Abstract
Mutations in genes involved in DNA methylation (DNAme; for example, TET2 and DNMT3A) are frequently observed in hematological malignancies1,2,3 and clonal hematopoiesis4,5. Applying single-cell sequencing to murine hematopoietic stem and progenitor cells, we observed that these mutations disrupt hematopoietic differentiation, causing opposite shifts in the frequencies of erythroid versus myelomonocytic progenitors following Tet2 or Dnmt3a loss. Notably, these shifts trace back to transcriptional priming skews in uncommitted hematopoietic stem cells. To reconcile genome-wide DNAme changes with specific erythroid versus myelomonocytic skews, we provide evidence in support of differential sensitivity of transcription factors due to biases in CpG enrichment in their binding motif. Single-cell transcriptomes with targeted genotyping showed similar skews in transcriptional priming of DNMT3A-mutated human clonal hematopoiesis bone marrow progenitors. These data show that DNAme shapes the topography of hematopoietic differentiation, and support a model in which genome-wide methylation changes are transduced to differentiation skews through biases in CpG enrichment of the transcription factor binding motif.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data used in this publication is available at the Gene Expression Omnibus (GEO) database under accession number GSE124822.
References
Ley, T. J. et al. DNMT3A mutations in acute myeloid leukemia. N. Engl. J. Med. 363, 2424–2433 (2010).
Delhommeau, F. et al. Mutation in TET2 in myeloid cancers. N. Engl. J. Med. 360, 2289–2301 (2009).
Gross, S. et al. Cancer-associated metabolite 2-hydroxyglutarate accumulates in acute myelogenous leukemia with isocitrate dehydrogenase 1 and 2 mutations. J. Exp. Med. 207, 339–344 (2010).
Busque, L. et al. Recurrent somatic TET2 mutations in normal elderly individuals with clonal hematopoiesis. Nat. Genet. 44, 1179–1181 (2012).
Abelson, S. et al. Prediction of acute myeloid leukaemia risk in healthy individuals. Nature 559, 400–404 (2018).
Chang, H. H., Hemberg, M., Barahona, M., Ingber, D. E. & Huang, S. Transcriptome-wide noise controls lineage choice in mammalian progenitor cells. Nature 453, 544–547 (2008).
Velten, L. et al. Human haematopoietic stem cell lineage commitment is a continuous process. Nat. Cell Biol. 19, 271–281 (2017).
Graf, T. & Stadtfeld, M. Heterogeneity of embryonic and adult stem cells. Cell Stem Cell 3, 480–483 (2008).
Yu, V. W. C. et al. Epigenetic memory underlies cell-autonomous heterogeneous behavior of hematopoietic stem cells. Cell 168, 944–945 (2017).
Bintu, L. et al. Dynamics of epigenetic regulation at the single-cell level. Science 351, 720–724 (2016).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Domcke, S. et al. Competition between DNA methylation and transcription factors determines binding of NRF1. Nature 528, 575–579 (2015).
Stone, A. et al. DNA methylation of oestrogen-regulated enhancers defines endocrine sensitivity in breast cancer. Nat. Commun. 6, 7758 (2015).
Prendergast, G. C. & Ziff, E. B. Methylation-sensitive sequence-specific DNA binding by the c-Myc basic region. Science 251, 186–189 (1991).
Yin, Y. et al. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science 356, eaaj2239 (2017).
Kribelbauer, J. F. et al. Quantitative analysis of the DNA methylation sensitivity of transcription factor complexes. Cell Rep. 19, 2383–2395 (2017).
Yang, L. et al. DNMT3A loss drives enhancer hypomethylation in FLT3-ITD-associated leukemias. Cancer Cell 30, 363–365 (2016).
Bock, C. et al. DNA methylation dynamics during in vivo differentiation of blood and skin stem cells. Mol. Cell 47, 633–647 (2012).
Ji, H. et al. Comprehensive methylome map of lineage commitment from haematopoietic progenitors. Nature 467, 338–342 (2010).
Xu, W. et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of alpha-ketoglutarate-dependent dioxygenases. Cancer Cell 19, 17–30 (2011).
Abdel-Wahab, O. & Levine, R. L. Mutations in epigenetic modifiers in the pathogenesis and therapy of acute myeloid leukemia. Blood 121, 3563–3572 (2013).
