Abstract
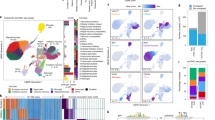
Genome-wide association studies have discovered hundreds of loci associated with complex brain disorders, but it remains unclear in which cell types these loci are active. Here we integrate genome-wide association study results with single-cell transcriptomic data from the entire mouse nervous system to systematically identify cell types underlying brain complex traits. We show that psychiatric disorders are predominantly associated with projecting excitatory and inhibitory neurons. Neurological diseases were associated with different cell types, which is consistent with other lines of evidence. Notably, Parkinson’s disease was genetically associated not only with cholinergic and monoaminergic neurons (which include dopaminergic neurons) but also with enteric neurons and oligodendrocytes. Using post-mortem brain transcriptomic data, we confirmed alterations in these cells, even at the earliest stages of disease progression. Our study provides an important framework for understanding the cellular basis of complex brain maladies, and reveals an unexpected role of oligodendrocytes in Parkinson’s disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All single-cell expression data are publicly available. Most summary statistics used in this study are publicly available. The migraine GWAS60 can be obtained by contacting the authors of that study. The full Parkinson’s disease summary statistics from 23andMe can be obtained under an agreement that protects the privacy of 23andMe research participants (https://research.23andme.com/collaborate/#publication). The 10,000 most associated SNPs from the 23andMe cohort are available in Supplementary Table 12.
Code availability
The code used to generate these results is available at https://github.com/jbryois/scRNA_disease. An R package for performing cell-type enrichments using MAGMA is also available from https://github.com/NathanSkene/MAGMA_Celltyping.
References
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Lee, J. J., Wedow, R. & Okbay Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121 (2018).
Nagel, M. et al. Meta-analysis of genome-wide association studies for neuroticism in 449,484 individuals identifies novel genetic loci and pathways. Nat. Genet. 50, 920–927 (2018).
Yengo, L. et al. Meta-analysis of genome-wide association studies for height and body mass index in ∼700000 individuals of European ancestry. Hum. Mol. Genet. 27, 3641–3649 (2018).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Akbarian, S. et al. The PsychENCODE project. Nat. Neurosci. 18, 1707–1712 (2015).
Aguet, F. et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–329 (2015).
Ongen, H. et al. Estimating the causal tissues for complex traits and diseases. Nat. Genet. 49, 1676–1683 (2017).
Skene, N. G. & Grant, S. G. N. Identification of vulnerable cell types in major brain disorders using single cell transcriptomes and expression weighted cell type enrichment. Front. Neurosci. 10, 1–11 (2016).
Skene, N. G. et al. Genetic identification of brain cell types underlying schizophrenia. Nat. Genet. 50, 825–833 (2018).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
Calderon, D. et al. Inferring relevant cell types for complex traits by using single-cell gene expression. Am. J. Hum. Genet. 101, 686–699 (2017).
Savage, J. E. et al. Genome-wide association meta-analysis in 269,867 individuals identifies new genetic and functional links to intelligence. Nat. Genet. 50, 912–919 (2018).
Coleman, J. R. I. et al. Biological annotation of genetic loci associated with intelligence in a meta-analysis of 87,740 individuals. Mol. Psychiatry 24, 182–197 (2019).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Nalls, M. A. et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 18, 1091–1102 (2019).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, 1–19 (2015).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Jevtic, S., Sengar, A. S., Salter, M. W. & McLaurin, J. A. The role of the immune system in Alzheimer disease: etiology and treatment. Ageing Res. Rev. 40, 84–94 (2017).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
O’Leary, D. H. et al. Carotid-artery intima and media thickness as a risk factor for myocardial infarction and stroke in older adults. N. Engl. J. Med. 340, 14–22 (1999).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014.e22 (2018).
Anttila, V. et al. Analysis of shared heritability in common disorders of the brain. Science 360, (2018).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Braak, H. et al. Staging of brain pathology related to sporadic Parkinson’s disease. Neurobiol. Aging 24, 197–211 (2003).
Sulzer, D. & Surmeier, D. J. Neuronal vulnerability, pathogenesis, and Parkinson’s disease. Mov. Disord. 28, 41–50 (2013).
Poewe, W. et al. Parkinson disease. Nat. Rev. Dis. Primers 3, 17013 (2017).
Halliday, G. M. et al. Neuropathology of immunohistochemically identified brainstem neurons in Parkinson’s disease. Ann. Neurol. 27, 373–385 (1990).
Delaville, C., de Deurwaerdère, P. & Benazzouz, A. Noradrenaline and Parkinson’s disease. Front. Syst. Neurosci. https://doi.org/10.3389/fnsys.2011.00031 (2011).
