
Cell-surface proteins were purified using the EZ-Link Sulfo-NHS-SS-Biotin system (Thermo Fisher Scientific). Cells were rinsed with PBS ( + CaCl2/MgCl2) and incubated in a 0.5 mg ml−1 biotin solution for 1 h on ice. The free biotin was quenched by three 5-min washes with a 50 mM glycine solution. Cells were rinsed with PBS and lyzed in an immunoprecipitation buffer (Cell Signaling Technology) containing protease inhibitors. After centrifugation of the lysate, Neutravidin beads (Thermo Fisher Scientific) were added 1:2 to the supernatant. The beads/lysate mixture was incubated at 4 °C for 3 h under constant rotation. Beads were washed once with the immunoprecipitation buffer and then three times for 5 min with a 50 mM Tris-HCl solution. After on-bead digestion with trypsin, purified peptides were labeled and analyzed by tandem mass spectrometry using a 180-min MS3 method on an Orbitrap Fusion Lumos mass spectrometer.
Gene expression analyses
Read length was 150 bp pair ended. Quality control of RNA-sequencing datasets was performed by FastQC and Cutadapt to remove adaptor sequences and low-quality regions. The high-quality reads were aligned to UCSC build hg19 of human genome using Tophat 2.0.11 without novel splicing form calls. Transcript abundance and differential expression were calculated with Cufflinks 2.2.1 (ref. 55). Raw counts were not normalized but FPKM values were used to normalize and quantify each transcript. Genes were filtered by log fold change and q-value. Differentially regulated pathways were analyzed by Gene Ontology56,57 and gene set enrichment analysis58,59.
Statistics
Statistical tests used in this study are indicated in figure legends. Comparison of Kaplan–Meier survival curves was performed by a log-rank (Mantel–Cox) test. Statistical differences in cell culture experiments were estimated using paired two-tailed t-test. For other experiments, a two-tailed t-test was used. Data distribution was assumed to be normal, but this was not formally tested. An F test was used to compare variances. For gene expression analyses, statistical methods are describes in the original Gene Ontology56,57 and gene set enrichment analysis58,59 papers. For each statistical test, the P value is reported on the figure. Significance was defined as P value < 0.05. Dispersion of the data and precision measures are specified in figure legends. Generally, graphs presented in this study show the mean ± s.d. The number of replicates indicated on figures or in figure legends refers to biological replicates. For cell line work, no statistical methods were used to predetermine sample size, but our sample sizes are similar to those reported in previous publications. Experiments were repeated four times to ensure robustness, unless statistical significance could be reached with three replicates only. For zebrafish transplantation experiments, a target sample size of 16 primary tumors per arm was calculated such that twofold differences in tumor spreading could be observed with a power of 80% and with statistical significance (5% type I error) using two-sided Student’s t-test. This sample size was never attained because statistical significance was reached with smaller samples due to the larger size of the observed effects. For human melanoma sections, sample size was limited by availability. Analyses of the screens for growth factors and hormones and migration inhibitors were blinded. Data collection was also performed blinded in the case of transplantation experiments in zebrafish. Blinding was only lifted at the time of data analysis.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The TCGA dataset for cutaneous melanoma used in this study is available at https://portal.gdc.cancer.gov/projects/TCGA-SKCM.
Copy-number analyses were performed with GISTIC 2.0 through the interface of the Broad Institute (https://portals.broadinstitute.org/tcga/home) using 2014-04-28 stddata release.
The RNA-sequencing data of the consequences of NECTIN1 loss and the effect of serum depletion on A375 human melanoma cells generated during this study have been deposited in the Gene Expression Omnibus under the accession code GSE174150 and are publicly available.
The tandem mass spectrometry data of the cell-surface proteins of NECTIN1-deficient and NECTIN1-proficient A375 human melanoma cells upon serum depletion generated during this study have been deposited in MassIVE (Center for Computational Mass Spectrometry, UCSD) under the accession code MSV000090170 and are publicly available. Source data are provided with this paper.
Code availability
All data processing and analysis were performed by existing software packages, which have been included in the relevant Methods section. No custom code or software was used for any aspect of data processing or analysis.
References
Steeg, P. S. Targeting metastasis. Nat. Rev. Cancer 16, 201–218 (2016).
Schadendorf, D. et al. Melanoma. Lancet 392, 971–984 (2018).
Kim, M. et al. Comparative oncogenomics identifies NEDD9 as a melanoma metastasis gene. Cell 125, 1269–1281 (2006).
