Abstract
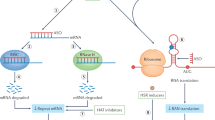
Noncoding repeat expansions cause various neuromuscular diseases, including myotonic dystrophies, fragile X tremor/ataxia syndrome, some spinocerebellar ataxias, amyotrophic lateral sclerosis and benign adult familial myoclonic epilepsies. Inspired by the striking similarities in the clinical and neuroimaging findings between neuronal intranuclear inclusion disease (NIID) and fragile X tremor/ataxia syndrome caused by noncoding CGG repeat expansions in FMR1, we directly searched for repeat expansion mutations and identified noncoding CGG repeat expansions in NBPF19 (NOTCH2NLC) as the causative mutations for NIID. Further prompted by the similarities in the clinical and neuroimaging findings with NIID, we identified similar noncoding CGG repeat expansions in two other diseases: oculopharyngeal myopathy with leukoencephalopathy and oculopharyngodistal myopathy, in LOC642361/NUTM2B-AS1 and LRP12, respectively. These findings expand our knowledge of the clinical spectra of diseases caused by expansions of the same repeat motif, and further highlight how directly searching for expanded repeats can help identify mutations underlying diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The genotyping microarray data and sequence data obtained by massively parallel sequencing analysis, including whole-genome sequencing and transcriptome analyses, are available on request from the corresponding author. Since whole-genome sequence data are protected by the Personal Information Protection Law, these data are available under regulation by the institutional review board.
References
Loureiro, J. R., Oliveira, C. L. & Silveira, I. Unstable repeat expansions in neurodegenerative diseases: nucleocytoplasmic transport emerges on the scene. Neurobiol. Aging 39, 174–183 (2016).
Vissers, L. E. et al. A de novo paradigm for mental retardation. Nat. Genet. 42, 1109–1112 (2010).
Lindenberg, R., Rubinstein, L. J., Herman, M. M. & Haydon, G. B. A light and electron microscopy study of an unusual widespread nuclear inclusion body disease. A possible residuum of an old herpesvirus infection. Acta Neuropathol. 10, 54–73 (1968).
Haltia, M., Somer, H., Palo, J. & Johnson, W. G. Neuronal intranuclear inclusion disease in identical twins. Ann. Neurol. 15, 316–321 (1984).
Sone, J. et al. Clinicopathological features of adult-onset neuronal intranuclear inclusion disease. Brain 139, 3170–3186 (2016).
Takahashi-Fujigasaki, J., Nakano, Y., Uchino, A. & Murayama, S. Adult-onset neuronal intranuclear hyaline inclusion disease is not rare in older adults. Geriatr. Gerontol. Int. 16, 51–56 (2016).
Kimber, T. E. et al. Familial neuronal intranuclear inclusion disease with ubiquitin positive inclusions. J. Neurol. Sci. 160, 33–40 (1998).
Sone, J. et al. Neuronal intranuclear hyaline inclusion disease showing motor-sensory and autonomic neuropathy. Neurology 65, 1538–1543 (2005).
Yamaguchi, N. et al. An autopsy case of familial neuronal intranuclear inclusion disease with dementia and neuropathy. Intern. Med. 57, 3459–3462 (2018).
Sone, J. et al. Neuronal intranuclear inclusion disease cases with leukoencephalopathy diagnosed via skin biopsy. J. Neurol. Neurosurg. Psychiatry 85, 354–356 (2014).
Sone, J. et al. Skin biopsy is useful for the antemortem diagnosis of neuronal intranuclear inclusion disease. Neurology 76, 1372–1376 (2011).
Nakano, Y. et al. PML nuclear bodies are altered in adult-onset neuronal intranuclear hyaline inclusion disease. J. Neuropathol. Exp. Neurol. 76, 585–594 (2017).
Takumida, H. et al. Case of a 78-year-old woman with a neuronal intranuclear inclusion disease. Geriatr. Gerontol. Int. 17, 2623–2625 (2017).
Sugiyama, A. et al. MR imaging features of the cerebellum in adult-onset neuronal intranuclear inclusion disease: 8 cases. Am. J. Neuroradiol. 38, 2100–2104 (2017).
Hunsaker, M. R. et al. Widespread non-central nervous system organ pathology in fragile X premutation carriers with fragile X-associated tremor/ataxia syndrome and CGG knock-in mice. Acta Neuropathol. 122, 467–479 (2011).
Hagerman, R. J. et al. Intention tremor, parkinsonism, and generalized brain atrophy in male carriers of fragile X. Neurology 57, 299–301 (2001).
Doi, K. et al. Rapid detection of expanded short tandem repeats in personal genomics using hybrid sequencing. Bioinformatics 30, 815–822 (2014).
