Abstract
Mesenchymal (stromal) stem cells (MSCs) constitute populations of mesodermal multipotent cells involved in tissue regeneration and homeostasis in many different organs. Here we performed comprehensive characterization of the transcriptional and epigenomic changes associated with osteoblast and adipocyte differentiation of human MSCs. We demonstrate that adipogenesis is driven by considerable remodeling of the chromatin landscape and de novo activation of enhancers, whereas osteogenesis involves activation of preestablished enhancers. Using machine learning algorithms for in silico modeling of transcriptional regulation, we identify a large and diverse transcriptional network of pro-osteogenic and antiadipogenic transcription factors. Intriguingly, binding motifs for these factors overlap with SNPs related to bone and fat formation in humans, and knockdown of single members of this network is sufficient to modulate differentiation in both directions, thus indicating that lineage determination is a delicate balance between the activities of many different transcription factors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data generated from BM-hMSC-TERT4, AT-hMSC-TERT, A549, EAhy926, HEK293, PANC-1 and SH-SY5Y cells have been deposited in the GEO database under accession code GSE113253. Primary stromal cells from bone marrow were obtained from volunteers (males, 20–28 years) in the local community of Odense, recruited through advertisements in newspapers. Exclusion criteria were known metabolic bone diseases; use of medications influencing bone and fat metabolism, such as glucocorticoids and anabolic drugs; and contraindications for bone marrow biopsies. The study was performed according to the Declaration of Helsinki and approved by the Scientific Ethics committee at Odense University Hospital (S-20150013), and signed informed consent was obtained from all participants. Primary stromal cells from muscle were obtained from previous study cohorts91, whereas WAT-derived ones were obtained from volunteers (females, 48–67 years) in the local community of Copenhagen, recruited through advertisements in newspapers. Volunteers were healthy, with normal weight and normal glucose tolerance. Both studies were performed according to the Declaration of Helsinki and approved by the Scientific Ethics committee of the Capital Regions of Copenhagen and Fredriksberg Municipalities (muscle, H-A-2009-020; WAT, H-A-2008-081, H-KF-01-141/04), and signed informed consent was obtained from all participants. RNA-seq and ATAC-seq raw reads are available after approval by the local scientific ethics committee and the data-responsible person (WAT, C.S.; muscle, B. K. Pedersen; bone marrow, M.K.). Processed data including ATAC-seq quantification at ATAC sites and gene count tables are available under GSE113253. Accession codes for published datasets used in this study are as follows: processed RNA-seq data for primary bone marrow–derived MSCs from mice are available from GEO under accession code GSE79814 (ref. 29); raw RNA-seq data for uncultured primary human MSCs31, osteoblasts26 and adipocytes27 are available through the corresponding author upon approval by the respective data/ethics access committee; processed RNA-seq read counts are available from ENCODE (Supplementary Table 3)30; and processed microarray data for human mesenchymal biopsies are available from GEO under accession codes GSE35959 (ref. 79), GSE27951 (ref. 80), GSE15790 (ref. 81), GSE73108 (ref. 82), GSE44000 (ref. 83), GSE48964 (ref. 84), GSE29718 (ref. 85), GSE15773 (ref. 86), GSE9624 (ref. 87), GSE15524 (ref. 88), GSE39540 (ref. 89) and GSE12274 (ref. 90).
Change history
25 March 2019
In the version of this article initially published, in the graph keys in Fig. 1i, the colors indicating ‘Ob’ and ‘Ad’ were red and blue, respectively, but should have been blue and red, respectively; the shapes indicating ‘MUS’ and ‘BM’ were a triangle and a square, respectively, but should have been a square and a triangle, respectively. The errors have been corrected in the HTML and PDF versions of the article.
References
Kassem, M. & Marie, P. J. Senescence-associated intrinsic mechanisms of osteoblast dysfunctions. Aging Cell 10, 191–197 (2011).
Squillaro, T., Peluso, G. & Galderisi, U. Clinical trials with mesenchymal stem cells: an update. Cell Transplant. 25, 829–848 (2016).
Shafiee, A. et al. A comparison between osteogenic differentiation of human unrestricted somatic stem cells and mesenchymal stem cells from bone marrow and adipose tissue. Biotechnol. Lett. 33, 1257–1264 (2011).