Rinne, J. O., Ma, S. Y., Lee, M. S., Collan, Y. & Röyttä, M. Loss of cholinergic neurons in the pedunculopontine nucleus in Parkinson’s disease is related to disability of the patients. Parkinsonism Relat. Disord. 14, 553–557 (2008).
Braak, H., Rüb, U., Gai, W. P. & Del Tredici, K. Idiopathic Parkinson’s disease: possible routes by which vulnerable neuronal types may be subject to neuroinvasion by an unknown pathogen. J. Neural Transm. 110, 517–536 (2003).
Liddle, R. A. Parkinson’s disease from the gut. Brain Res. 1693, 201–206 (2018).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030.e16 (2018).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955 (2017).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Lesnick, T. G. et al. A genomic pathway approach to a complex disease: axon guidance and Parkinson disease. PLoS Genet. 3, 0984–0995 (2007).
Moran, L. B. et al. Whole genome expression profiling of the medial and lateral substantia nigra in Parkinson’s disease. Neurogenetics 7, 1–11 (2006).
Kannarkat, G. T., Boss, J. M. & Tansey, M. G. The role of innate and adaptive immunity in Parkinson’s disease. J. Parkinsons Dis. 3, 493–514 (2013).
Gagliano, S. A. et al. Genomics implicates adaptive and innate immunity in Alzheimer’s and Parkinson’s diseases. Ann. Clin. Transl. Neurol. 3, 924–933 (2016).
Dijkstra, A. A. et al. Evidence for immune response, axonal dysfunction and reduced endocytosis in the substantia nigra in early stage Parkinson’s disease. PLoS ONE 10, e0128651 (2015).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Sathyamurthy, A. et al. Massively parallel single nucleus transcriptional profiling defines spinal cord neurons and their activity during behavior. Cell Rep. 22, 2216–2225 (2018).
Lake, B. B. et al. A comparative strategy for single-nucleus and single-cell transcriptomes confirms accuracy in predicted cell-type expression from nuclear RNA. Sci. Rep. 7, 6031 (2017).
Caspi, A. et al. The p factor: one general psychopathology factor in the structure of psychiatric disorders? Clin. Psychol. Sci. 2, 119–137 (2014).
Sullivan, P. F. & Geschwind, D. H. Defining the genetic, genomic, cellular, and diagnostic architectures of psychiatric disorders. Cell 177, 162–183 (2019).
Reynolds, R. H. et al. Moving beyond neurons: the role of cell type-specific gene regulation in Parkinson’s disease heritability. NPJ Parkinsons Dis. 5, 6 (2019).
Singaram, C. et al. Dopaminergic defect of enteric nervous system in Parkinson’s disease patients with chronic constipation. Lancet 346, 861–864 (1995).
Wakabayashi, K., Takahashi, H., Takeda, S., Ohama, E. & Ikuta, F. Lewy bodies in the enteric nervous system in Parkinson’s disease. Arch. Histol. Cytol. 52, 191–194 (1989).
Stokholm, M. G., Danielsen, E. H., Hamilton-Dutoit, S. J. & Borghammer, P. Pathological α-synuclein in gastrointestinal tissues from prodromal Parkinson disease patients. Ann. Neurol. 79, 940–949 (2016).
Svensson, E. et al. Vagotomy and subsequent risk of Parkinson’s disease. Ann. Neurol. 78, 522–529 (2015).
Gilman, S. et al. Second consensus statement on the diagnosis of multiple system atrophy. Neurology 71, 670–676 (2008).
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat. Genet. 46, 989–993 (2014).
Wakabayashi, K., Hayashi, S., Yoshimoto, M., Kudo, H. & Takahashi, H. NACP/α-synuclein-positive filamentous inclusions in astrocytes and oligodendrocytes of Parkinson’s disease brains. Acta Neuropathol. 99, 14–20 (2000).
Seidel, K. et al. The brainstem pathologies of Parkinson’s disease and dementia with Lewy bodies. Brain Pathol. 25, 121–135 (2015).
Stahl, E. A. et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat. Genet. 51, 793–803 (2019).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681 (2018).
Perry, J. R. B. et al. Parent-of-origin-specific allelic associations among 106 genomic loci for age at menarche. Nature 514, 92–97 (2014).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Gormley, P. et al. Meta-analysis of 375,000 individuals identifies 38 susceptibility loci for migraine. Nat. Genet. 48, 1296 (2016).
Van Rheenen, W. et al. Genome-wide association analyses identify new risk variants and the genetic architecture of amyotrophic lateral sclerosis. Nat. Genet. 48, 1043–1048 (2016).
Demontis, D. et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat. Genet. 51, 63–75 (2019).
Day, F. R. et al. Large-scale genomic analyses link reproductive aging to hypothalamic signaling, breast cancer susceptibility and BRCA1-mediated DNA repair. Nat. Genet. 47, 1294–1303 (2015).
