Abstract
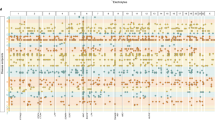
Genetic risk factors frequently affect multiple common human diseases, providing insight into shared pathophysiological pathways and opportunities for therapeutic development. However, systematic identification of genetic profiles of disease risk is limited by the availability of both comprehensive clinical data on population-scale cohorts and the lack of suitable statistical methodology that can handle the scale of and differential power inherent in multi-phenotype data. Here, we develop a disease-agnostic approach to cluster the genetic risk profiles for 3,025 genome-wide independent loci across 19,155 disease classification codes from 320,644 participants in the UK Biobank, representing a large and heterogeneous population. We identify 339 distinct disease association profiles and use multiple approaches to link clusters to the underlying biological pathways. We show how clusters can decompose the variance and covariance in risk for disease, thereby identifying underlying biological processes and their impact. We demonstrate the use of clusters in defining disease relationships and their potential in informing therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data shown in this paper are available at https://www.treewas.org/.
Code availability
The code for the TreeWAS analysis is available at https://github.com/mcveanlab/TreeWASDir.
References
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Pickrell, J. K. et al. Detection and interpretation of shared genetic influences on 42 human traits. Nat. Genet. 48, 709–717 (2016).
Malik, R. et al. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat. Genet. 50, 524–537 (2018).
Warren, H. R. et al. Genome-wide association analysis identifies novel blood pressure loci and offers biological insights into cardiovascular risk. Nat. Genet. 49, 403–415 (2017).
Lee, S. H. et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013).
Ellinghaus, D. et al. Analysis of five chronic inflammatory diseases identifies 27 new associations and highlights disease-specific patterns at shared loci. Nat. Genet. 48, 510–518 (2016).
Parkes, M., Cortes, A., van Heel, D. A. & Brown, M. A. Genetic insights into common pathways and complex relationships among immune-mediated diseases. Nat. Rev. Genet. 14, 661–673 (2013).
Inshaw, J. R. J., Cutler, A. J., Burren, O. S., Stefana, M. I. & Todd, J. A. Approaches and advances in the genetic causes of autoimmune disease and their implications. Nat. Immunol. 19, 674–684 (2018).
Cortes, A. et al. Bayesian analysis of genetic association across tree-structured routine healthcare data in the UK Biobank. Nat. Genet. 49, 1311–1318 (2017).
Oprea, T. I. et al. Unexplored therapeutic opportunities in the human genome. Nat. Rev. Drug Discov. 17, 317–332 (2018).
Dendrou, C. A. et al. Resolving TYK2 locus genotype-to-phenotype differences in autoimmunity. Sci. Transl. Med. 8, 363ra149 (2016).
Beecham, A. H. et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat. Genet. 45, 1353–1360 (2013).
Cortes, A. et al. Identification of multiple risk variants for ankylosing spondylitis through high-density genotyping of immune-related loci. Nat. Genet. 45, 730–738 (2013).
Timpson, N. J., Greenwood, C. M. T., Soranzo, N., Lawson, D. J. & Richards, J. B. Genetic architecture: the shape of the genetic contribution to human traits and disease. Nat. Rev. Genet. 19, 110–124 (2018).
Solovieff, N., Cotsapas, C., Lee, P. H., Purcell, S. M. & Smoller, J. W. Pleiotropy in complex traits: challenges and strategies. Nat. Rev. Genet. 14, 483–495 (2013).
Udler, M. S. et al. Type 2 diabetes genetic loci informed by multi-trait associations point to disease mechanisms and subtypes: a soft clustering analysis. PLoS Med. 15, e1002654 (2018).
Sanseau, P. et al. Use of genome-wide association studies for drug repositioning. Nat. Biotechnol. 30, 317–320 (2012).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
MacArthur, J. et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 45, D896–D901 (2017).