Vishnubalaji, R., Al-Nbaheen, M., Kadalmani, B., Aldahmash, A. & Ramesh, T. Comparative investigation of the differentiation capability of bone-marrow- and adipose-derived mesenchymal stem cells by qualitative and quantitative analysis. Cell Tissue Res. 347, 419–427 (2012).
Hayashi, O., Katsube, Y., Hirose, M., Ohgushi, H. & Ito, H. Comparison of osteogenic ability of rat mesenchymal stem cells from bone marrow, periosteum, and adipose tissue. Calcif. Tissue Int. 82, 238–247 (2008).
Siersbaek, M. S. et al. Genome-wide profiling of peroxisome proliferator-activated receptor gamma in primary epididymal, inguinal, and brown adipocytes reveals depot-selective binding correlated with gene expression. Mol. Cell. Biol. 32, 3452–3463 (2012).
Liu, T. M. et al. Identification of common pathways mediating differentiation of bone marrow– and adipose tissue–derived human mesenchymal stem cells into three mesenchymal lineages. Stem Cells 25, 750–760 (2007).
Komori, T. et al. Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell 89, 755–764 (1997).
Otto, F. et al. Cbfa1, a candidate gene for cleidocranial dysplasia syndrome, is essential for osteoblast differentiation and bone development. Cell 89, 765–771 (1997).
Tontonoz, P., Hu, E. & Spiegelman, B. M. Stimulation of adipogenesis in fibroblasts by PPARγ2, a lipid-activated transcription factor. Cell 79, 1147–1156 (1994).
Rosen, E. D. et al. PPARγ is required for the differentiation of adipose tissue in vivo and in vitro. Mol. Cell 4, 611–617 (1999).
Akune, T. et al. PPARγ insufficiency enhances osteogenesis through osteoblast formation from bone marrow progenitors. J. Clin. Invest. 113, 846–855 (2004).
Zhang, X. et al. Runx2 overexpression enhances osteoblastic differentiation and mineralization in adipose-derived stem cells in vitro and in vivo. Calcif. Tissue Int. 79, 169–178 (2006).
Mikkelsen, T. S. et al. Comparative epigenomic analysis of murine and human adipogenesis. Cell 143, 156–169 (2010).
Nielsen, R. et al. Genome-wide profiling of PPARγ:RXR and RNA polymerase II occupancy reveals temporal activation of distinct metabolic pathways and changes in RXR dimer composition during adipogenesis. Genes Dev. 22, 2953–2967 (2008).
Siersbaek, R. et al. Extensive chromatin remodelling and establishment of transcription factor ‘hotspots’ during early adipogenesis. EMBO J. 30, 1459–1472 (2011).
Meyer, M. B., Benkusky, N. A. & Pike, J. W. The RUNX2 cistrome in osteoblasts: characterization, down-regulation following differentiation, and relationship to gene expression. J. Biol. Chem. 289, 16016–16031 (2014).
Thompson, B., Varticovski, L., Baek, S. & Hager, G. L. Genome-wide chromatin landscape transitions identify novel pathways in early commitment to osteoblast differentiation. PLoS One 11, e0148619 (2016).
Wu, H. et al. Chromatin dynamics regulate mesenchymal stem cell lineage specification and differentiation to osteogenesis. Biochim. Biophys. Acta 1860, 438–449 (2017).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Siersbaek, R. et al. Dynamic rewiring of promoter-anchored chromatin loops during adipocyte differentiation. Mol. Cell 66, 420–435 (2017).
Siersbaek, R. et al. Transcription factor cooperativity in early adipogenic hotspots and super-enhancers. Cell Rep. 7, 1443–1455 (2014).
Stergachis, A. B. et al. Developmental fate and cellular maturity encoded in human regulatory DNA landscapes. Cell 154, 888–903 (2013).
Simonsen, J. L. et al. Telomerase expression extends the proliferative life-span and maintains the osteogenic potential of human bone marrow stromal cells. Nat. Biotechnol. 20, 592–596 (2002).
Wiwie, C. et al. Elucidation of time-dependent systems biology cell response patterns with time course network enrichment. Preprint at https://arxiv.org/abs/1710.10262 (2017).
Fujita, K. et al. Isolation and characterization of human osteoblasts from needle biopsies without in vitro culture. Osteoporos. Int. 25, 887–895 (2014).
Allum, F. et al. Characterization of functional methylomes by next-generation capture sequencing identifies novel disease-associated variants. Nat. Commun. 6, 7211 (2015).
