Abstract
The UK Biobank Exome Sequencing Consortium (UKB-ESC) is a private–public partnership between the UK Biobank (UKB) and eight biopharmaceutical companies that will complete the sequencing of exomes for all ~500,000 UKB participants. Here, we describe the early results from ~200,000 UKB participants and the features of this project that enabled its success. The biopharmaceutical industry has increasingly used human genetics to improve success in drug discovery. Recognizing the need for large-scale human genetics data, as well as the unique value of the data access and contribution terms of the UKB, the UKB-ESC was formed. As a result, exome data from 200,643 UKB enrollees are now available. These data include ~10 million exonic variants—a rich resource of rare coding variation that is particularly valuable for drug discovery. The UKB-ESC precompetitive collaboration has further strengthened academic and industry ties and has provided teams with an opportunity to interact with and learn from the wider research community.
Similar content being viewed by others
Main
Developing novel therapies to address unmet medical needs is a resource-intensive challenge characterized by high attrition rates1. While biopharmaceutical companies have recently improved overall research and development productivity and success rates for drug candidates in late-stage clinical development, the probability of a drug candidate proceeding from phase 1 clinical trials through approval and launch remains near 10%. Most clinical failures (~75%) are attributable to safety concerns or a lack of efficacy2.
Why are drug developers interested in human genetics?
The potential for human genetics to increase the likelihood of successful drug discovery has long been recognized, albeit largely in anecdotal form. Anti-PCSK9 cholesterol-lowering drugs are a highly publicized example wherein human genetic evidence for a target contributed to technical and regulatory success, providing rationale for further investment3. Human genetics may also identify potential liability phenotypes associated with a target. For example, it is plausible that gastrointestinal adverse events observed in clinical trials of DGAT1 inhibitors4,5 could have been predicted based on the causal link between rare, highly penetrant DGAT1 variants and congenital diarrheal disorder6. Genetics has also proven its value in bringing more precision to drug development, particularly in the context of large trials. Stratification of patients to enrich for signal has shown success in PCSK9 inhibitors7,8, providing the ability to select genetically defined patient populations.
Recently, several studies systematically characterized the role of human genetics in drug discovery9,10,11. These retrospective analyses of drug development successes and failures demonstrate that drug–target pairs with human genetic evidence are at least twice as likely to reach approval as those without. Moreover, there is evidence to suggest that targets with clear causal relationships to disease exhibit even higher success rates10. Furthermore, genetic association with a non-relevant phenotype increases the likelihood of corresponding adverse events12.
Building on these and similar experiences, several frameworks for the systematic evaluation of genetically motivated targets have emerged. For example, the allelic series model leverages the existence of multiple, independent genetic alterations in a gene of interest to assess its potential as a drug target13. Genetic dose–response curves can be used to investigate the relationship between variants in the allelic series and phenotypes of interest to assess the potential for a tractable therapeutic window14. In addition, several promising drug development candidate targets with robust genetic evidence were identified based on associations between disease phenotypes, biomarker endo-phenotypes and functionally consequential genetic variants. Recent examples include HSD17B13 for chronic liver disease15, TYK2 for multiple autoimmune disorders16,17,18, NRXN1 for neuropsychiatric disease19 and ASGR1 for cardiovascular disease20.
Large-scale biobanks have emerged to catalyze biomedical research, in part due to widespread adoption of electronic health records and the maturation of experimental and computational platforms capable of cost-effective population-scale sequencing and analysis. These advances create a unique opportunity for the scientific community to build on these foundations and accelerate the use of human genetics to inform drug discovery. Innovative public–private partnerships in the precompetitive space have demonstrated value for furthering this acceleration21. Here, we describe one such effort—the UK Biobank Exome Sequencing Consortium (UKB-ESC)—focused on generating whole-exome sequencing (WES) data for all participants (~500,000) in the UK Biobank (UKB).
Why are drug developers interested in the UKB?