Sperling, A. S., Gibson, C. J. & Ebert, B. L. The genetics of myelodysplastic syndrome: from clonal haematopoiesis to secondary leukaemia. Nat. Rev. Cancer 17, 5–19 (2017).
Steensma, D. P. et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 126, 9–16 (2015).
Jaiswal, S. et al. Clonal hematopoiesis and risk of atherosclerotic cardiovascular disease. N. Engl. J. Med. 377, 111–121 (2017).
Genovese, G., Jaiswal, S., Ebert, B. L. & McCarroll, S. A. Clonal hematopoiesis and blood-cancer risk. N. Engl. J. Med. 372, 1071–1072 (2015).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Couronne, L., Bastard, C. & Bernard, O. A. TET2 and DNMT3A mutations in human T-cell lymphoma. N. Engl. J. Med. 366, 95–96 (2012).
Li, W. et al. DNMT3A mutations and prognostic significance in childhood acute lymphoblastic leukemia. Leuk. Lymphoma 56, 1066–1071 (2015).
Mayle, A. et al. Dnmt3a loss predisposes murine hematopoietic stem cells to malignant transformation. Blood 125, 629–638 (2015).
Kramer, A. C. et al. Dnmt3a regulates T-cell development and suppresses T-ALL transformation. Leukemia 31, 2479–2490 (2017).
Pan, F. et al. Tet2 loss leads to hypermutagenicity in haematopoietic stem/progenitor cells. Nat. Commun. 8, 15102 (2017).
Paul, F. et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell 163, 1663–1677 (2015).
Wilson, N. K. et al. Combined single-cell functional and gene expression analysis resolves heterogeneity within stem cell populations. Cell Stem Cell 16, 712–724 (2015).
Mildner, A. et al. Genomic characterization of murine monocytes reveals C/EBPβ transcription factor dependence of Ly6C− cells. Immunity 46, 849–862.e7 (2017).
Olsson, A. et al. Single-cell analysis of mixed-lineage states leading to a binary cell fate choice. Nature 537, 698–702 (2016).
Yanez, A. et al. Granulocyte-monocyte progenitors and monocyte-dendritic cell progenitors independently produce functionally distinct monocytes. Immunity 47, 890–902.e4 (2017).
Drissen, R. et al. Distinct myeloid progenitor-differentiation pathways identified through single-cell RNA sequencing. Nat. Immunol. 17, 666–676 (2016).
Ward, P. S. et al. The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting alpha-ketoglutarate to 2-hydroxyglutarate. Cancer Cell 17, 225–234 (2010).
Shih, A. H., Abdel-Wahab, O., Patel, J. P. & Levine, R. L. The role of mutations in epigenetic regulators in myeloid malignancies. Nat. Rev. Cancer 12, 599–612 (2012).
Sugiyama, T., Kohara, H., Noda, M. & Nagasawa, T. Maintenance of the hematopoietic stem cell pool by CXCL12-CXCR4 chemokine signaling in bone marrow stromal cell niches. Immunity 25, 977–988 (2006).
Tzeng, Y. S. et al. Loss of Cxcl12/Sdf-1 in adult mice decreases the quiescent state of hematopoietic stem/progenitor cells and alters the pattern of hematopoietic regeneration after myelosuppression. Blood 117, 429–439 (2011).
Hwang, H. S. et al. Enhanced anti-leukemic effects through induction of immunomodulating microenvironment by blocking CXCR4 and PD-L1 in an AML mouse model. Immunol. Invest. 48, 96–105 (2019).
Cho, B. S., Kim, H. J. & Konopleva, M. Targeting the CXCL12/CXCR4 axis in acute myeloid leukemia: from bench to bedside. Korean J. Intern. Med. 32, 248–257 (2017).
Pujato, M., Kieken, F., Skiles, A. A., Tapinos, N. & Fiser, A. Prediction of DNA binding motifs from 3D models of transcription factors; identifying TLX3 regulated genes. Nucleic Acids Res. 42, 13500–13512 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Gaiti, F. et al. Epigenetic evolution and lineage histories of chronic lymphocytic leukaemia. Nature 569, 576–580 (2019).
Kulakovskiy, I. V. et al. HOCOMOCO: towards a complete collection of transcription factor binding models for human and mouse via large-scale ChIP-Seq analysis. Nucleic Acids Res. 46, D252–D259 (2018).