Tatrai, P. et al. Combined introduction of Bmi-1 and hTERT immortalizes human adipose tissue–derived stromal cells with low risk of transformation. Biochem. Biophys. Res. Commun. 422, 28–35 (2012).
Meyer, M. B., Benkusky, N. A., Sen, B., Rubin, J. & Pike, J. W. Epigenetic plasticity drives adipogenic and osteogenic differentiation of marrow-derived mesenchymal stem cells. J. Biol. Chem. 291, 17829–17847 (2016).
The ENCODE Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Roforth, M. M. et al. Global transcriptional profiling using RNA sequencing and DNA methylation patterns in highly enriched mesenchymal cells from young versus elderly women. Bone 76, 49–57 (2015).
Zhu, Y. et al. Predicting enhancer transcription and activity from chromatin modifications. Nucleic Acids Res. 41, 10032–10043 (2013).
Sanyal, A., Lajoie, B. R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012).
Madsen, J. G. S. et al. Integrated analysis of motif activity and gene expression changes of transcription factors. Genome Res. 28, 243–255 (2018).
Rauch, A. et al. Glucocorticoids suppress bone formation by attenuating osteoblast differentiation via the monomeric glucocorticoid receptor. Cell Metab. 11, 517–531 (2010).
Steger, D. J. et al. Propagation of adipogenic signals through an epigenomic transition state. Genes Dev. 24, 1035–1044 (2010).
Ahrends, R. et al. Controlling low rates of cell differentiation through noise and ultrahigh feedback. Science 344, 1384–1389 (2014).
Miyamoto, T., Furusawa, C. & Kaneko, K. Pluripotency, differentiation, and reprogramming: a gene expression dynamics model with epigenetic feedback regulation. PLoS Comput. Biol. 11, e1004476 (2015).
Signorelli, M., Vinciotti, V. & Wit, E. C. NEAT: an efficient network enrichment analysis test. BMC Bioinformatics 17, 352 (2016).
Jaager, K., Islam, S., Zajac, P., Linnarsson, S. & Neuman, T. RNA-seq analysis reveals different dynamics of differentiation of human dermis- and adipose-derived stromal stem cells. PLoS One 7, e38833 (2012).
Ducy, P., Schinke, T. & Karsenty, G. The osteoblast: a sophisticated fibroblast under central surveillance. Science 289, 1501–1504 (2000).
Corces, M. R. et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat. Genet. 48, 1193–1203 (2016).
Gonzalez, A. J., Setty, M. & Leslie, C. S. Early enhancer establishment and regulatory locus complexity shape transcriptional programs in hematopoietic differentiation. Nat. Genet. 47, 1249–1259 (2015).
Ye, M. et al. Hematopoietic stem cells expressing the myeloid lysozyme gene retain long-term, multilineage repopulation potential. Immunity 19, 689–699 (2003).
Zhao, L. et al. Smurf1 inhibits mesenchymal stem cell proliferation and differentiation into osteoblasts through JunB degradation. J. Bone Miner. Res. 25, 1246–1256 (2010).
Wagegg, M. et al. Hypoxia promotes osteogenesis but suppresses adipogenesis of human mesenchymal stromal cells in a hypoxia-inducible factor-1 dependent manner. PLoS One 7, e46483 (2012).
Lambertini, E. et al. Slug gene expression supports human osteoblast maturation. Cell. Mol. Life Sci. 66, 3641–3653 (2009).
Leupin, O. et al. Control of the SOST bone enhancer by PTH using MEF2 transcription factors. J. Bone Miner. Res. 22, 1957–1967 (2007).
Choy, L. & Derynck, R. Transforming growth factor-β inhibits adipocyte differentiation by Smad3 interacting with CCAAT/enhancer-binding protein (C/EBP) and repressing C/EBP transactivation function. J. Biol. Chem. 278, 9609–9619 (2003).
Zhao, X. et al. Comparative analyses by sequencing of transcriptomes during skeletal muscle development between pig breeds differing in muscle growth rate and fatness. PLoS One 6, e19774 (2011).
Uccelli, A., Moretta, L. & Pistoia, V. Mesenchymal stem cells in health and disease. Nat. Rev. Immunol. 8, 726–736 (2008).