Nominating, evaluating and prioritizing potential drug targets based on human genetics is data intensive. The most actionable novel insights are likely to come from rare or private functional genetic variants leading to highly penetrant gains or losses of protein function22. These variants are most effectively discovered through direct sequencing of large populations or smaller populations likely to be enriched for variants of interest based on genetic architecture or the prevalence of specific traits. Human genetic data alone, however, are insufficient. Comprehensive, consistent, longitudinal phenotypic data that can be linked to the genetic data are needed to explore the breadth of biological consequences of genetic variation. Essential phenotypes may include accurate disease diagnosis, molecular and cellular biomarkers, treatment and outcome information, imaging endpoints, self-reported conditions and environmental and lifestyle data.
Beyond target identification, access to large-scale human genetic data linked to phenotypes enables a variety of other key opportunities for drug development efforts. In addition to direct human genetic discovery, these cohorts are incredibly valuable for instant replication of insights from other human genetics studies, disentangling causal relationships using methods such as Mendelian randomization, prediction of potential side effects, recall sub-studies, rapid validation of targets proposed by other means, and unbiased study of the natural history of rare monogenic diseases of interest to drug developers. Fifteen years ago, the Wellcome Trust Case Control Consortium shifted the field from candidate gene studies to genome-wide association studies23. Now, large biobanks are moving us even further in the unbiased spectrum by recruiting population-based cohorts without regard to disease status. The disease-agnostic approach and collection of a wide variety of data and specimens allow an immense number of hypotheses to be tested. The UKB is an exemplar of such data—a unique resource that provides a rich substrate for a broad spectrum of biomedical research24.
The UKB design allows for recontact of participants to enroll them in new scientific research projects. This mechanism is highly valuable to the scientific community (including drug developers), as it allows researchers to engage with cohorts of specific, well-characterized participants, to address emerging scientific questions. As an example, the UKB recently launched a serology study to track the extent of infection of severe acute respiratory syndrome coronavirus 2.
The objective of the UKB-ESC is a comprehensive assessment of the protein-coding genetic variation in the half-million UKB participants. As the exome is enriched for variants that are most readily interpretable and actionable, this consortium has prioritized deep sequencing of the protein-coding portions of the genome. This is the highest-value assay to have been added to the UKB genotype resource, pairing the rarest coding variation with the common and rare variation captured via chip genotyping and imputation. The speed of WES is enabling a rapid acceleration of actionable scientific discoveries. While WES is the assay of choice for Mendelian disease discovery25 and has provided important novel target discoveries15, we are also enthusiastic for the arrival of the UKB whole-genome sequencing data26 and recognize the value of continued investment in growing the UKB resource.
The enhancement of the UKB with WES data is not without limitations. It is well documented that human genetics studies are highly skewed towards populations of European ancestry27,28. Table 1 describes the clinical and demographic characteristics of the first 50,000 sequenced individuals (50,000 WES cohort), the current ~200,000 sequenced individuals (200,000 WES cohort) and the total of 500,000 participants (full UKB cohort), including information on self-reported ethnic background, which was captured in data field 21000. While the United Kingdom’s population includes individuals from diverse backgrounds, its largest ethnic group is described as White British. In addition, the full UKB cohort is not representative of the UK population as a whole when considering ethnicity, age, sex, general health status and other factors29. Researchers need to be mindful that these data include a fraction of all human genetic variation and that signals derived from the UKB may not generalize to other populations.