Raychaudhuri, S. et al. Five amino acids in three HLA proteins explain most of the association between MHC and seropositive rheumatoid arthritis. Nat. Genet. 44, 291–296 (2012).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Deloukas, P. et al. Large-scale association analysis identifies new risk loci for coronary artery disease. Nat. Genet. 45, 25–33 (2013).
Willer, C. J. et al. Discovery and refinement of loci associated with lipid levels. Nat. Genet. 45, 1274–1283 (2013).
Li, Y. et al. Genetic variants associated with deep vein thrombosis: the F11 locus. J. Thromb. Haemost. 7, 1802–1808 (2009).
Bertina, R. M. et al. Mutation in blood coagulation factor V associated with resistance to activated protein C. Nature 369, 64–67 (1994).
Klarin, D. et al. Genetic analysis of venous thromboembolism in UK Biobank identifies the ZFPM2 locus and implicates obesity as a causal risk factor. Circ. Cardiovasc. Genet. 10, e001643 (2017).
Gerhardt, A. et al. Prothrombin and factor V mutations in women with a history of thrombosis during pregnancy and the puerperium. N. Engl. J. Med. 342, 374–380 (2000).
Clarke, R. et al. Genetic variants associated with Lp(a) lipoprotein level and coronary disease. N. Engl. J. Med. 361, 2518–2528 (2009).
Thanassoulis, G. et al. Genetic associations with valvular calcification and aortic stenosis. N. Engl. J. Med. 368, 503–512 (2013).
McPherson, R. et al. A common allele on chromosome 9 associated with coronary heart disease. Science 316, 1488–1491 (2007).
Zhao, W. et al. Identification of new susceptibility loci for type 2 diabetes and shared etiological pathways with coronary heart disease. Nat. Genet. 49, 1450–1457 (2017).
Abifadel, M. et al. Mutations in PCSK9 cause autosomal dominant hypercholesterolemia. Nat. Genet. 34, 154–156 (2003).
Lewontin, R. C. The interaction of selection and linkage. I. General considerations; heterotic models. Genetics 49, 49–67 (1964).
Frot, B., Jostins, L. & McVean, G. Graphical model selection for Gaussian conditional random fields in the presence of latent variables. J. Am. Stat. Assoc. 114, 723–734 (2018).
Evangelou, E. et al. Genetic analysis of over 1 million people identifies 535 new loci associated with blood pressure traits. Nat. Genet. 50, 1412–1425 (2018).
Davey Smith, G. & Hemani, G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum. Mol. Genet. 23, R89–R98 (2014).
Trochet, H. et al. Bayesian meta-analysis across genome-wide association studies of diverse phenotypes. Genet. Epidemiol. 43, 532–547 (2019).
Giambartolomei, C. et al. A Bayesian framework for multiple trait colocalization from summary association statistics. Bioinformatics 34, 2538–2545 (2018).
Stephens, M. A unified framework for association analysis with multiple related phenotypes. PLoS ONE 8, e65245 (2013).
Richardson, T. G., Harrison, S., Hemani, G. & Davey Smith, G. An atlas of polygenic risk score associations to highlight putative causal relationships across the human phenome. eLife 8, e43657 (2019).
Ding, L. et al. Modeling of multivariate longitudinal phenotypes in family genetic studies with Bayesian multiplicity adjustment. BMC Proc. 8, S69 (2014).
Wain, L. V. et al. Novel insights into the genetics of smoking behaviour, lung function, and chronic obstructive pulmonary disease (UK BiLEVE): a genetic association study in UK Biobank. Lancet Respir. Med. 3, 769–781 (2015).
Acknowledgements
This research has been conducted using the UK Biobank Resource (application no. 10625). This work uses data provided by patients and collected by the National Health Service (NHS) as part of their care and support. Computation used the Biomedical Research Computing facility, a joint development between the Wellcome Centre for Human Genetics and the Big Data Institute supported by the National Institute for Health Research (NIHR) Oxford Biomedical Research Centre (BRC). The views expressed are those of the author(s) and not necessarily those of the NHS, NIHR or the Department of Health. This research has been conducted with the support of the Wellcome Trust (grant nos. 100956/Z/13/Z and 090532/Z/09/Z to G.M. and grant no. 100308/Z/12/Z to L.F.). L.F. was also supported by the Danish National Research Foundation, Takeda, the Medical Research Council (grant no. MC_UU_12010/3), the Oak Foundation (grant no. OCAY-15-520) and the NIHR Oxford BRC. C.A.D. was supported by the Wellcome Trust/Royal Society (grant no. 204290/Z/16/Z). G.M. was supported by the Li Ka Shing Foundation.