Pillai, I. C. et al. Cardiac fibroblasts adopt osteogenic fates and can be targeted to attenuate pathological heart calcification. Cell Stem Cell 20, 218–232 (2017).
Jespersen, N. Z. et al. A classical brown adipose tissue mRNA signature partly overlaps with brite in the supraclavicular region of adult humans. Cell Metab. 17, 798–805 (2013).
Broholm, C. et al. LIF is a contraction-induced myokine stimulating human myocyte proliferation. J. Appl. Physiol. 111, 251–259 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010).
Uhlen, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 29.1–29.9 (2015).
Li, Q. H., Brown, J. B., Huang, H. Y. & Bickel, P. J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
MacArthur, J. et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 45, D896–D901 (2017).
Horikoshi, M. et al. Discovery and fine-mapping of glycaemic and obesity-related trait loci using high-density imputation. PLoS Genet. 11, e1005230 (2015).
Dastani, Z. et al. Novel loci for adiponectin levels and their influence on type 2 diabetes and metabolic traits: a multi-ethnic meta-analysis of 45,891 individuals. PLoS Genet. 8, e1002607 (2012).
Chu, A. Y. et al. Multiethnic genome-wide meta-analysis of ectopic fat depots identifies loci associated with adipocyte development and differentiation. Nat. Genet. 49, 125–130 (2017).
Surakka, I. et al. The impact of low-frequency and rare variants on lipid levels. Nat. Genet. 47, 589–597 (2015).
Kemp, J. P. et al. Identification of 153 new loci associated with heel bone mineral density and functional involvement of GPC6 in osteoporosis. Nat. Genet. 49, 1468–1475 (2017).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Demenais, F. et al. Multiancestry association study identifies new asthma risk loci that colocalize with immune-cell enhancer marks. Nat. Genet. 50, 42–53 (2018).
Jun, G. R. et al. Transethnic genome-wide scan identifies novel Alzheimer’s disease loci. Alzheimers Dement. 13, 727–738 (2017).
Day, F. R. et al. Large-scale genomic analyses link reproductive aging to hypothalamic signaling, breast cancer susceptibility and BRCA1-mediated DNA repair. Nat. Genet. 47, 1294–1303 (2015).
Pers, T. H., Timshel, P. & Hirschhorn, J. N. SNPsnap: a web-based tool for identification and annotation of matched SNPs. Bioinformatics 31, 418–420 (2015).
Davis, S. & Meltzer, P. S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847 (2007).
Benisch, P. et al. The transcriptional profile of mesenchymal stem cell populations in primary osteoporosis is distinct and shows overexpression of osteogenic inhibitors. PLoS One 7, e45142 (2012).
Keller, P. et al. Gene-chip studies of adipogenesis-regulated microRNAs in mouse primary adipocytes and human obesity. BMC Endocr. Disord. 11, 7 (2011).
van Tienen, F. H. et al. Preadipocytes of type 2 diabetes subjects display an intrinsic gene expression profile of decreased differentiation capacity. Int. J. Obes. 35, 1154–1164 (2011).
Wentworth, J. M. et al. GM3 ganglioside and phosphatidylethanolamine-containing lipids are adipose tissue markers of insulin resistance in obese women. Int. J. Obes. 40, 706–713 (2016).
Deng, T. et al. Class II major histocompatibility complex plays an essential role in obesity-induced adipose inflammation. Cell Metab. 17, 411–422 (2013).
Onate, B. et al. Stem cells isolated from adipose tissue of obese patients show changes in their transcriptomic profile that indicate loss in stemcellness and increased commitment to an adipocyte-like phenotype. BMC Genomics 14, 625 (2013).
Tam, C. S. et al. An early inflammatory gene profile in visceral adipose tissue in children. Int. J. Pediatr. Obes. 6, e360–e363 (2011).
Hardy, O. T. et al. Body mass index–independent inflammation in omental adipose tissue associated with insulin resistance in morbid obesity. Surg. Obes. Relat. Dis. 7, 60–67 (2011).
Aguilera, C. M. et al. Genome-wide expression in visceral adipose tissue from obese prepubertal children. Int. J. Mol. Sci. 16, 7723–7737 (2015).
MacLaren, R. E., Cui, W., Lu, H., Simard, S. & Cianflone, K. Association of adipocyte genes with ASP expression: a microarray analysis of subcutaneous and omental adipose tissue in morbidly obese subjects. BMC Med. Genomics 3, 3 (2010).