Lessons from drug discovery collaboration in genomics
Large-scale collaborative projects have driven transformative scientific discoveries, particularly in genetics and genomics24,30,31,32,33,34,35 (see also https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga). Such collaborations provide a unique opportunity to unify scientific communities by enabling a diversity of voices and perspectives to produce insights that would be inaccessible to smaller, more homogeneous groups. Historically, large-scale collaborative life science projects have typically been funded by governments and non-profit organizations to engage academic groups. More recently, government agencies, non-profit organizations and academic institutions have sought industry partners for large-scale scientific projects (https://www.nih.gov/research-training/accelerating-medicines-partnership-amp, https://www.alzdiscovery.org/). The UKB-ESC seeks to further strengthen the ties between academia and industry through a precompetitive collaboration. This model is enabled by a project structure that engages all parties as scientific contributors and incentivizes industry investment in a sharable data resource. We expect that the output of this partnership will catalyze novel scientific discoveries, accelerate the development of new therapies and ultimately improve patient outcomes. As both the co-funders of the UKB WES data and the scientific teams who are analyzing them, we have found the following principles essential to the success of this effort.
First and foremost, the UKB-ESC is uniquely enabled by the UKB open data access policy. The biomedical research community has already made great use of UKB data, as evidenced by the rapidly growing body of scientific literature (https://www.ukbiobank.ac.uk/enable-your-research/publications). The UKB data access model is valuable in how it enables researchers to openly publish and commercialize results from and derivatives of the UKB data and incentivizes contributions back to the resource. This cycle enables academia and industry to be engaged and benefit from a body of work larger than any one entity can achieve alone.
Second, the UKB-ESC scale and scope will enable unique, valuable scientific discoveries. The search for actionable genetic variants (for example, rare with profound functional consequences or common with informative phenotypic associations) depends on the scale of available data, in terms of both sample size and phenotypic characterization. Building large, diverse and deeply characterized cohorts is an enormous multidisciplinary undertaking. Such work is particularly challenging in high-value phenotyping wherein different modalities (for example, imaging, proteomics and electronic medical record data extraction) require diverse expertise, and the data often require extensive curation and processing before research use. The UKB excels at such tasks within a user-friendly framework attractive to both academic and industrial researchers, creating a roadmap for other projects with similar aspirations.
Third, the UKB’s data access and contribution terms invite precompetitive collaboration by industry partners. The generation of sequencing data at this scale requires a substantial investment of time, personnel and financial resources36. The UKB policy of providing a window of data exclusivity for data generators (also a common feature of large-scale academic collaborations) was essential to the business case for an investment of this scale. In addition to the scientific drivers described above, participation in a consortium and the exclusivity period both provide tangible benefits. While the competitive commercial interests of companies may seem to preclude cooperation, an appropriately managed collaboration mitigates individual risk to each partner and provides value larger than the sum of the individual investments. Moreover, the finite window of exclusivity is an incentive for the partner companies to focus their initial efforts on projects that are likely to have near-term impact on their pipelines.
Finally, the value of engagement in a large, precompetitive industry collaborative project provides additional value to the participating institutions. Building expertise and functional excellence in important new areas is a key benefit to participating institutions. In our experience, drug discovery teams typically have deep experience in areas such as disease biology, chemistry and translational research. In contrast, the inclusion of human genetics expertise varies between companies and therapeutic areas and is often rate limited by the availability of relevant datasets. Participation in the UKB-ESC provided member companies with a unique opportunity to build out or enhance the expertise needed to conduct population-scale genomic medicine research without each incurring the full overhead of creating the underlying resource. The resulting combination of scientific expertise and a critical mass of data serves as a multiplier for extant research programs, complementing existing genomics data, such as targeted clinical sequencing of probands and families, and providing a roadmap for at-scale genomics in therapeutic areas that remain under-represented in genetic data. We anticipate a virtuous cycle between the increasing availability of large, well-annotated genetics resources and increased industry investment in genetics.
In summary, we have identified the following key features of an effective precompetitive industry collaboration: (1) build a large dataset with rich phenotypic characterization, noting that participant recontact is highly valued for answering new questions that emerge from the data; (2) provide researchers with the opportunity to derive both academic and commercial value from the data in an unencumbered way; (3) provide some exclusivity/first-mover advantage to build a compelling business justification for participation and financing of the project; (4) enable data access providing insight generation for key internal therapeutic-focused scientists and research and development stakeholders within the collaborating institutions; and (5) provide opportunities for constructive engagement with academic partners, as well as low-friction data-sharing terms and platforms to enable the broadest possible suite of research projects. We are hopeful that others will follow the UKB model and build substantial data collections designed to engage a wide variety of stakeholders.