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Turajlic, S. et al. Tracking cancer evolution reveals constrained routes to metastases: TRACERx Renal. Cell 173, 581–594 (2018).
Reiter, J. G. et al. Minimal functional driver gene heterogeneity among untreated metastases. Science 361, 1033–1037 (2018).
Priestley, P. et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 575, 210–216 (2019).
McAllister, S. S. & Weinberg, R. A. The tumour-induced systemic environment as a critical regulator of cancer progression and metastasis. Nat. Cell Biol. 16, 717–727 (2014).
Vanharanta, S. & Massagué, J. Origins of metastatic traits. Cancer Cell 24, 410–421 (2013).
Cavallaro, U. & Christofori, G. Cell adhesion and signalling by cadherins and Ig-CAMs in cancer. Nat. Rev. Cancer 4, 118–132 (2004).
Peri, A. K., Wilgenbus, P., Dahl, U., Semb, H. & Christofori, G. A causal role for E-cadherin in the transition from adenoma to carcinoma. Nature 392, 190–193 (1998).
Nieto, M. A., Huang, R. Y. Y. J., Jackson, R. A. A. & Thiery, J. P. P. EMT: 2016. Cell 166, 21–45 (2016).
Dongre, A. & Weinberg, R. A. New insights into the mechanisms of epithelial–mesenchymal transition and implications for cancer. Nat. Rev. Mol. Cell Biol. 20, 69–84 (2019).
Mort, R. L., Jackson, I. J. & Elizabeth Patton, E. The melanocyte lineage in development and disease. Development 142, 620–632 (2015).
Hsu, M. Y. et al. E-cadherin expression in melanoma cells restores keratinocyte-mediated growth control and down-regulates expression of invasion-related adhesion receptors. Am. J. Pathol. 156, 1515–1525 (2000).
Hsu, M. Y. Shifts in cadherin profiles between human normal melanocytes and melanomas. J. Investig. Dermatol. Symp. Proc. 1, 188–194 (1996).
Takahashi, K. et al. Nectin/PRR: An immunoglobulin-like cell adhesion molecule recruited to cadherin-based adherens junctions through interaction with afadin, a PDZ domain-containing protein. J. Cell Biol. 145, 539–545 (1999).
Satoh-Horikawa, K. et al. Nectin-3, a new member of immunoglobulin-like cell adhesion molecules that shows homophilic and heterophilic cell-cell adhesion activities. J. Biol. Chem. 275, 10291–10299 (2000).
Tachibana, K. et al. Two cell adhesion molecules, nectin and cadherin, interact through their cytoplasmic domain-associated proteins. J. Cell Biol. 150, 1161–1176 (2000).
Beroukhim, R. et al. Assessing the significance of chromosomal aberrations in cancer: Methodology and application to glioma. Proc. Natl Acad. Sci. USA 104, 20007–20012 (2007).
Akbani, R. et al. Genomic classification of cutaneous melanoma. Cell 161, 1681–1696 (2015).
Ablain, J., Durand, E. M., Yang, S., Zhou, Y. & Zon, L. I. A CRISPR/Cas9 vector system for tissue-specific gene disruption in zebrafish. Dev. Cell 32, 756–764 (2015).
Ablain, J. et al. Human tumor genomics and zebrafish modeling identify SPRED1 loss as a driver of mucosal melanoma. Science 362, 1055–1060 (2018).
Fazio, M. et al. Satb2 induction of a neural crest mesenchyme-like program drives melanoma invasion and drug resistance. Elife 10, e64370 (2021).
Heilmann, S. et al. A quantitative system for studying metastasis using transparent zebrafish. Cancer Res. 75, 4272–4282 (2015).
Tsoi, J. et al. Multi-stage differentiation defines melanoma subtypes with differential vulnerability to drug-induced iron-dependent oxidative Stress. Cancer Cell 33, 890–904 (2018).
Sato, T. et al. Regulation of the assembly and adhesion activity of E-cadherin by nectin and afadin for the formation of adherens junctions in Madin-Darby canine kidney cells. J. Biol. Chem. 281, 5288–5299 (2006).
Arozarena, I. & Wellbrock, C. Phenotype plasticity as enabler of melanoma progression and therapy resistance. Nat. Rev. Cancer 19, 377–391 (2019).
Humphries, J. D., Byron, A. & Humphries, M. J. Integrin ligands at a glance. J. Cell Sci. 119, 3901–3903 (2006).