Alves, H. et al. A mesenchymal stromal cell gene signature for donor age. PLoS One 7, e42908 (2012).
Wagner, W. et al. Aging and replicative senescence have related effects on human stem and progenitor cells. PLoS One 4, e5846 (2009).
Green, C. J., Pedersen, M., Pedersen, B. K. & Scheele, C. Elevated NF-κB activation is conserved in human myocytes cultured from obese type 2 diabetic patients and attenuated by AMP-activated protein kinase. Diabetes 60, 2810–2819 (2011).
Acknowledgements
We thank B. Klarlund Pedersen (Centre of Inflammation and Metabolism and Centre for Physical Activity Research, Rigshospitalet University Hospital of Copenhagen, Copenhagen, Denmark) for providing primary stromal cells from muscle tissue; T. E. Andersen (Research Unit of Clinical Microbiology, Odense University Hospital and Department of Clinical Research University of Southern Denmark, Odense, Denmark) for EAhy926 cells; and B. S. Andresen (Department of Biochemistry and Molecular Biology, University of Southern Denmark, Odense, Denmark) for PANC-1, A549 and SH-SY5Y cells. This work was carried out mainly at the Villum Center for Bioanalytical Sciences, Department of Biochemistry and Molecular Biology, SDU, with support from the Villum Foundation. The work was supported by grants from the Danish Independent Research Council | Natural Sciences, the Novo Nordisk Foundation and the Lundbeck Foundation and by the Danish National Research Foundation. The Novo Nordisk Foundation Center for Basic Metabolic Research is supported by an unconditional grant from the Novo Nordisk Foundation to the University of Copenhagen. A.R. was supported by an EMBO Long-Term Fellowship (ALTF 1544-2011). A.R., M.T. and N.Z.J. received a postdoc stipend and N.Z.J. and I.F. received a PhD stipend from the Danish Diabetes Academy supported by the Novo Nordisk Foundation. C.W. and J.B. were supported by a VILLUM Young Investigator grant (13154). M.K. was supported by the Novo Nordisk Foundation (NNF15OC0016284).
Author information
Authors and Affiliations
Contributions
Conceptualization: A.R., S.M., A.K.H. and M.K.; software: J.G.S.M., C.W., R.R. and J.B.; formal analysis: A.R., A.K.H., J.G.S.M. and C.W.; investigation: A.R., M.L., I.F., E.L.V.H. and M.R.M.; primary cell isolation: N.Z.J., C.S. and M.T.; resources: A.R., R.N., B.D.L., J.G.S.M., C.W., J.B. and M.K.; data curation: A.R. and A.K.H.; writing, original draft: A.R. and S.M.; writing, review and editing: A.R., S.M., A.K.H., M.K., J.G.S.M., M.L. and E.L.V.H.; visualization: A.R.; supervision: S.M.; project administration: A.R. and S.M.; funding acquisition: A.R., J.B., M.K. and S.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
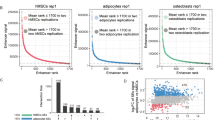
Supplementary Figure 1 Adipogenesis requires more dramatic changes in the transcriptome than osteogenesis.