Results
The release of the first 200,633 sequenced exomes represents a milestone in the availability of large-scale genomics data. This release is inclusive of the first 50,000 UKB samples37, the sequencing and initial release of which were funded through a separate mechanism independent of the UKB-ESC. Here, we provide an overview of the phenotypes and genotypes available as part of this resource. Table 1 includes a summary of the clinical characteristics of the 50,000 WES, 200,000 WES and full UKB cohorts. Definitions of the UKB phenotypes are provided in ref. 37. Table 2 summarizes the variants observed in the first 200,000 exomes sequenced from the UKB (that is, the 200,000 WES cohort). The targeted region covered 38,997,831 bases and coverage exceeded 20× on 95.6% of the sites on average. Approximately 10 million variants were observed within the targeted regions. These include 8,086,176 single-nucleotide variants (SNVs), 370,958 indels and 1,596,984 multi-allelic variants. Of the ~8 million SNVs observed, 84.5% are coding variants and include 2,139,318 (25.3%) synonymous variants, 4,549,694 (53.8%) missense variants and 453,733 (5.4%) predicted loss-of-function (LOF) variants (initiation codon loss, premature stop codons, stop codon loss or splicing and frameshift variants) affecting at least one coding transcript. Analysis of allele frequencies revealed 98% of synonymous, 99% of missense and 99% of LOF (at least one transcript) variants with a minor allele frequency (MAF) of <1%. On a per-individual basis, we observed a median number of 8,586 synonymous, 7,724 missense and 202 LOF variants. Restricting our analysis to variants that affect canonical transcripts, we observed a median of 142 LOFs per individual. The numbers are largely similar to those in published exome studies37,38.
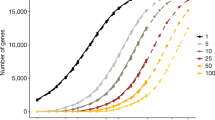
In addition, we also analyzed the increase in the number of genes with heterozygous and homozygous LOFs with the increase in the number of sequenced samples. Previously, 17,718 genes with heterozygous LOF variants were observed in the 50,000 WES data37; therefore, is it not surprising that we only observed a modest increase in that number (18,045 genes) in the 200,000 WES data. A total of 789 genes with at least one homozygous LOF variant were reported in the 50,000 WES data. The number of genes with homozygous LOFs still appears to be increasing, with a projection of 1,981 genes with at least one homozygous LOF variant in the 500,000 participants of the full UKB cohort (1,492 genes with at least one homozygous LOF seen in the 200,000 WES cohort; Table 3). The tolerance of homozygous LOF variants is of particular interest to target development programs, although characterization of homozygous LOFs in all human genes will not be accomplished by scale alone39. Rather, the increasing number of genes tolerating homozygous LOFs identified in the UKB exomes can complement smaller study designs, such as ancestry-specific population sequencing and consanguineous cohorts.
Discussion
The UKB exists to enable scientific research with the ultimate aim of improving human health. The UKB-ESC is proud of its contribution to the scientific community, enthusiastic about making these data broadly available and excited to see what the future holds in terms of discoveries from and contributions to the UKB and UKB-ESC resources, particularly as additional insight is gained into functionally consequential variants that can meaningfully inform drug development. Data and methods developed by the UKB-ESC have already contributed to multiple publications40,41,42,43,44.
With the UKB-ESC exome sequencing nearly complete, we hope the key features of this collaboration will be adopted as the preferred model for similar projects in the future. We expect that the value of the WES data will be enhanced by layering deeper and richer phenotypes, which can provide important insights into disease biology, characterize responses to marketed therapies and identify novel targets. Thoughtfully designed projects that maximize the engagement of academia, non-profit organizations and industry will yield valuable data resources and scientific insights that accelerate drug discovery and personalized health care. As an example, the recently announced Pharma Proteomics Project will use a model similar to that of the UKB-ESC to generate high-dimensional proteomics data on approximately 53,000 participants45. The groundwork laid by these private–public partnerships will nucleate further partnerships by lowering entry costs: existing partners already have relationships established and new partners have example data and contractual frameworks to draw on to make the business case.
