Abstract
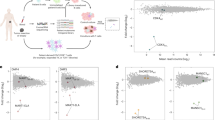
Somatic mutations can result in the formation of neoantigens, immunogenic peptides that are presented on the tumor cell surface by HLA molecules. These mutations are expected to be under negative selection pressure, but the extent of the resulting neoantigen depletion remains unclear. On the basis of HLA affinity predictions, we annotated the human genome for its translatability to HLA binding peptides and screened for reduced single nucleotide substitution rates in large genomic data sets from untreated cancers. Apparent neoantigen depletion signals become negligible when taking into consideration trinucleotide-based mutational signatures, owing to lack of power or to efficient immune evasion mechanisms that are active early during tumor evolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
This study is based on public data (open or controlled access) from The Cancer Genome Atlas Network. Downstream data and source code are available at zenodo (https://doi.org/10.5281/zenodo.2621365 and https://doi.org/10.5281/zenodo.3461642, respectively).
References
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Cancer Genome Atlas Research Network et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Dunn, G. P., Old, L. J. & Schreiber, R. D. The three Es of cancer immunoediting. Annu. Rev. Immunol. 22, 329–360 (2004).
DuPage, M., Mazumdar, C., Schmidt, L. M., Cheung, A. F. & Jacks, T. Expression of tumour-specific antigens underlies cancer immunoediting. Nature 482, 405–409 (2012).
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012).
Hodi, F. S. et al. Improved survival with ipilimumab in patients with metastatic melanoma. N. Engl. J. Med. 363, 711–723 (2010).
Sharma, P. & Allison, J. P. Immune checkpoint targeting in cancer therapy: toward combination strategies with curative potential. Cell 161, 205–214 (2015).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Shukla, S. A. et al. Comprehensive analysis of cancer-associated somatic mutations in class I HLA genes. Nat. Biotechnol. 33, 1152–1158 (2015).
McGranahan, N. et al. Allele-specific HLA loss and immune escape in lung cancer evolution. Cell 171, 1259–1271.e11 (2017).
Davoli, T., Uno, H., Wooten, E. C. & Elledge, S. J. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science 355, eaaf8399 (2017).
Rutledge, W. C. et al. Tumor-infiltrating lymphocytes in glioblastoma are associated with specific genomic alterations and related to transcriptional class. Clin. Cancer Res. 19, 4951–4960 (2013).
Brown, S. D. et al. Neo-antigens predicted by tumor genome meta-analysis correlate with increased patient survival. Genome Res. 24, 743–750 (2014).
Rosenthal, R. et al. Neoantigen-directed immune escape in lung cancer evolution. Nature 567, 479–485 (2019).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Zapata, L. et al. Negative selection in tumor genome evolution acts on essential cellular functions and the immunopeptidome. Genome Biol. 19, 67 (2018).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949.e15 (2017).
Van den Eynden, J. & Larsson, E. Mutational signatures are critical for proper estimation of purifying selection pressures in cancer somatic mutation data when using the dN/dS metric. Front. Genet. 8, 74 (2017).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041.e21 (2017).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Marty, R. et al. MHC-I genotype restricts the oncogenic mutational landscape. Cell 17, 1272–1283.e15 (2017).
Paul, S. et al. HLA class I alleles are associated with peptide-binding repertoires of different size, affinity, and immunogenicity. J. Immunol. 191, 5831–5839 (2013).
Chowell, D. et al. TCR contact residue hydrophobicity is a hallmark of immunogenic CD8+ T cell epitopes. Proc. Natl Acad. Sci. USA 112, E1754–E1762 (2015).
Ellrott, K. et al. Scalable open science approach for mutation calling of tumor exomes using multiple genomic pipelines. Cell Syst. 6, 271–281.e7 (2018).
Turajlic, S. et al. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol. 18, 1009–1021 (2017).
Mandal, R. et al. Genetic diversity of tumors with mismatch repair deficiency influences anti-PD-1 immunotherapy response. Science 364, 485–491 (2019).