Nam, A. S. et al. Somatic mutations and cell identity linked by Genotyping of Transcriptomes. Nature 571, 355–360 (2019).
Kunimoto, H. et al. Tet2-mutated myeloid progenitors possess aberrant in vitro self-renewal capacity. Blood 123, 2897–2899 (2014).
Verbist, K. C. et al. Metabolic maintenance of cell asymmetry following division in activated T lymphocytes. Nature 532, 389–393 (2016).
Wilson, A. et al. c-Myc controls the balance between hematopoietic stem cell self-renewal and differentiation. Genes Dev. 18, 2747–2763 (2004).
Giladi, A. et al. Single-cell characterization of haematopoietic progenitors and their trajectories in homeostasis and perturbed haematopoiesis. Nat. Cell Biol. 20, 836–846 (2018).
Zhang, X. et al. DNMT3A and TET2 compete and cooperate to repress lineage-specific transcription factors in hematopoietic stem cells. Nat. Genet. 48, 1014–1023 (2016).
Emperle, M. et al. Mutations of R882 change flanking sequence preferences of the DNA methyltransferase DNMT3A and cellular methylation patterns. Nucleic Acids Res. 47, 11355–11367 (2019).
Viner, C. et al. Modeling methyl-sensitive transcription factor motifs with an expanded epigenetic alphabet. Preprint at bioRxiv https://doi.org/10.1101/043794 (2016).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Tetteh, P. W. et al. Replacement of lost Lgr5-positive stem cells through plasticity of their enterocyte-lineage daughters. Cell Stem Cell 18, 203–213 (2016).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Lhoumaud, P. et al. EpiMethylTag: simultaneous detection of ATAC-seq or ChIP-seq signals with DNA methylation. Genome Biol. 20, 248 (2019).
Liu, T. Use Model-based Analysis of ChIP-Seq (MACS) to analyze short reads generated by sequencing protein-DNA interactions in embryonic stem cells. Methods Mol. Biol. 1150, 81–95 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Akalin, A. et al. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 13, R87 (2012).
Yoshida, H. et al. The cis-regulatory atlas of the mouse immune system. Cell 176, 897–912.e20 (2019).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Moran-Crusio, K. et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell 20, 11–24 (2011).
Nguyen, S., Meletis, K., Fu, D., Jhaveri, S. & Jaenisch, R. Ablation of de novo DNA methyltransferase Dnmt3a in the nervous system leads to neuromuscular defects and shortened lifespan. Dev. Dyn. 236, 1663–1676 (2007).
Shih, A. H. et al. Combination targeted therapy to disrupt aberrant oncogenic signaling and reverse epigenetic dysfunction in IDH2- and TET2-mutant acute myeloid lLeukemia. Cancer Discov. 7, 494–505 (2017).
Kuhn, R., Schwenk, F., Aguet, M. & Rajewsky, K. Inducible gene targeting in mice. Science 269, 1427–1429 (1995).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Hafemeister, C. S. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Han, X. et al. Mapping the Mouse Cell Atlas by Microwell-Seq. Cell 172, 1091–1107.e17 (2018).
Sun, H., Zhou, Y., Fei, L., Chen, H. & Guo, G. scMCA: a tool to define mouse cell types based on single-cell digital expression. Methods Mol. Biol. 1935, 91–96 (2019).
Bolker, B. M. et al. Generalized linear mixed models: a practical guide for ecology and evolution. Trends Ecol. Evol. 24, 127–135 (2009).
Martin, J. C. et al. Single-cell analysis of Crohn’s disease lesions identifies a pathogenic cellular module associated with resistance to anti-TNF therapy. Cell 178, 1493–1508.e20 (2019).
Orlanski, S. et al. Tissue-specific DNA demethylation is required for proper B-cell differentiation and function. Proc. Natl Acad. Sci. USA 113, 5018–5023 (2016).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Macaulay, I. C. et al. G&T–seq: parallel sequencing of single-cell genomes and transcriptomes. Nat. Methods 12, 519–522 (2015).
Picelli, S. et al. Full-length RNA–seq from single cells using Smart–seq2. Nat. Protoc. 9, 171–181 (2014).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359 (2012).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res 22, 1760–1774 (2012).