Developing new therapies to treat unmet medical needs is among the most important scientific challenges we face. Large-scale collaborations such as the UKB-ESC play an essential role by generating unique, accessible resources that can be used by a diverse community of researchers to address questions critical to advancing human health.
Data availability
The UKB aims to encourage and provide the widest possible access to its data and samples for health-related research in the public interest performed by all bona fide researchers from the academic, charity, public and commercial sectors, both in the United Kingdom and internationally, without preferential access for any user. The UKB’s publicly available Data Showcase (http://biobank.ndph.ox.ac.uk/showcase/) presents the univariate distributions and methods used for collection of all of the variables available for health-related research, enabling potential research users to explore which data are available and to plan research applications. All researchers who wish to access the resource must register with the UKB via its online access management system (https://bbams.ndph.ox.ac.uk/ams/). Once approved, researchers may apply (via the access management system) to access the resource for specific, well-defined research projects. At the time of publication, over 16,500 researchers were registered with the UKB and over 2,000 research applications were approved (see https://www.ukbiobank.ac.uk/enable-your-research/approved-research for a summary of research that is currently underway).
References
DiMasi, J. A., Grabowski, H. G. & Hansen, R. W. Innovation in the pharmaceutical industry: new estimates of R&D costs. J. Health Econ. 47, 20–33 (2016).
Dowden, H. & Munro, J.Trends in clinical success rates and therapeutic focus. Nat. Rev. Drug Discov. 18, 495–496 (2019).
Furtado, R. H. M. & Giugliano, R. P.What lessons have we learned and what remains to be clarified for PCSK9 inhibitors? A review of FOURIER and ODYSSEY outcomes trials. Cardiol. Ther. 9, 59–73 (2020).
Denison, H. et al. Proof of mechanism for the DGAT1 inhibitor AZD7687: results from a first-time-in-human single-dose study. Diabetes Obes. Metab. 15, 136–143 (2013).
Meyers, C. D. et al. Effect of the DGAT1 inhibitor pradigastat on triglyceride and apoB48 levels in patients with familial chylomicronemia syndrome. Lipids Health Dis. 14, 8 (2015).
Haas, J. T. et al. DGAT1 mutation is linked to a congenital diarrheal disorder. J. Clin. Invest. 122, 4680–4684 (2012).
Sabatine, M. S. et al. Evolocumab and clinical outcomes in patients with cardiovascular disease. N. Engl. J. Med. 376, 1713–1722 (2017).
Schwartz, G. G. et al. Alirocumab and cardiovascular outcomes after acute coronary syndrome. N. Engl. J. Med. 379, 2097–2107 (2018).
Cook, D. et al. Lessons learned from the fate of AstraZeneca’s drug pipeline: a five-dimensional framework. Nat. Rev. Drug Discov. 13, 419–431 (2014).
King, E. A., Wade Davis, J. & Degner, J. F. Are drug targets with genetic support twice as likely to be approved? Revised estimates of the impact of genetic support for drug mechanisms on the probability of drug approval. PLoS Genet. 15, e1008489 (2019).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Nguyen, P. A., Born, D. A., Deaton, A. M., Nioi, P. & Ward, L. D. Phenotypes associated with genes encoding drug targets are predictive of clinical trial side effects. Nat. Commun. 10, 1579 (2019).
McKusick, V. A.Phenotypic diversity of human diseases resulting from allelic series. Am. J. Hum. Genet. 25, 446–456 (1973).
Plenge, R. M., Scolnick, E. M. & Altshuler, D.Validating therapeutic targets through human genetics. Nat. Rev. Drug Discov. 12, 581–594 (2013).