Ishiura, H. et al. Expansions of intronic TTTCA and TTTTA repeats in benign adult familial myoclonic epilepsy. Nat. Genet. 50, 581–590 (2018).
Vandepoele, K., Van Roy, N., Staes, K., Speleman, F. & van Roy, F. A novel gene family NBPF: intricate structure generated by gene duplication during primate evolution. Mol. Biol. Evol. 22, 2265–2275 (2005).
Fiddes, I. T. et al. Human-specific NOTCH2NL genes affect Notch signaling and cortical neurogenesis. Cell 173, 1356–1369 (2018).
Suzuki, I. K. et al. Human-specific NOTCH2NL genes expand cortical neurogenesis through Delta/Notch regulation. Cell 173, 1370–1384 (2018).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Flusberg, B. A. et al. Direct detection of DNA methylation during single-molecule, real-time sequencing. Nat. Methods 7, 461–465 (2010).
Suzuki, Y. et al. Agln: measuring the landscape of CpG methylation of individual repetitive elements. Bioinformatics 32, 2911–2919 (2016).
Schuffler, M. D., Bird, T. D., Sumi, S. M. & Cook, A. A familial neuronal disease presenting as intestinal pseudoobstruction. Gastroenterology 75, 889–898 (1978).
Satoyoshi, E. & Kinoshita, M. Oculopharyngodistal myopathy. Arch. Neurol. 34, 89–92 (1977).
Durmus, H. et al. Oculopharyngodistal myopathy is a distinct entity: clinical and genetic features of 47 patients. Neurology 76, 227–235 (2011).
Zhao, J. et al. Clinical and muscle imaging findings in 14 mainland Chinese patients with oculopharyngodistal myopathy. PLoS ONE 10, e0128629 (2015).
Satoyoshi, E. Distal myopathy. Tohoku J. Exp. Med. 161, 1–19 (1990).
Brais, B. et al. Short GCG expansions in the PABP2 gene cause oculopharyngeal muscular dystrophy. Nat. Genet. 18, 164–167 (1998).
Seltzer, M. M. et al. Prevalence of CGG expansions of the FMR1 gene in a US population-based sample. Am. J. Med. Genet. B Neuropsychiatr. Genet. 159B, 589–597 (2012).
Beck, J. et al. Large C9orf72 hexanucleotide repeat expansions are seen in multiple neurodegenerative syndromes and are more frequent than expected in the UK population. Am. J. Hum. Genet. 92, 345–353 (2013).
Renton, A. E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72, 257–268 (2011).
Jacquemont, S. et al. Penetrance of the fragile X-associated tremor/ataxia syndrome in a premutation carrier population. J. Am. Med. Assoc. 291, 460–469 (2004).
Coffey, S. M. et al. Expanded clinical phenotype of women with the FMR1 premutation. Am. J. Med. Genet. A 146A, 1009–1016 (2008).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Fratta, P. et al. Screening a UK amyotrophic lateral sclerosis cohort provides evidence of multiple origins of the C9orf72 expansion. Neurobiol. Aging 36, 546.e1–546.e7 (2015).
Buxton, J. et al. Detection of an unstable fragment of DNA specific to individuals with myotonic dystrophy. Nature 355, 547–548 (1992).
Zu, T. et al. Non-ATG-initiated translation directed by microsatellite expansions. Proc. Natl Acad. Sci. USA 108, 260–265 (2011).
Todd, P. K. et al. CGG repeat-associated translation mediates neurodegeneration in fragile X tremor ataxia syndrome. Neuron 78, 440–455 (2013).
Uyama, E., Uchino, M., Chateau, D. & Tomé, F. M. Autosomal recessive oculopharyngodistal myopathy in light of distal myopathy with rimmed vacuoles and oculopharyngeal muscular dystrophy. Neuromuscul. Disord. 8, 119–125 (1998).
Jin, P. et al. Pur α binds to rCGG repeats and modulates repeat-mediated neurodegeneration in a Drosophila model of fragile X tremor/ataxia syndrome. Neuron 55, 556–564 (2007).
Sofola, O. A. et al. RNA-binding proteins hnRNP A2/B1 and CUGBP1 suppress fragile X CGG premutation repeat-induced neurodegeneration in a Drosophila model of FXTAS. Neuron 55, 565–571 (2007).
Bahlo, M. et al. Recent advances in the detection of repeat expansions with short-read next-generation sequencing. F1000Res. 7, 736 (2018).
Mitsuhashi, S. et al. Tandem-genotypes: robust detection of tandem repeat expansions from long DNA reads. Genome Biol. 20, 58 (2019).
Sznajder, J. et al. Intron retention induced by microsatellite expansions as a disease biomarker. Proc. Natl Acad. Sci. USA 115, 4234–4239 (2018).
Fukuda, Y. et al. SNP HiTLink: a high-throughput linkage analysis system employing dense SNP data. BMC Bioinformatics. 10, 121 (2009).