Acknowledgements
R.C. is supported by Lymphoma Research Foundation and Marie Skłodowska-Curie fellowships. S.C.L. is supported by the ASH Scholar Award, the Edward P. Evans Foundation Young Investigator Award and the NIH K99/R00 Pathway to Independence Award (no. NCI R00 CA218896). P.L. is supported by the AACR Takeda Multiple Myeloma Fellowship and the NCC National Cancer Center Postdoctoral fellowship. O.A.-W. is supported by grants from NIH/NHLBI (no. R01 HL128239), the Henry & Marilyn Taub Foundation, the Edward P. Evans Foundation and the Leukemia and Lymphoma Society. This work was also funded by grant no. P30 CA008748. J.A.S. is supported by National Institute of General Medical Sciences grant no. 1R35GM122515. A.D.V. is supported by a National Cancer Institute career development grant no. K08 CA215317, the William Raveis Charitable Fund Fellowship of the Damon Runyon Cancer Research Foundation (grant no. DRG 117-15) and an Evans MDS Young Investigator grant from the Edward P. Evans Foundation. D.A.L. is supported by the Burroughs Wellcome Fund Career Award for Medical Scientists, the Pershing Square Sohn Prize for Young Investigators in Cancer Research and the National Institutes of Health Director’s New Innovator Award (no. DP2-CA239065). This work was enabled by the Weill Cornell Epigenomics Core and Flow Cytometry Core. This work was also supported by the Leukemia and Lymphoma Society Translational Research Program, National Heart Lung and Blood Institute (grant no. R01HL145283-01) and Stand Up To Cancer Innovative Research Grant (no. SU2C-AACR-IRG-0616).
Author information
Authors and Affiliations
Contributions
F.I., S.C.L., E.K., O.A.-W. and D.A.L. devised the research strategy. F.I., S.C.L., A.P., R.C., C.A., M.K., P.W.J., A.D.V. and A.S.N. performed the experiments. F.I., S.C.L., A.P., F.G., B.G., R.R.M., S.D.D., K.-T.K., S.K.-H. and R.C.S. performed the analyses. F.I., P.W.J., P.L., J.A.S. and D.A.L. developed the ATAC-seq plus bisulfite conversion (ATAC-Bseq) method. F.I. and D.A.L. wrote the manuscript. R.L.L., E.K., O.A.-W. and D.A.L. helped to interpret results. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
O.A.-W. has served as a consultant for H3B Biomedicine, Foundation Medicine Inc., Merck and Janssen, and is on the Scientific Advisory Board of Envisagenics Inc. O.A.-W. has received prior research funding from H3B Biomedicine unrelated to the current manuscript. R.L.L. is on the supervisory board of Qiagen and is a scientific advisor to Loxo (until 2/2019), Imago, C4 Therapeutics and Isoplexis, which each include an equity interest. He receives research support from and consulted for Celgene and Roche, he has received research support from Prelude Therapeutics and he has consulted for Lilly, Incyte, Novartis, Astellas, Morphosys and Janssen. He has received honoraria from Lilly and Amgen for invited lectures and from Gilead for grant reviews.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
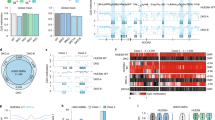
Extended Data Fig. 1 Chromium 10x data summary.
a, Summary of Chromium 10x data (pIpC = polyinosinic-polycytadylic acid). b, Number of genes detected as a function of the number of unique molecular identifiers (UMIs) per cell barcode. Red dots = cell barcodes with mitochondrial content > 20%; blue dots = cell barcode with lower than expected complexity (lower than two standard deviations from linear fit); dashed red line = linear fit. c, Percentage of cell barcodes removed per sample after filtering low complexity barcodes and barcodes with mitochondrial UMIs > 20%. d, Quality control of scRNA-seq (n = 13 biological independent animals) after filtering. e, PCR validation of Tet2 exon 3 deletion 4 weeks after pIpC administration. Genomic DNA was isolated from Lin− bone marrow cells and amplified using the primers Tet2-F1, Tet2-R1 or Tet2-R-Lox, (Supplementary Table 5). One representative example of n = 3 independent experiments is shown. f, PCR validation of Dnmt3a exon 17 and 18 deletion 4 weeks after pIpC administration. Genomic DNA was isolated from Lin− bone marrow cells and amplified using the primers Dnmt3a-F1, Dnmt3a-R1 or Dnmt3a-R-Lox, shown in Supplementary Table 5. One representative example of n = 3 independent experiments is shown. g, Uniform Manifold Approximation and Projection (UMAP) dimensionality reduction showing joint embedding of WT (17,702 cells; n = 7 mice), Tet2 KO (18,651 cells; n = 7 mice), Dnmt3a KO (13,858 cells, n = 4 mice) and Idh2-R140Q (9,883 cells, n = 3 mice).