Abul-Husn, N. S. et al. A protein-truncating HSD17B13 variant and protection from chronic liver disease. N. Engl. J. Med. 378, 1096–1106 (2018).
Dendrou, C. A. et al. Resolving TYK2 locus genotype-to-phenotype differences in autoimmunity. Sci. Transl. Med. 363, 363ra149 (2016).
Diogo, D. et al. TYK2 protein-coding variants protect against rheumatoid arthritis and autoimmunity, with no evidence of major pleiotropic effects on non-autoimmune complex traits. PLoS ONE 10, e0122271 (2015).
Minegishi, Y. et al. Human tyrosine kinase 2 deficiency reveals its requisite roles in multiple cytokine signals involved in innate and acquired immunity. Immunity 25, 745–755 (2006).
Noh, H. J. et al. Integrating evolutionary and regulatory information with a multispecies approach implicates genes and pathways in obsessive-compulsive disorder. Nat. Commun. 8, 774 (2017).
Nioi, P. et al. Variant ASGR1 associated with a reduced risk of coronary artery disease. N. Engl. J. Med. 374, 2131–2141 (2016).
Dolgin, E. Massive NIH–industry project opens portals to target validation. Nat. Rev. Drug Discov. https://doi.org/10.1038/d41573-019-00033-8 (2019).
McCarthy, M. I. et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat. Rev. Genet. 9, 356–369 (2008).
Burton, P. R. et al. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Yang, Y. et al. Clinical whole-exome sequencing for the diagnosis of Mendelian disorders. N. Engl. J. Med. 369, 1502–1511 (2013).
UK Biobank leads the way in genetics research to tackle chronic diseases. UK Biobank https://www.ukbiobank.ac.uk/learn-more-about-uk-biobank/news/uk-biobank-leads-the-way-in-genetics-research-to-tackle-chronic-diseases-1 (2019).
Diversity matters. Nat. Rev. Genet. 20, 495 (2019).
Gurdasani, D., Barroso, I., Zeggini, E. & Sandhu, M. S.Genomics of disease risk in globally diverse populations. Nat. Rev. Genet. 20, 520–535 (2019).
Fry, A. et al. Comparison of sociodemographic and health-related characteristics of UK Biobank participants with those of the general population. Am. J. Epidemiol. 186, 1026–1034 (2017).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Methé, B. A. et al. A framework for human microbiome research. Nature 486, 215–221 (2012).
Holden, A. L., Contreras, J. L., John, S. & Nelson, M. R.The international serious adverse events consortium. Nat. Rev. Drug Discov. 13, 795–796 (2014).
Regeneron announces major collaboration to exome sequence UK Biobank genetic data more quickly. UK Biobank https://www.ukbiobank.ac.uk/learn-more-about-uk-biobank/news/regeneron-announces-major-collaboration-to-exome-sequence-uk-biobank-genetic-data-more-quickly (2018).
Van Hout, C. V. et al. Exome sequencing and characterization of 49,960 individuals in the UK Biobank. Nature 586, 749–756 (2020).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Minikel, E. V. et al. Evaluating drug targets through human loss-of-function genetic variation. Nature 581, 459–464 (2020).
Liu, J. Z. et al. The burden of rare protein-truncating genetic variants on human lifespan. Preprint at bioRxiv https://doi.org/10.1101/2020.06.02.129908 (2020).
Povysil, G. et al. Assessing the role of rare genetic variation in patients with heart failure. J. Am. Med. Assoc. Cardiol. 6, 379–386 (2021).
Carss, K. J. et al. Spontaneous coronary artery dissection: insights on rare genetic variation from genome sequencing. Circ. Genom. Precis. Med. 13, e003030 (2020).
Dhindsa, R. et al. Identification of a novel missense variant in SPDL1 associated with idiopathic pulmonary fibrosis. Commun. Biol. 4, 392 (2021).