Gudbjartsson, D. F., Thorvaldsson, T., Kong, A., Gunnarsson, G. & Ingolfsdottir, A. Allegro version 2. Nat. Genet. 37, 1015–1016 (2005).
Kent, W. J. BLAT—the blast-like alignment tool. Genome Res. 14, 656–664 (2002).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Benson, G. Tandem repeat finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Frey, U. H., Bachmann, H. S., Peters, J. & Siffert, W. PCR-amplification of GC-rich regions: ‘slowdown PCR’. Nat. Protoc. 3, 1312–1317 (2008).
Su, J. et al. CpG_MP2: identification of CpG methylation patterns of genomic regions from high-throughput bisulfite sequencing data. Nucleic Acids Res. 41, e4 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-Seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Robinson, J. T. et al. Integrative genomic viewer. Nat. Biotechnol. 29, 24–26 (2011).
Miyazawa, H. et al. Homozygosity haplotype allows a genomewide search for the autosomal segments shared among patients. Am. J. Hum. Genet. 80, 1090–1102 (2007).
Acknowledgements
We thank the patients and family members for participating in the study. We also thank the neurologists, pathologists and radiologists of the patients for collecting and providing clinical information, K. J. L. Porto for proofreading, and K. Hirayama, M. Takeyama, Z. Wu and K. Wakabayashi for technical support. This work was supported in part by KAKENHI (Grants-in-Aid for Scientific Research on Innovative Areas (numbers 22129001 and 22129002) from the Ministry of Education, Culture, Sports, Science and Technology of Japan, Grants-in-Aid (H23-Jitsuyoka (Nanbyo)-Ippan-004 and H26-Jitsuyoka (Nanbyo)-Ippan-080) from the Ministry of Health, Labour and Welfare, Japan, and grants (numbers 15ek0108065h0002, 16kk0205001h0001, 17kk0205001h0002 and 17ek0109279h0001) from the Japan Agency for Medical Research and Development; all to S.T.). This work was also supported by KAKENHI (Grants-in-Aid for Young Scientists (numbers 15K20941 and 17H05085) from the Japan Society for the Promotion of Science; to H.I) and the Advanced Genome Research and Bioinformatics Study to Facilitate Medical Innovation (GRIFIN) from the Japan Agency for Medical Research and Development (to S. Morishita).
Author information
Authors and Affiliations
Contributions
H.I. and S.T. designed the research. H.I., S.S., M.A.A., J.K., M.Taira, J.M., Y.Takahashi, Y.I., T.Matsukawa, M.Tanaka, H.D., J.G., T.T. and S.T. performed the experiments and/or analyzed the data. H.I., S.S., J.Y., Y.Suzuki., W.Q., K.D. and S.Morishita performed the computational analysis. H.I., S.S., J.K., M.Taira, J.M., Y.Takahashi, Y.I., T.Mano, A.I., Y.H., M.K.M., T.Matsukawa, M.Tanaka, Y.Shirota, R.O., H.K., A.Mitsue., H.H., S.Morimoto, S.Murayama, Y.Shiio, Y.Saito, A.Mitsutake, M.K., T.S., Y.Sugiyama, M.H., G.O., Y.Terao, Y.N., A.T., Y.Sakiyama, Y.U.-K., J.Shinmi, K.O., Y.K., S.-Y.L., A.H.T., J.Shimizu, J.G., I.N., T.T. and S.T. collected and analyzed the clinical data and/or provided patients’ samples. H.I. and S.T. wrote the manuscript, together with contributions from S.S. and S.Morishita. All authors contributed to and critically reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–17, Supplementary Note and Supplementary Tables 1–10
Rights and permissions
About this article
Cite this article
Ishiura, H., Shibata, S., Yoshimura, J. et al. Noncoding CGG repeat expansions in neuronal intranuclear inclusion disease, oculopharyngodistal myopathy and an overlapping disease. Nat Genet 51, 1222–1232 (2019). https://doi.org/10.1038/s41588-019-0458-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0458-z
This article is cited by
-
RExPRT: a machine learning tool to predict pathogenicity of tandem repeat loci
Genome Biology (2024)
-
Plasma neurofilament light as a promising biomarker in neuronal intranuclear inclusion disease
Journal of Neurology (2024)
-
Letter to the editor on: Hornerin deposits in neuronal intranuclear inclusion disease: direct identification of proteins with compositionally biased regions in inclusions by Park et al. (2022)
Acta Neuropathologica Communications (2024)
-
A large pedigree study confirmed the CGG repeat expansion of RILPL1 Is associated with oculopharyngodistal myopathy
BMC Medical Genomics (2023)
-
Characterization of large-scale genomic differences in the first complete human genome
Genome Biology (2023)