Carboni, J. M. et al. Tumor development by transgenic expression of a constitutively active insulin-like growth factor I receptor. Cancer Res. 65, 3781–3787 (2005).
Satyamoorthy, K. et al. Insulin-like growth factor-1 induces survival and growth of biologically early melanoma cells through both the mitogen-activated protein kinase and β-catenin pathways. Cancer Res. 61, 7318–7324 (2001).
Sakamoto, Y. et al. Interaction of integrin αvβ3 with nectin: Implication in cross-talk between cell-matrix and cell-cell junctions. J. Biol. Chem. 281, 19631–19644 (2006).
Lopez, T. & Hanahan, D. Elevated levels of IGF-1 receptor convey invasive and metastatic capability in a mouse model of pancreatic islet tumorigenesis. Cancer Cell 1, 339–353 (2002).
Castaño, Z. et al. Stromal EGF and IGF-I together modulate plasticity of disseminated triple-negative breast tumors. Cancer Discov. 3, 922–935 (2013).
Le Coz, V. et al. IGF-1 contributes to the expansion of melanoma-initiating cells through an epithelial-mesenchymal transition process. Oncotarget 7, 82511–82527 (2016).
Gupta, G. P. & Massagué, J. Cancer metastasis: Building a framework. Cell 127, 679–695 (2006).
White, R. M. et al. DHODH modulates transcriptional elongation in the neural crest and melanoma. Nature 471, 518–522 (2011).
Kaufman, C. K. et al. A zebrafish melanoma model reveals emergence of neural crest identity during melanoma initiation. Science 351, aad2197 (2016).
Suzuki, K. et al. Mutations of PVRL1, encoding a cell-cell adhesion molecule/herpesvirus receptor, in cleft lip/palate-ectodermal dysplasia. Nat. Genet. 25, 427–430 (2000).
Ciarlo, C. et al. A chemical screen in zebrafish embryonic cells establishes that Akt activation is required for neural crest development. Elife 6, e29145 (2017).
Li, G., Satyamoorthy, K. & Herlyn, M. N-cadherin-mediated intercellular interactions promote survival and migration of melanoma cells. Cancer Res. 61, 3819–3825 (2001).
Piskounova, E. et al. Oxidative stress inhibits distant metastasis by human melanoma cells. Nature 527, 186–191 (2015).
Konieczkowski, D. J. et al. A melanoma cell state distinction influences sensitivity to MAPK pathway inhibitors. Cancer Discov. 4, 816–827 (2014).
Ahn, S., Woo, J. W., Lee, K. & Park, S. Y. HER2 status in breast cancer: Changes in guidelines and complicating factors for interpretation. J. Pathol. Transl. Med. 54, 34–44 (2020).
Kawakami, K. et al. A transposon-mediated gene trap approach identifies developmentally regulated genes in zebrafish. Dev. Cell 7, 133–144 (2004).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Hartley, J. L., Temple, G. F. & Brasch, M. A. DNA cloning using in vitro site-specific recombination. Genome Res. 10, 1788–1795 (2000).
Ceol, C. J. et al. The histone methyltransferase SETDB1 is recurrently amplified in melanoma and accelerates its onset. Nature 471, 513–517 (2011).
White, R. M. et al. Transparent adult zebrafish as a tool for in vivo transplantation analysis. Cell Stem Cell 2, 183–189 (2008).
Berdyyeva, T. K., Woodworth, C. D. & Sokolov, I. Human epithelial cells increase their rigidity with ageing in vitro: Direct measurements. Phys. Med. Biol. 50, 81 (2005).
Guz, N., Dokukin, M., Kalaparthi, V. & Sokolov, I. If cell mechanics can be described by elastic modulus: Study of different models and probes used in indentation experiments. Biophys. J. 107, 564–575 (2014).
A-Hassan, E. et al. Relative microelastic mapping of living cells by atomic force microscopy. Biophys. J. 74, 1564–1578 (1998).
Sokolov, I., Zorn, G. & Nichols, J. M. A study of molecular adsorption of a cationic surfactant on complex surfaces with atomic force microscopy. Analyst 141, 1017–1026 (2016).
Dokukin, M. E. & Sokolov, I. On the measurements of rigidity modulus of soft materials in nanoindentation experiments at small depth. Macromolecules 45, 4277–4288 (2012).