Author information
Authors and Affiliations
Contributions
A.C. and G.M. performed the analyses with contributions from C.A.D. and L.F. A.C., L.F. and G.M. conceived the study. A.C., C.A.D., L.F. and G.M. wrote the manuscript. P.K.A. designed and created the website https://www.treewas.org/ and prepared the manuscript figures.
Corresponding author
Ethics declarations
Competing interests
G.M. is a director of and shareholder in GENOMICS plc. He is also a partner in Peptide Groove LLP. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Comparison of estimated log10(BFtree) in the two implementations of TreeWAS for 25,000 SNPs in the hospital episode statistics data set.
Pearson correlation between the two analysis is noted in the text.
Extended Data Fig. 2 Derivation of an allele frequency-specific log10(BFtree) significance threshold to maintain a false positive rate below 1%.
The threshold for each allele frequency bin was set to be at least log10(BFtree) = 5.
Extended Data Fig. 3 Concordance of TreeWAS analysis results in the two sources of phenotype data from the UK Biobank, self-reported (SR) data-field 20002 and hospitalisation in-patient records (HES) data-fields 41142 and 41078.
We observed high concordance of the observed evidence of association (log10(BFtree)) for 3,025 independent SNPs and 25,640 GWAS catalog SNPs, with Pearson’s correlation of 0.87 and 0.56, respectively.
Extended Data Fig. 4 Hierarchical clustering of 3,025 SNP risk profiles across the ICD-10 classification tree in the UK Biobank HES data set.
Y-axis is the distance between pairs. Blue line is at height value 0 and red line at height value -5.
Extended Data Fig. 5 Estimates of relationship between the genetic risk profiles for 339 clusters.
For all pairwise comparisons we computed the |D’| statistic and the Jaccard index (see Section Disease ontology analyses in the Supplementary Note).
Extended Data Fig. 6 Schematic illustration of the model that is used to motivate the focal phenotype analysis.
We hypothesize that a set of variants, G, that influences risk for a common set of disease phenotypes, Z, can be acting through a single underlying biological process, X. Typically, we are unlikely to have direct measurement of this variable, though of those disease codes that are mediated by this latent variable, some are likely to be closer to it than others, where closer means a larger absolute value for the regression coefficient of the latent variable on the observed outcome (See Supplementary Note).
Extended Data Fig. 7 Principal component analysis of genome-wide genotype data in the UK Biobank cohort.
Each plot corresponds to a projection into two dimensions of the principal component analysis. Individuals in blue were determined to be of recent and genome-wide British Isles ancestry.
Supplementary information
Rights and permissions
About this article
Cite this article
Cortes, A., Albers, P.K., Dendrou, C.A. et al. Identifying cross-disease components of genetic risk across hospital data in the UK Biobank. Nat Genet 52, 126–134 (2020). https://doi.org/10.1038/s41588-019-0550-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0550-4
This article is cited by
-
Identifying genetic subtypes of disease from hospital diagnosis records
Nature Genetics (2023)
-
Age-dependent topic modeling of comorbidities in UK Biobank identifies disease subtypes with differential genetic risk
Nature Genetics (2023)
-
Inflammatory human leucocyte antigen genotypes are not a risk factor in chronic subdural hematoma development
Acta Neurochirurgica (2023)
-
A tissue-level phenome-wide network map of colocalized genes and phenotypes in the UK Biobank
Communications Biology (2022)
-
Identification of putative genetic variants in major depressive disorder patients in Pakistan
Molecular Biology Reports (2022)