Nelson, C. P. et al. Association analyses based on false discovery rate implicate new loci for coronary artery disease. Nat. Genet. 49, 1385–1391 (2017).
Wheeler, E. et al. Impact of common genetic determinants of Hemoglobin A1c on type 2 diabetes risk and diagnosis in ancestrally diverse populations. PLoS Med. 14, 1–30 (2017).
Hibar, D. P. et al. Novel genetic loci associated with hippocampal volume. Nat. Commun. 8, 13624 (2017).
de Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Adams, H. H. H. et al. Novel genetic loci underlying human intracranial volume identified through genome-wide association. Nat. Neurosci. 19, 1569–1582 (2016).
Malik, R. et al. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat. Genet. 50, 524–537 (2018).
Scott, R. A. et al. An expanded genome-wide association study of type 2 diabetes in Europeans. Diabetes 66, 2888–2902 (2017).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Watson, H. J. et al. Genome-wide association study identifies eight risk loci and implicates metabo-psychiatric origins for anorexia nervosa. Nat. Genet. 51, 1207–1214 (2019).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Watanabe, K., Umićević Mirkov, M., de Leeuw, C. A., van den Heuvel, M. P. & Posthuma, D. Genetic mapping of cell type specificity for complex traits. Nat. Commun. 10, 3222 (2019).
Cajigas, I. J. et al. The local transcriptome in the synaptic neuropil revealed by deep sequencing and high-resolution imaging. Neuron 74, 453–466 (2012).
Alexa, A. & Rahnenfuhrer, J. topGO: enrichment analysis for Gene Ontology v.2.36 (2016).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. Affy - analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Acknowledgements
J.B. was funded by a grant from the Swiss National Science Foundation (P400PB_180792). N.G.S. was supported by the Wellcome Trust (108726/Z/15/Z). N.G.S. and L.B. performed part of the work at the Systems Genetics of Neurodegeneration summer school funded by BMBF as part of the e:Med programme (FKZ 01ZX1704). J.H.-L. was funded by the Swedish Research Council (Vetenskapsrådet, award 2014-3863), StratNeuro, the Wellcome Trust (108726/Z/15/Z) and the Swedish Brain Foundation (Hjärnfonden). P.F.S. was supported by the Swedish Research Council (Vetenskapsrådet, award D0886501), the Horizon 2020 Program of the European Union (COSYN, RIA grant agreement no. 610307) and US NIMH (U01 MH109528 and R01 MH077139). E.A. was supported by the Swedish Research Council (VR 2016-01526), Swedish Foundation for Strategic Research (SLA SB16-0065), Karolinska Institutet (SFO Strat. Regen., Senior grant 2018), Cancerfonden (CAN 2016/572), Hjärnfonden (FO2017-0059) and Chen Zuckeberg Initiative: Neurodegeneration Challenge Network (2018-191929-5022). C.M.B. acknowledges funding from the Swedish Research Council (Vetenskapsrådet, award: 538-2013-8864) and the Klarman Family Foundation. This work is supported by the UK Dementia Research Institute, which receives its funding from UK DRI Ltd, funded by the UK Medical Research Council, Alzheimer’s Society and Alzheimer’s Research UK. We thank the research participants from 23andMe and other cohorts for their contribution to this study.
Author information
Authors and Affiliations
Consortia
Contributions
J.B., N.G.S., J.H.-L. and P.F.S. designed the study, and wrote and reviewed the manuscript; J.B. performed the analyses pertaining to Figs. 1–4, Extended Data Figs. 1–10, Supplementary Figs. 1–5 and 7–20, and Supplementary Tables 1–9 and 12–17; N.G.S performed the analyses pertaining to Fig. 5, Supplementary Fig. 6 and Supplementary Tables 10 and 11; T.F.H., L.J.A.K. and the I.H.G.C. provided the migraine GWAS summary statistics; H.J.W., the E.D.W.G.P.G.C., G.B. and C.M.B. performed the anorexia GWAS; Z.L. contributed to the revision of the manuscript, the 23andMe R.T. provided GWAS summary statistics for Parkinson’s disease in the 23andMe cohort. L.B. contributed to the post-mortem differential expression analysis (Fig. 5); E.A. provided expert knowledge on Parkinson’s disease and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
P.F.S. reports the following potentially competing financial interests: current—Lundbeck (advisory committee, grant recipient); past three years—Pfizer (scientific advisory board), Element Genomics (consultation fee) and Roche (speaker reimbursement). C.M.B. reports: Shire (grant recipient, Scientific Advisory Board member); Pearson and Walker (author, royalty recipient).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Enrichment of immune genes in GTEx tissues.