Trapnell, C. et al. Transcript assembly and abundance estimation from RNA-Seq reveals thousands of new transcripts and switching among isoforms. Nat. Biotechnol. 28, 511–515 (2011).
Ashburner, M. et al. Gene ontology: Tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Carbon, S. et al. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank A. Weng and S. Gusscott for sharing their CD8-IGF1R plasmid, C. Yuan for technical support with immunohistochemistry, S. Rowbotham for comments on the manuscript, N. Osmani for advice regarding integrin signaling and S. Avagyan for critical reading of the manuscript, discussions on the data and logistical support between the Lyon and Boston laboratories. This work was supported by grants from the National Institutes of Health/National Cancer Institute (R01 CA103846 and P01 CA163222 to L.I.Z. and K99 CA201465 to J.A.), the Sloan Kettering Institute for Cancer Research (BD527651 to L.I.Z.), the Melanoma Research Alliance, the National Science Foundation (CMMI 1435655 and CMMI 1937373 to I.S.), the Centre National de la Recherche Scientifique (ATIP Avenir 2019 to J.A.), the Institut National de la Santé et de la Recherche Médicale, the Ligue contre le Cancer, the Fondation ARC pour la recherche sur le cancer and the Fondation Tourre. L.I.Z. is a Howard Hughes Medical Institute Investigator.
Author information
Authors and Affiliations
Contributions
J.A. designed the study, performed cell culture and zebrafish experiments, analyzed results, discussed findings and wrote the manuscript, A.A. performed cell culture experiments and analyzed results, H.R. helped with cell culture and zebrafish experiments, M.P. and S.Y. performed RNA-sequencing analyses, S.A. participated in cell culture experiments, M.D. performed initial zebrafish transplantation experiments, S.X. and C.G.L. performed and analyzed immunohistochemistry experiments and M.E.D. and I.S. performed and analyzed atomic force microscopy experiments. L.I.Z. designed the study, discussed findings and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
L.I.Z. is a founder and stock holder of Fate Therapeutics, CAMP4 Therapeutics, Scholar Rock, and Amagma, outside of the present work. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Rodney Stewart, Victoria Sanz-Moreno, and Inna Smalley for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genetics of NECTIN1 in human melanoma.
(a) List of the 18 most significant chromosomal deletions found in the Cancer Genome Atlas (TCGA) cohort of human cutaneous melanomas (363 samples). (b) Diagram representing the frequency of chromosomal deletions in the TCGA cohort of human cutaneous melanomas (363 samples). Frequent loss of CDKN2A on chromosome 9, and PTEN on chromosome 10 are highlighted. NECTIN1 was identified by GISTIC analysis in a recurrent focal deletion on chromosome 11. (c) Linear copy number in 202 human melanoma samples exhibiting deep or shallow deletions of the NECTIN1 locus. (d) Sequences of shallow deletions affecting NECTIN1 in three examples of human melanoma samples. In TCGA-D9-A148 (Patient 1) and TCGA-FS-A1ZQ (Patient 2), the deletion encompasses the entire NECTIN1 gene, while in TCGA-FW-A3TU (Patient 3), the deletion starts in exon 2. (e) Driver lesions in oncogenes and tumor-suppressor genes occurring with NECTIN1 alterations in the TCGA cohort of human cutaneous melanomas (363 samples).
Extended Data Fig. 2 NECTIN1 expression in human melanoma.
(a) Dot plot representing NECTIN1 mRNA expression and linear copy number in the TCGA cohort of human cutaneous melanomas (363 samples). Spearman correlation: r = 0.38, p = 3.8E-14. Four outliers with high NECTIN1 expression were removed from the graph but retained in the analysis. (b) Box-and-whisker plot representing NECTIN1 mRNA expression depending on NECTIN1 copy-number status (363 samples). Min-max, 5 and 95 percentiles, median. Brown-Forsythe ANOVA. Unpaired two-tailed Welch’s t-test: deep deletion vs diploid: p = 0.0013, shallow deletion vs diploid: p = 0.0006. Four outliers with high NECTIN1 expression were removed from the graph but retained in the analysis. (c) Representative histology and scoring system used for the evaluation of NECTIN1 expression by immunohistochemistry, related to Fig. 1d. 0 = Negative (<1% tumor cells immunoreactive); 1= weak and incomplete staining in < 10% of cells, 2 = weak/moderate heterogeneous staining in > 10% of cells, or 3 = strong and complete homogenous staining in > 10% of cells. Scale bar: 50 μm. Representative of 3 independent tissue-microarrays. (d) Distribution of NECTIN1 staining level by immunohistochemistry in 253 tissue sections of human primary melanomas or metastases. Correlation was measured by Chi-square test. (e) Dot plot representing NECTIN1 and CDH1 mRNA expression in the TCGA cohort of human cutaneous melanomas (363 samples). Spearman correlation: r = −0.08, p = 0.107. Four outliers with high NECTIN1 expression were removed from the graph but retained in the analysis. (f) Immunofluorescence analysis of E-cadherin (green) and NECTIN1 (red) on tissue sections of 4 different human melanomas exhibiting different staining patterns: from left to right and top to bottom: double positive, single E-cadherin-positive, single NECTIN1-positive, double negative. Scale bar: 10 μm. BV, blood vessel. Representative examples of the cases described in g. (g) Distribution of NECTIN1 and E-cadherin positivity by immunofluorescence in 20 sections of human melanoma. Correlation was measured by Chi-square test.