Cameron-Christie, S. et al. A broad exome study of the genetic architecture of asthma reveals novel patient subgroups. Preprint at bioRxiv https://doi.org/10.1101/2020.12.10.419663 (2020).
UK Biobank launches one of the largest scientific studies measuring circulating proteins, to better understand the link between genetics and human disease. UK Biobank https://www.ukbiobank.ac.uk/learn-more-about-uk-biobank/news/uk-biobank-launches-one-of-the-largest-scientific-studies (2020).
Acknowledgements
We thank everyone who made this work possible, including the UKB team, their funders and the dedicated professionals from the member institutions who contributed to and supported this work. We are especially grateful to the UKB participants who generously volunteered to take part in this research. This research has been conducted using the UKB resource under application number 26041.
Author information
Authors and Affiliations
Consortia
Contributions
J.D.S., P.G.B., E.W., J.W.D., C.H., A.K.L., R.M., H.J.N., S.W., E.N.S., J.F.W., C.D.W., E.A.T., J.D.O., W.J.S., H.J., S.S., H.R., G.H., P.N., S.P., M.R.M., A.B., L.J.M. and J.G.R. jointly supervised the research. J.D.S., W.J.S., A.B. and J.G.R. conceived of and designed the experiments. A.E.L. and J.D.O. performed the experiments. J.D.S., S.B., A.S., S.K., E.K., D.L., X.B., A.H., O.K. and W.J.S. analyzed the data. X.B., A.H., O.K., R.U., W.J.S. and J.G.R. contributed reagents, materials and/or analysis tools. J.D.S., S.B., A.S., P.G.B., E.K., W.J.S., S.S., H.R., P.N., S.P., M.R.M. and J.G.R. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
J.D.S. is employed by and owns stocks in Bristol Myers Squibb. C.D.W. and H.R. are employees of Biogen and hold stock options at Biogen. E.N.S. is an employee of Takeda California. S.S. is employed by Takeda California and is a shareholder of Takeda and Johnson & Johnson. H.J.N. and J.W.D. are employed by AbbVie and may own AbbVie stock and/or options. A.K.L. is an employee of Pfizer. M.R.M. is an employee and stockholder of Pfizer. P.N. is an employee of and stockholder in Alnylam Pharmaceuticals. G.H. is a paid consultant to 54gene—a Nigerian genomics company. C.H. and S.P. are employees and stockholders of AstraZeneca. W.J.S., R.U., A.H., O.K., S.K., D.L., X.B., A.E.L., S.B., A.B., J.D.O. and L.J.M. are full-time employees of the Regeneron Genetics Center of Regeneron Pharmaceuticals and receive stock options and restricted stock units as compensation. J.G.R. is a full-time employee of the Regeneron Genetics Center of Regeneron Pharmaceuticals and receives stock options and restricted stock units as compensation. J.G.R. also provides (unpaid) advice, support and guidance to the UKB as a member of the UKB International Scientific Advisory Board. The other authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks Amalio Telenti, Matthew Nelson and Kaja Wasik for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Note
Rights and permissions
About this article
Cite this article
Szustakowski, J.D., Balasubramanian, S., Kvikstad, E. et al. Advancing human genetics research and drug discovery through exome sequencing of the UK Biobank. Nat Genet 53, 942–948 (2021). https://doi.org/10.1038/s41588-021-00885-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00885-0
This article is cited by
-
Genetic associations of protein-coding variants in venous thromboembolism
Nature Communications (2024)
-
Inherited polygenic effects on common hematological traits influence clonal selection on JAK2V617F and the development of myeloproliferative neoplasms
Nature Genetics (2024)
-
Large-scale whole-exome sequencing of neuropsychiatric diseases and traits in 350,770 adults
Nature Human Behaviour (2024)
-
RNA interference in the era of nucleic acid therapeutics
Nature Biotechnology (2024)
-
Exome-wide analysis implicates rare protein-altering variants in human handedness
Nature Communications (2024)