Extended Data Fig. 2 Drop-seq data summary.
a, Summary of Drop-seq data showing PCR pool, genotype, sorting strategy, time after recombination (n = 14 biologically independent animals) and number of cells captured after filtering (pIpC = polyinosinic-polycytadylic acid). b, Number of unique molecular identifiers (UMIs) and genes detected per cell barcode per sample. c, Overview of number of genes detected as a function of the number of UMIs per cell barcode. Red dots = cell barcodes with mitochondrial content > 20%; blue dots = cell barcode with lower than expected complexity (lower than two standard deviations from linear fit); dashed red line = linear fit. d, Percentage of cell barcodes removed per sample (n = 14 biologically independent animals) after filtering out low complexity barcodes and barcodes with mitochondrial UMIs > 20%. e, Percentage of mitochondrial UMIs per cell per sample (n = 14 biologically independent animals) after filtering.
Extended Data Fig. 3 Quality control of joint embedding across single cell technologies.
a, Left panel: Uniform Manifold Approximation and Projection (UMAP) dimensionality reduction showing joint embedding of WT (17,702 cells; n = 7 mice), Tet2 KO (18,651 cells; n = 7 mice), Dnmt3a KO (13,858 cells, n = 4 mice) and Idh2-R140Q (9,883 cells, n = 3 mice) lineage-negative hematopoietic progenitors. Right panels: UMAP embedding obtained for each scRNA-seq method is shown separately. b, Gene expression correlation between cells obtained by different scRNA-seq methods (Chromium v2, Chromium v3 and Drop-seq) that were mapped to the same cell cluster. The gene expression frequency was calculated as the number of unique molecular identifiers (UMIs) mapping to a given gene relative to the total number of UMIs detected for a given cluster, and multiplied by a factor of 105. The log2 of the pseudo-bulk gene expression is shown (R2 values were obtained from Pearson correlation; red dots highlight the top gene markers for each cluster). c, Gene expression correlation between WT cells and expression profiles from the Mouse Cell Atlas66 dataset, as obtained by scMCA67 (see methods).
Extended Data Fig. 4 Cluster annotation, supporting evidence for HSC self-renewal and Lineage-negative, c-Kit+ cells validation in Tet2 KO.
a, Differentially expressed genes for WT cluster HSC-1 (492 cells), HSC-2 (288 cells) or HSC-3 (384 cells), relative to the remaining HSC clusters from Chromium data (n = 4 mice; logistic regression with Bonferroni correction; FDR < 0.05). b, Drop-seq data for Lin−, c-Kit positive cells for WT (2,986 cells, n = 2 mice) or Tet2 KO (1,425 cells, n = 2 mice) progenitors 4 weeks after recombination (HSCs = Hematopoietic stem cells; IMP = Immature myeloid progenitors; MD = Monocytic-dendritic progenitors; NP = Neutrophil progenitors; EP = Erythroid progenitors; MkP = Megakaryocyte progenitors; CLP = Common lymphoid progenitors; Ba = Basophil progenitors; Eo = Eosinophil progenitors; B-cellP = B-cell progenitors; T-cellP = T-cell progenitors). c, Frequency changes for HSCs, MDs and EPs 4 weeks after recombination (Permutation test on 1,425 randomly sampled cells from each genotype, with 105 iterations). d, Quiescence score per cell cycle category (above/below median) in WT HSCs (n = 1,982 cells; two-sided Wilcoxon rank sum test). e, Flow cytometry of cell cycle in LT-HSCs as measured by Mki67 expression for WT (n = 4 mice) or Tet2 KO (n = 3 mice) 4 weeks after recombination (two-sided Student t-test). f, Serial re-plating colony-formation assays for WT (n = 11) and Tet2 KO (n = 7) Lin−, c-Kit+ bone marrow hematopoietic (CFU = colony-formation unit; dots represent the mean; error bars represent standard deviation; two-sided Students t-test). g, Differentially expressed genes per WT cluster Mono-1 (n = 344 cells), Mono-2 (n = 345 cells) or Mono-3 (n = 284 cells), relative to the remaining monocyte clusters. Differentially expressed genes were defined from Chromium data (n = 4 mice; logistic regression with Bonferroni correction; FDR < 0.05). h, Expression of Ly6c2 and H2-Ab1 in WT Mono-1 (n = 344 cells), Mono-2 (n = 345 cells) and Mono-3 (n = 284 cells) clusters (logistic regression with Bonferroni correction).