Extended Data Fig. 3 NECTIN1 inactivation in zebrafish and human melanoma.
(a) Box-and-whisker plot representing nectin1a and nectin1b expression levels in primary zebrafish melanomas. Min-max, 1st and 4th quartiles, median, n = 10. Two-tailed t-test. (b) Type and proportion of nectin1a and nectin1b mutant alleles in 3 primary zebrafish melanomas, their corresponding allografts in casper zebrafish, and matched disseminated patches. D: deletion, I: insertion, numbers denote affected base pairs. Unlabeled slice represents all other minor alleles. (c) Elasticity (Young’s modulus) of primary zebrafish melanomas, as measured by atomic force microscopy. Mean ± SD of 4184 and 3134 measurements performed on 20 and 26 areas of 3 control-knockout and 3 nectin1-knockout tumors, respectively. Two-tailed Kolmogorov-Smirnov test. (d) Evolution of nectin1b variant allele fraction in 2 zebrafish melanoma cell lines over 7 passages in culture in the presence or absence of zebrafish media. wt: wild-type; 5D: 5 base-pair deletion. (e) Images of cultures of the supernatant of a nectin1-knockout cell line subjected to zebrafish media withdrawal. Scale bar: 50 μm. Representative of 3 independent experiments. (f) Spheroids formed by control or nectin1-knockout (KO) zebrafish melanoma cells after 4 days in low-attachment conditions in the absence of zebrafish media. Scale bar: 200 μm. Representative of 3 independent experiments. (g) Western blot analysis of NECTIN1 in A375 cells stably expressing a control shRNA (C) or an shRNA against NECTIN1 (Sh), and either GFP or a NECTIN1 construct mutated in the shRNA target site (NECTIN1*). Representative of 3 independent experiments. (h) Sequences of the NECTIN1 CRISPR loci in wild-type (WT) A375 cells and 3 NECTIN1-knockout (KO) cell lines derived from A375. Blue: PAM sequence; red: insertion/deletion; yellow: mutation. (i) Western blot analysis of NECTIN1 in 3 NECTIN1-wild-type and 3 NECTIN1-knockout (KO) cell lines derived from A375. Representative example of 2 independent experiments. (j) NECTIN1 mRNA expression levels (FPKM) in the cells described in i. Two-tailed t-test.
Extended Data Fig. 4 Migratory phenotype associated with NECTIN1 inactivation in human melanoma cell lines.
(a) Structures formed by 5,000 A375 cells stably expressing a control shRNA (shCTRL) or an shRNA against NECTIN1 (shNECTIN1) and stained with fluorescent dyes, after 4 days in low-attachment conditions in the absence of serum. Scale bar: 300 μm. Representative of 3 independent experiments. (b) Spheroids formed by 20,000 cells of 3 NECTIN1-wild-type (WT) and 3 NECTIN1-knockout (KO) cell lines derived from A375 cells, after 4 days in low-attachment conditions in the absence of serum. Scale bar: 500 μm. Representative of 5 independent experiments. (c) Analysis of the images shown in b. (d) Circularity of the colonies described in b, as measured with ImageJ. Mean ± SD of 5 independent colonies per condition. Two-tailed t-test. (e) Migration of the cells described in b in a transwell assay after 12 hours of serum starvation. Cells were allowed to migrate for 6 hours. Mean ± SD of 3 independent experiments. Two-tailed t-test. (f) Migration of the cells described in a stably expressing either GFP or an shRNA-resistant NECTIN1 gene (NECTIN1*) in a transwell assay after 12 hours of serum starvation. Cells were allowed to migrate for 6 hours. Mean ± SD of 3 independent experiments. Paired two-tailed t-test. (g) Diagram representing the position of 6 human melanoma cell lines on the spectrum of differentiation states from undifferentiated, migratory, AXLhigh/MITFlow to melanocytic, proliferative, AXLlow/MITFhigh. NC-like, neural crest-like. (h) Western Blot analysis of NECTIN1 in 6 human melanoma cell lines. Representative example of 3 independent experiments. (i) Migration of 6 human melanoma cell lines stably expressing a control shRNA (shCTRL) or an shRNA against NECTIN1 (shNECTIN1) in a transwell assay after 12 hours of serum starvation. Cells were allowed to migrate for 10 hours and were counted in a 2.35 ×2.35 mm field (a.u.). Mean ± SD of 4 independent experiments. Paired two-tailed t-test.