Enrichment pvalues of genes belonging to the GO term ‘Immune System Process’ in the 10% most specific genes in each tissue. The one-sided pvalues were computed using linear regression, testing whether the average specificity metric of the gene set was higher than 0 (z-scaled specificity metrics per tissue). The GO term was selected because it is the most associated with inflammatory bowel disease using MAGMA.
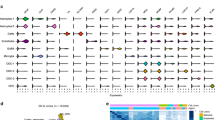
Extended Data Fig. 2 Associations of brain related traits with cell types from the entire mouse nervous system.
Associations of the top 15 most associated cell types are shown. The mean strength of association (-log10P) of MAGMA and LDSC is shown and the bar color indicates whether the cell type is significantly associated with both methods, one method or none (significance threshold: 5% false discovery rate).
Extended Data Fig. 3 Correlation in cell type associations across traits.
The Spearman rank correlations between the cell types associations across traits (-log10P) are shown. SCZ (schizophrenia), EDU (educational attainment), INT (intelligence), BMI (body mass index), BIP (bipolar disorder), NEU (neuroticism), PAR (Parkinson’s disease), MDD (Major depressive disorder), MEN (age at menarche), ICV (intracranial volume), ASD (autism spectrum disorder), STR (stroke), AN (anorexia nervosa), MIG (migraine), ALS (amyotrophic lateral sclerosis), ADHD (attention deficit hyperactivity disorder), ALZ (Alzheimer’s disease), HIP (hippocampal volume).
Extended Data Fig. 4 Associations of brain related traits with neurons from the central nervous system.
Associations of the 15 most associated neurons from the central nervous system (CNS) are shown. The specificity metrics were computed only using neurons from the CNS. The mean strength of association (-log10P) of MAGMA and LDSC is shown and the bar color indicates whether the cell type is significantly associated with both methods, one method or none (significance threshold: 5% false discovery rate).
Extended Data Fig. 5 Associations of cell types with schizophrenia/cognitive traits conditioning on gene-level genetic association of cognitive traits/schizophrenia.
MAGMA association strength for each cell type before and after conditioning on gene-level genetic association for another trait. The black bar represents the significance threshold (5% false discovery rate). SCZ (schizophrenia), INT (intelligence), EDU (educational attainment).
Extended Data Fig. 6 Replication of cell type—trait associations in 88 cell types from 9 different brain regions.
The mean strength of association (-log10P) of MAGMA and LDSC is shown for the 15 top cell types for each trait. The bar color indicates whether the cell type is significantly associated with both methods, one method or none (significance threshold: 5% false discovery rate).
Extended Data Fig. 7 Correlation in cell type associations across traits in a replication data set (88 cell types, 9 brain regions).
Spearman rank correlations for cell types associations (-log10P) across traits are shown. SCZ (schizophrenia), EDU (educational attainment), INT (intelligence), BMI (body mass index), BIP (bipolar disorder), NEU (neuroticism), PAR (Parkinson’s disease), MDD (Major depressive disorder), MEN (age at menarche), ICV (intracranial volume), ASD (autism spectrum disorder), STR (stroke), AN (anorexia nervosa), MIG (migraine), ALS (amyotrophic lateral sclerosis), ADHD (attention deficit hyperactivity disorder), ALZ (Alzheimer’s disease), HIP (hippocampal volume).
Extended Data Fig. 8 Associations of brain related traits with neurons from 9 different brain regions.
Trait—neuron association are shown for neurons of the 9 different brain regions. The specificity metrics were computed only using neurons. The mean strength of association (-log10P) of MAGMA and LDSC is shown and the bar color indicates whether the cell type is significantly associated with both methods, one method or none (significance threshold: 5% false discovery rate).
Extended Data Fig. 9 Top associated cell types with brain related traits among 24 cell types from 5 different brain regions.
The mean strength of association (-log10P) of MAGMA and LDSC is shown for the 15 top cell types for each trait. The bar color indicates whether the cell type is significantly associated with both methods, one method or none (significance threshold: 5% false discovery rate).
Extended Data Fig. 10 Top associated neurons with brain related traits among 16 neurons from 5 different brain regions.
The specificity metrics were computed only using neurons. The mean strength of association (-log10P) of MAGMA and LDSC is shown for the top 15 cell types for each trait. The bar color indicates whether the cell type is significantly associated with both methods, one method or none (significance threshold= 5% false discovery rate).
Supplementary information
Supplementary Information
Supplementary Figs. 1–20 and Note
Supplementary Tables
Supplementary Tables 1–17
Rights and permissions
About this article
Cite this article
Bryois, J., Skene, N.G., Hansen, T.F. et al. Genetic identification of cell types underlying brain complex traits yields insights into the etiology of Parkinson’s disease. Nat Genet 52, 482–493 (2020). https://doi.org/10.1038/s41588-020-0610-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0610-9