Extended Data Fig. 5 Flow cytometry validation, peripheral blood counts, in vitro colony-forming assay and 20 weeks post pIpC validations of Tet2 KO frequency changes.
a, Frequency changes for Lin− Tet2 KO (18,651 cells, n = 7 mice) relative to WT (17,702 cells, n = 7 mice) 4 weeks after recombination. Red dots indicate significant frequency changes; red error bars represent standard deviation; dashed line indicates WT reference frequencies; grey shadow region indicates ± standard deviation (LMM followed by ANOVA; * P < 0.05; ** P < 0.01; *** P < 0.001). b, Flow cytometry for WT (n = 15) and Tet2 KO (n = 23) mice 4 weeks after recombination (two-sided Students t-test; bars represent the mean value, error bars represent the standard deviation; LT-HSC = long-term hematopoietic stem cell; MPP = Multi-potent progenitor; HPC = Hematopoietic progenitor cell; CMP = common myeloid progenitor; GMP = Granulocyte-monocyte progenitor; MEP = megakaryocyte-erythrocyte progenitor). c, Peripheral blood cell counts from WT (n = 10) or Tet2 KO (n = 10) mice, either 4 or 20 weeks after Cre-mediated recombination (two-sided Students t-test; bars represent the mean and error bars represent the standard deviation. Each dot represents a mouse replicate; RBC = red blood cells; MCV = mean corpuscular volume). d, Erythroid colony-forming assay for WT (n = 4) or Tet2 KO (n = 4) mice, 4 weeks after recombination (two-sided Student t-test; bars represent the mean number of colonies for each genotype; error bars represent standard deviation). e, Drop-seq data showing 2,478 randomly sampled cells from Lin− cells for WT (2,757 cells) or Tet2 KO (2,875 cells) 20 weeks after recombination (HSCs = Hematopoietic stem cells; IMP = Immature myeloid progenitors; MD = Monocytic-dendritic progenitors; NP = Neutrophil progenitors; EP = Erythroid progenitors; MkP = Megakaryocyte progenitors; CLP = Common lymphoid progenitors; Ba = Basophil progenitors; Eo = Eosinophil progenitors; B-cellP = B-cell progenitors; T-cellP = T-cell progenitors). f, Frequency changes for monocyte (Mono 1–3) and erythroid (Ery 1–3) progenitor clusters (permutation test).
Extended Data Fig. 6 Validation of cell cluster frequency changes in Idh2-R140Q mutant mice and Dnmt3a KO mice.