Extended Data Fig. 5 Changes in cell adhesion and proliferation upon NECTIN1 loss in human melanoma cell lines.
(a) Immunofluorescence analysis of α-catenin (green) and F-actin (red) in A375 human melanoma cells stably expressing a control shRNA in the absence of serum for 12 hours. Scale bar: 10 μm. Representative images of 5 independent experiments. (b) Immunofluorescence analysis of N-cadherin (green) and α-actinin (red) in A375 human melanoma cells stably expressing a control shRNA (shCTRL) or an shRNA directed against NECTIN1 (shNECTIN1) and cultured in the presence or absence of serum (FBS) for 12 hours. Scale bar: 10 μm. Representative images of 5 independent experiments. (c) Quantification of the number of stress fibers per cell based on α-actinin staining on immunofluorescence images as shown in b. Mean ± SD of n = 29, 26, 24, 20 cells for shCTRL+FBS, shCTRL-FBS, shNECTIN1+FBS, and shNECTIN1-FBS, respectively. Paired two-tailed t-test. (d) Migration of NECTIN1-deficient cells depending on cell density in a transwell assay after 12 hours of serum starvation. Data was pooled from 3 NECTIN1-knockout (KO) cell lines relative to 3 NECTIN1-wild-type (WT) cell lines derived from A375 cells. Cells were seeded at high density (40,000 cells) or low density (4,000 cells) and allowed to migrate for 6 hours. Mean ± SD of 3 independent experiments. Paired two-tailed t-test. (e) Proliferation of the cell lines described in d, measured as the cell count 4 days after seeding relative to the average cell count in wild-type lines. Mean ± SD of 4 independent experiments. Two-tailed t-test. (f) Number of colonies formed in matrigel by the cells described in e after 14 days in culture. Mean ± SD of 3 independent experiments. Two-tailed t-test.
Extended Data Fig. 6 Adhesion and motility program expressed by NECTIN1-deficient cells.
(a) Most represented categories of biological processes in a Gene Ontology (GO) analysis of the set of genes more significantly differentially expressed between NECTIN1-knockout and NECTIN1-wild-type A375 human melanoma cells in response to 12 hours of serum starvation. The GO terms in bold correspond to adhesion and migration signatures. (b-c) Enrichment plots of Integrin and Focal adhesion gene signatures found by Gene Set Enrichment Analysis (GSEA) among the most significantly enriched pathways in NECTIN1-knockout cells compared to NECTIN1-wild-type cells in the A375 human melanoma line under serum starvation. FDR p-values were calculated by GSEA using a permutation test. (d) Migration of A375 human melanoma cells stably expressing an shRNA directed against NECTIN1 (shNECTIN1) relative to cells expressing a control shRNA (shCTRL) in transwells coated with the indicated Extracellular Matrix (ECM) proteins and after 12 hours of serum starvation. Cells were allowed to migrate for 6 hours. Mean ± SD of 4 independent experiments. Paired two-tailed t-test. (e) Expression levels of various ECM genes in 3 clones of A375 human melanoma cells, as measured by RNA-sequencing. (f) Immunofluorescence analysis of fibronectin or laminin (green) in A375 cells cultured in the absence of serum (FBS) for 12 hours. Scale bar: 10 μm. Representative of 3 independent experiments. (g-h) Immunofluorescence analysis of fibronectin (g) and laminin (h) in adult casper zebrafish transplanted with primary zebrafish melanoma cells. Scale bar: 20 μm. S: skin; D: dorsal cavity; M: muscle. Representative of 8 independent zebrafish tissue sections.