Stein, A. & Folprecht, G. Immunotherapy of colon cancer. Oncol. Res. Treat. 41, 282–285 (2018).
Van den Eynden, J., Basu, S. & Larsson, E. Somatic mutation patterns in hemizygous genomic regions unveil purifying selection during tumor evolution. PLoS Genet. 12, e1006506 (2016).
Weghorn, D. & Sunyaev, S. Bayesian inference of negative and positive selection in human cancers. Nat. Genet. 49, 1785–1788 (2017).
Rosenbloom, K. R. et al. The UCSC Genome Browser database: 2015 update. Nucleic Acids Res. 43, D670–D681 (2014).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Birger, C. et al. FireCloud, a scalable cloud-based platform for collaborative genome analysis: strategies for reducing and controlling costs. Preprint at bioRxiv https://doi.org/10.1101/209494 (2017).
González-Galarza, F. F. et al. Allele frequency net 2015 update: new features for HLA epitopes, KIR and disease and HLA adverse drug reaction associations. Nucleic Acids Res. 43, D784–D788 (2015).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Nielsen, M. & Andreatta, M. NetMHCpan-3.0; improved prediction of binding to MHC class I molecules integrating information from multiple receptor and peptide length datasets. Genome Med. 8, 33 (2016).
Acknowledgements
The results published here are in whole or in part based on data generated by TCGA. Information about TCGA and the investigators and institutions who constitute the TCGA research network can be found at https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga. We are most grateful to the patients, investigators, clinicians, technical personnel, and funding bodies who contributed to TCGA, thereby making this study possible. This work was supported by grants from the Wenner-Gren Foundation (SSv2017-005 to E.L.), the Swedish Cancer Society (CAN 2015/541 to E.L.), the Knut and Alice Wallenberg Foundation (KAW 2014.0057 and KAW 2015.0144 to E.L.), the Swedish Medical Research Council (2018-02852 to E.L.), the Swedish Foundation for Strategic Research (RB13-0204 to E.L.), EMBO (STF7729 to J.V.d.E.) and Cancer Research UK (C14303/A17197 and A21141 to M.L.M.).
Author information
Authors and Affiliations
Contributions
J.V.d.E., E.L. and M.L.M. designed and conceptualized the study. A.J.-S. provided input on study design and contributed to the interpretation of the results. J.V.d.E. was responsible for data analysis and drafted the manuscript. All authors discussed the results, edited and finalized the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–14 and Table 1
Supplementary Table 2
Different metrics calculated for each sample and tumor type in this study. HBMR (first tab) and dNHLA/dNnonHLA (second tab for TCGA samples and third tab for different tumor types). HBMR was calculated considering the HLA-binding annotation of the prototypical HLA genotype, while dNHLA/dNnonHLA was determined using HLA affinities from mutated peptides and TCGA sample-specific HLA genotypes. Normalizations were performed under a trinucleotide substitution model. See Supplementary Table 1 for cancer type abbreviations and main Methods for other abbreviations.
Rights and permissions
About this article
Cite this article
Van den Eynden, J., Jiménez-Sánchez, A., Miller, M.L. et al. Lack of detectable neoantigen depletion signals in the untreated cancer genome. Nat Genet 51, 1741–1748 (2019). https://doi.org/10.1038/s41588-019-0532-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0532-6
This article is cited by
-
Benchmark of tools for in silico prediction of MHC class I and class II genotypes from NGS data
BMC Genomics (2023)
-
MHC II immunogenicity shapes the neoepitope landscape in human tumors
Nature Genetics (2023)
-
Spatial biology of cancer evolution
Nature Reviews Genetics (2023)
-
Antigen presentation in cancer — mechanisms and clinical implications for immunotherapy
Nature Reviews Clinical Oncology (2023)
-
Cancer genomes tolerate deleterious coding mutations through somatic copy number amplifications of wild-type regions
Nature Communications (2023)