a, Frequency changes for Lin− Idh2-R140Q (n = 3) relative to WT (n = 6) mice 4 weeks post-recombination (linear mixed model (LMM) followed by ANOVA; *P < 0.05; ***P < 0.01). b, E/B and Ery 1–3 frequencies 4 weeks after recombination for WT (n = 6) and Idh2-R140Q (n = 3) mice. Error bars represent standard error of the mean (SEM; LMM followed by ANOVA). c, Ratio between erythroid (E/B, Ery-1 and ERy-2) and monocytic (IMP-1 and Mono-1) clusters for WT (n = 6) and Idh2-R140Q (n = 3) mice 4 weeks post-recombination. Error bars indicate SEM (LMM followed by ANOVA). d, Flow cytometry of hematopoietic progenitors from WT (n = 10) and Idh2-R140Q (n = 8) mice 4 weeks post-recombination (two-sided Students t-test; LT-HSC = long-term hematopoietic stem cell; MPP = Multi-potent progenitor; HPC = Hematopoietic progenitor cell; CMP = common myeloid progenitor; GMP = Granulocyte-monocyte progenitor; MEP = megakaryocyte-erythrocyte progenitor). e, Peripheral blood monocytes for WT (n = 22) or Idh2-R140Q (n = 8) mice 4 weeks post-recombination (two-sided Students t-test). f, Differential gene expression between WT (n = 2,150 cells) and Idh2-R140Q (n = 1,184 cells) HSC 1–3 clusters. Red dots represent differentially expressed genes (permutation test followed by Benjamini-Hochberg (BH) correction, P < 0.05 and absolute log2 fold change > 1). g, Frequencies for Lin− Dnmt3a KO (n = 4) relative to WT (n = 4) mice, 4 weeks post-recombination (LMM followed by ANOVA; *P < 0.05; ***P < 0.001). h, Flow cytometry of WT (n = 5) and Dnmt3a KO (n = 8) mice 4 weeks post-recombination (two-sided Students t-test). i, Peripheral blood measurements for WT (n = 8) or Dnmt3a KO (n = 8) mice 4 weeks post-recombination (two-sided Students t-test; RBC = red blood cell; MCV = mean corpuscular volume). j, Frequency changes in HSCs (Hlf+), erythroid (Car1+) and monocyte (Ly6c2+; Irf8+) progenitors for WT (n = 6), Tet2 KO (n = 6) and Dnmt3a KO (n = 4) mice clustered independently for each technology. For bar plots, bars represent mean values, dots represent mouse replicates and error bars represent standard deviation unless indicated otherwise. For radar plots, red dots indicate significant frequency changes; red error bars represent standard deviation; dashed line indicates WT reference frequencies and shadow region indicates ± standard deviation.
Extended Data Fig. 7 Gene module analysis.
a, Schematic representation of the process for gene module identification. b, Correlation between gene module scores in HSC clusters (HSC 1–3), as calculated by the number of unique molecular identifiers (UMIs) mapping to the genes from each module per 10,000 total UMIs in the cell (Pearson correlation). c, Transcriptional priming values per biological replicate for Tet2 KO (n = 2,989 cells; n = 7 mice), WT (n = 2,150 cells; n = 7 mice) and Dnmt3a KO (n = 1,325 cells; n = 4 mice). Dots represent the mean value; error bars show the 95% confidence interval.
Extended Data Fig. 8 Mean CpG frequencies per base of erythroid and monocytic transcription factor binding motifs.
a, Schematic representation of the process for mean CpG frequency per base calculation for transcription factor binding motif position weight matrix. b, Scatter plot showing the correlation between the ratio of transcription factor regulon68 activity change between Tet2 KO (n = 7 mice) and WT (n = 7 mice), as calculated by the total number of molecules mapping to the genes comprising the regulons for the HSC 1–3 clusters per 10,000 UMIs in the cluster, and the product of the CpG frequency in the transcription factor motif and enrichment score as determined by SELEX15 (two-sided Students t-test). c, Mean CpG frequency per base differences between erythroid- and monocytic-associated transcription factors according to different thresholds used for expression change between clusters (n = 7 biologically independent animals; two-sided Wilcoxon rank sum test; FC = fold change). d, Examples of motif CpG content and methylation for Klf1 and Spi1 transcription factors as obtained from the MethMotif database69.
Extended Data Fig. 9 Mean CpG frequency per base correlates with methylation of motifs at accessible enhancer regions.
a, Gating for cell sorting for ATAC-Bseq experiments (LSK = lineage negative; Sca1 positive; c-Kit positive). b, Correlation between biological replicates for ATAC-Bseq experiments. Reads were downsampled to 30 ×106 reads per sample and the average read count per 10 kbp genomic windows was calculated (Pearson correlation). c, Examples of Homer output for de novo motif enrichment for either erythroid- or myelo-monocytic-associated accessible peaks within 10 kb of the closest transcriptional start site. d, Correlation between mean CpG frequency per base and the number of differentially (FDR<0.25, absolute methylation difference > 5%) hyper- or hypo-methylated CpGs between WT and Tet2 KO (n = 104,829 CpG sites) or Dnmt3a KO (250,353 CpG sites) respectively, per 100 motifs at accessible enhancers (upper panel) or accessible promoters (two-sided Students t-test; bottom panel; Spearman correlation). e, Number of hypermethylated CpGs per 10,000 motifs for erythroid- or monocyte-associated transcription factor motifs. 100 iterations of sampling without replacement were performed, sampling 10,000 motif sites each iteration, and measuring the number of differentially (FDR<0.25, absolute methylation difference > 5%) hypermethylated or hypomethylated sites captured in Tet2 KO (n = 2 mice) and Dnmt3a KO (n = 2 mice), respectively (two-sided Students t-test). f, Correlation between the percentage of hyper- or hypo-methylated CpGs between WT (n = 2 mice) and Tet2 KO (n = 2 mice) or Dnmt3a KO (n = 2 mice), respectively from total CpGs captured for each transcription factor DNA binding motif site and the mean CpG frequency per base, for motifs in accessible enhancers (middle panel) or accessible promoters (bottom panel; Spearman correlation; two-sided Students t-test).