Extended Data Fig. 7 Implication of a FAK/SRC axis in the migration of NECTIN1-deficient melanoma cells.
(a) Phospho-kinase array immunoblot of A375 cells stably expressing a control shRNA (shCTRL) or an shRNA against NECTIN1 (shNECTIN1) upon serum depletion. Boxes indicate phospho-FAK. Representative of 4 independent experiments. (b) Most differentially activated proteins between NECTIN1-proficient and deficient cells, as measured by phospho-kinase array immunoblot. Mean ± SD of 4 independent experiments. Paired two-tailed t-test. (c) Relative migration of the cells described in a in a transwell assay after 12 hours of serum starvation and upon treatment with a panel of inhibitors. Cells were allowed to migrate for 6 hours. Mean ± SD of 3 independent experiments. (d) Relative migration of cells treated with a FAK inhibitor (FAKi, GSK2256098) or a SRC inhibitor (SRCi, bosutinib) as described in c. Mean ± SD of 5 independent experiments. Paired two-tailed t-test. (e) Western blot analysis of FAK and SRC in A375 cells transfected with a control siRNA (siCTRL) or siRNAs targeting FAK (siFAK) or SRC (siSRC). Representative example of 4 independent experiments. (f) Relative migration of the cells described in e in the conditions described in c. Mean ± SD of 4 independent experiments. Paired two-tailed t-test. (g) Invasion through matrigel of cells treated with a FAK inhibitor (FAKi, PF562271) or a SRC inhibitor (SRCi, dasatinib) as described in c. Cells were allowed to invade for 10 hours. Mean ± SD of 4 independent experiments. Paired two-tailed t-test. (h) Western blot analysis of FAK and SRC phosphorylation in the cells described in a serum-starved for 12 hours and treated with an integrin inhibitor (ITGi, echistatin), a FAK inhibitor (FAKi, PF562271) or a SRC inhibitor (SRCi, dasatinib) for 6 hours. C: shCTRL; Sh: shNECTIN1; veh: vehicle-treated. Representative example of 4 independent experiments. Signal intensity was normalized to GAPDH. Mean ± SD. Paired two-tailed t-test. P-values are shown for comparisons to vehicle-treated shNECTIN1.
Extended Data Fig. 8 Impact of FAK signaling in NECTIN1-deficient melanoma in vivo.
(a) Representative images of sections of nectin1-wildype or nectin1-knockout zebrafish melanomas transplanted into adult casper recipients, stained for phospho-fak by immunohistochemistry. Scale bars: 50 and 500 μm. (b) Scoring of overall phospho-fak staining intensity in secondary casper recipients transplanted with nectin1-wildype (n = 19) or nectin1-knockout (n = 28) zebrafish melanomas. Mean ± SD. Two-tailed t-test. (c) Top: section of adult casper recipient transplanted with nectin1-knockout zebrafish melanoma cells, stained for phospho-fak by immunohistochemistry. Orientation from skin to muscle is indicated. Scale bar: 50 μm. Representative example of 10 independent zebrafish tissue sections displaying a gradient of phospho-fak staining. Bottom: histogram representing phospho-fak signal intensity along the black dotted line on the image shown on top. The red dotted line materializes the spatial increase in fak activity. (d) Proportion of nectin1a, nectin1b, fak1a, and fak1b mutant alleles in 9 primary zebrafish melanomas expressing CRISPR vectors targeting these genes. Each section of each bar represents a different mutant allele. (e) Tumor-free survival curves of Tg(mitf:BRAFV600E);tp53−/− zebrafish injected with vectors targeting either nectin1 and a control gene, or nectin1 and fak. Pooled data of 2 independent experiments. Log-rank test. (f) Quantification of overall p-H3 staining intensity in 7 nectin1/ctrl-knockout and 9 nectin1/fak-knockout primary zebrafish melanomas. Mean ± SD. Two-tailed t-test. (g) Western Blot analysis of NECTIN1 and phospho-FAK in 6 human melanoma cell lines. Representative example of 3 independent experiments. (h) Correlation plot of NECTIN1 and phospho-FAK levels in 6 human melanoma cell lines. Average values of 3 independent experiments. Pearson’s correlation: R2 = 0.73, p = 0.03. (i) Western Blot analysis of NECTIN1 and phospho-FAK in 4 human melanoma cell lines treated with vehicle (Veh.) or a FAK inhibitor (FAKi, PF562271) for 12 hours. Representative examples of 3 independent experiments.
Extended Data Fig. 9 Implication of IGF1 in the regulation of adherens junctions between melanoma cells.
(a) Immunofluorescence analysis of α-catenin (green) in A375 human melanoma cells stably expressing a control shRNA (shCTRL) or an shRNA against NECTIN1 (shNECTIN1) and cultured in the presence or absence of serum (FBS), or in the presence of dextran-treated/charcoal-stripped serum (treated FBS) for 12 hours. Scale bar: 10 μm. Representative of 4 independent experiments. (b) Western Blot analysis of FAK and SRC phosphorylation levels in the cells described in a cultured in the presence or absence of serum (+FBS or -FBS), or in the presence of dextran-treated/charcoal-stripped serum (tFBS) for 12 hours. Representative example of 3 independent experiments. (c) Most represented categories of biological processes in a Gene Ontology (GO) analysis of the transcriptional response to serum starvation in A375 human melanoma cells. The right column indicates the number of significant GO terms assigned to each category. (d) Enrichment plot of an IGF1/IGF2 signature found among the most significantly enriched pathways in A375 cells cultured in the presence vs absence of serum by GSEA. The FDR p-value was calculated by GSEA using a permutation test. (e) List of secreted factors tested for their ability to prevent the formation of adherens junctions between melanoma cells upon serum withdrawal. (f) Immunofluorescence analysis of α-catenin (green) and F-actin (red) in A375 cells cultured in the presence or absence of serum (FBS), complemented or not with various growth factors, for 12 hours. Scale bar: 10 μm. Representative of 3 independent experiments. (g) Western Blot analysis of IGF1R in A375 cells stably transduced with a doxycycline-inducible vector expressing either GFP or a constitutively active form of IGF1R (CD8-IGF1R, IGF1R*) and treated with doxycycline for 12 hours. CD8-IGF1R (estimated size: 70 kDa) appears below endogenous IGF1R (estimated size: 95 kDa). Representative example of 3 independent experiments.
Extended Data Fig. 10 Role of IGF1 signaling in zebrafish and human melanoma.
(a) Tumor-free survival curves of Tg(mitf:BRAFV600E);tp53−/− zebrafish injected with a vector targeting nectin1 and vectors expressing either GFP or CD8-IGF1R (IGF1R*). Pooled data of 2 independent experiments. Log-rank test. (b) Quantification of overall p-H3 staining intensity in 8 nectin1-knockout/GFP and 8 nectin1-knockout/IGF1R* primary zebrafish melanomas. Mean ± SD. Two-tailed t-test. (c) Quantification of organ involvement in 16-week-old zebrafish bearing nectin1-knockout/GFP (n = 8) or nectin1-knockout/IGF1R* (n = 8) primary melanoma. Two-tailed t-test. (d) Progression-free survival curves of patients with melanoma stratified according to IGF1R mRNA expression (top and bottom n = 104 samples). TCGA cohort. Log-rank test. (e) Immunofluorescence analysis of α-catenin (green) and phospho-IGF1R (red) on a tissue section of human melanoma. Scale bar: 10 μm. Dotted line demarcates areas of high and low IGF1 signaling. Arrows point to adherens junctions. Representative example of 9 independent tumors. (f) Histogram representing signal intensities of α-catenin (green) and phospho-IGF1R (red) along the dotted line in the human melanoma tissue section shown on the left. Note the mutual exclusivity of the two signals.
Supplementary information
Supplementary Information
Supplementary Notes and Tables 1–4.
Supplementary Table 1
Table 1. Gene set enrichment analysis of the transcriptional difference between NECTIN1-knockout and NECTIN1-wild-type A375 human melanoma cells under serum starvation. Table 2. Gene Ontology analysis of the transcriptional response of A375 human melanoma cells to serum depletion. Table 3. Gene set enrichment analysis of the transcriptional response of A375 human melanoma cells to serum depletion. Table 4. Sequences of oligonucleotides.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Statistical Source Data.
Source Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 9
Statistical Source Data.
Source Data Extended Data Fig. 9
Unprocessed western blots.
Source Data Extended Data Fig. 10
Statistical Source Data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ablain, J., Al Mahi, A., Rothschild, H. et al. Loss of NECTIN1 triggers melanoma dissemination upon local IGF1 depletion. Nat Genet 54, 1839–1852 (2022). https://doi.org/10.1038/s41588-022-01191-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01191-z