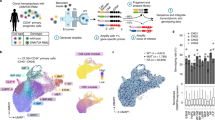
Extended Data Fig. 10 Single cell RNA and methylation reveals increased heterogeneity and links enhancer methylation with transcriptional priming.
a, LT-HSCs cell cycle scores for WT (n = 178 cells), Tet2 KO (n = 182 cells) and Dnmt3a KO (N =50 cells) as calculated by the number of UMIs mapping to the gene set per 10,000 total UMIs for each of the mapped clusters (two-sided Wilcoxon rank sum test). b, Single cell methylation percentage of CpG islands (CpGi), exon, intron and promoter regions for WT (n = 178 cells), Tet2 KO (n = 182 cells) or Dnmt3a KO (n = 50 cells) LT-HSCs. CpGi were robust to Tet2 deletion−induced hypermethylation, as previously reported70,71. c, Correlation between erythroid-to-monocytic transcriptional priming and mean enhancer methylation in WT (n = 178), Tet2 KO (n = 182) and Dnmt3a KO (n = 50) LT-HSCs (Spearman correlation; two-sided Students t-test). d, Average single cell enhancer methylation comparison between erythroid (n = 151 cells) or monocytic (n = 166 cells) primed LT-HSCs across genotypes (two-sided Wilcoxon rank sum test). e, CD34+ hematopoietic bone marrow progenitors from normal7 (n = 1,035 cells) or DNMT3A-F755S mutant affected (n = 7,338 cells) subjects. f, Clusters for the clonal hematopoiesis sample (HSC = hematopoietic stem cell; IMP = immature myeloid progenitor; Neu = neutrophil/granulocyte progenitor; Ery = erythroid progenitor; M/D = monocyte-dendritic progenitor; CLP = common lymphoid progenitor; MkP = megakaryocyte progenitor; cc = high cell cycle cluster; mt = high mitochondrial gene expression cluster). g, Differentially expressed genes per cluster (FDR < 0.05; logistic regression with Bonferroni correction; Supplementary Table 2) per cluster are shown. h, Gene marker expression from erythroid (GATA1, CA1), monocyte (IRF8, LGALS1), megakaryocyte (PF4, PLEK) and neutrophil (MPO, ELANE) cells. i, Frequency of GATA1+ cells for normal (n = 1,035 cells) and DNMT3A-F755S (n = 7,338 cells) clonal hematopoiesis subject. Cells were defined as positive when at least one UMI was detected for GATA1 (two-sided Fisher exact test).
Supplementary information
Supplementary Information
Supplementary Note and Fig. 1
Supplementary Tables
Supplementary Tables 1–6
Rights and permissions
About this article
Cite this article
Izzo, F., Lee, S.C., Poran, A. et al. DNA methylation disruption reshapes the hematopoietic differentiation landscape. Nat Genet 52, 378–387 (2020). https://doi.org/10.1038/s41588-020-0595-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0595-4
This article is cited by
-
Genetic variation across and within individuals
Nature Reviews Genetics (2024)
-
Increased iron uptake by splenic hematopoietic stem cells promotes TET2-dependent erythroid regeneration
Nature Communications (2024)
-
Clonal hematopoiesis and its impact on the aging osteo-hematopoietic niche
Leukemia (2024)
-
Cross-tissue patterns of DNA hypomethylation reveal genetically distinct histories of cell development
BMC Genomics (2023)
-
TET2 inhibits the proliferation and metastasis of lung adenocarcinoma cells via activation of the cGAS-STING signalling pathway
BMC Cancer (2023)