Related to Fig. 1. (a) Bar plot showing the p value for the overlap of BM-hMSC-TERT4 osteoblast-selective genes (left, Cl. 5, n = 291) or adipocyte-selective genes (right, Cl. 9, n = 358) with sample-selective gene expression (RNA-seq) in samples from different tissues and cell lines (172 ENCODE RNA-seq profiles and RNA-seq data from human primary stromal cells and freshly isolated human osteoblasts 26 and adipocytes 27). Sample-selective gene expression is evaluated by R package TissueEnrich and P value is based on a hypergeometric test. Samples were ordered according to P value and top 17 are shown. Bold letters indicate RNA-seq samples from human primary stromal cells from this study. (b) Alizarin red and Oil red-O staining of AT-hMSC-TERT cells after 14 days of osteoblast and adipocyte differentiation. Representative pictures of 3 independent experiments with similar results. (c) Scatter plot showing the expression levels of osteoblast (top) and adipocyte transcription factors (bottom) in undifferentiated BM-hMSC-TERT4 and AT-hMSC-TERT cells. (d) Normalized RNA-seq counts for genes selectively expressed in BM-hMSC-TERT4 or AT-hMSC-TERT cells as well as genes being selectively upregulated during osteoblast (Cl. 5) or adipocyte differentiation (Cl. 9) in differentiating BM-hMSC-TERT4 and AT-hMSC-TERT cells. RPS4Y1, ribosomal protein S4, Y-linked 1; MAGEC2, MAGE family member C2; CSAG1, chondrosarcoma associated gene 1; ZFY, zinc finger protein, Y-linked; HTATSF1P2, HIV-1 Tat specific factor 1 pseudogene 2; TSIX, TSIX transcript, XIST antisense RNA; HCLS1, hematopoietic cell-specific Lyn substrate 1; XIST, X inactive specific transcript; BGLAP, bone gamma-carboxyglutamate protein; ALPL, alkaline phosphatase, liver/bone/kidney; IGFBP3, insulin like growth factor binding protein 3; SCUBE3, signal peptide, CUB domain and EGF like domain containing 3; OMD, osteomodulin; FABP4, fatty acid binding protein 4; PCK1, phosphoenolpyruvate carboxykinase 1; PLIN1, perilipin 1; FASN, fatty acid synthase; ADIPOQ, adiponectin. (e) Heat maps showing changes in gene expression during adipocyte and osteoblast differentiation of BM-hMSC-TERT4 (left) and AT-hMSC-TERT cells (right). Differentially expressed genes (FDR < 0.01 in at least one cell line) during either osteoblast (1,267 genes) or adipocyte differentiation (4,116 genes) were subjected to hierarchical clustering. The order of genes is identical in both heat maps. Side bar indicating Pearson’s correlation of temporal expression between the two cell lines of each single gene. (f) Box plot (thicker band, mean; box, first and third quartiles; whiskers, 1.5 times of interquartile range) showing highest induction levels over MSCs for genes in the commonly upregulated (n = 405), osteoblast-selective (n = 291) and adipocyte-selective cluster (n = 358) (Cl. 3, 5 and 9) during osteoblast (Ob) and adipocyte (Ad) differentiation of AT-hMSC-TERT cells. Statistical analysis was performed using an unpaired two-tailed Wilcoxon-Mann-Whitney test. (g) Box plot (thicker band, mean; box first and third quartiles; whiskers, 1.5 times of interquartile range) showing the expression levels in undifferentiated AT-hMSC-TERT cells. Clusters and statistics as in e. (h) Alizarin red and Oil red-O staining of human primary stromal cells from white adipose tissue (WAT), muscle (MUS) and bone marrow (BM) following 9 days of osteoblast and adipocyte differentiation from three different donors for each tissue. Differentiation assay was done once. (i) Box plot (thicker band, mean; box, first and third quartiles; whiskers, 1.5 times of interquartile range) showing highest induction levels during osteoblast (Ob) and adipocyte (Ad) differentiation of primary human stromal cells from white adipose tissue (WAT), muscle (MUS) and bone marrow (BM) over undifferentiated cells (MSC). Clusters and statistics as in e. (j) Box plot (thicker band: mean; box: first and third quartiles; whiskers: 1.5 times of interquartile range) showing the expression of genes in Cl. 3, 5 and 9 in undifferentiated primary human stromal cells from white adipose tissue (WAT), muscle (MUS) and bone marrow (BM). Clusters and statistics as in e. (k) Principal component analysis of differentiating human primary stromal cells (Fig. 1i) was used to project MSC cell lines (BM-hMSC-TERT4 (n = 3) and AT-hMSC-TERT (n = 2)) and non-mesenchymal cell lines (n = 1) prior to differentiation (MSC) and after 7 days of osteoblast and adipocyte differentiation. (l) Principal component analysis representing the variances in gene expression (RNA-seq) patterns between undifferentiated and in vitro differentiated murine primary bone marrow -derived MSCs (n = 3) for the indicated time points 29.
Supplementary Figure 2 Osteoblast enhancers are already accessible in MSCs, whereas adipogenesis requires extensive chromatin remodeling.
Related to Fig. 2. (a) Temporal changes in the mRNA level of PPARG and CEBPA (RPKM from RNA-seq, top panel, n = 3) and in chromatin accessibility of dynamic enhancers ± 50 kb from the transcription start site of CEBPA and PPARG (DNase-seq tag counts, middle panel. n = 2). Enhancer coordinates (hg19) are indicated. Center values represent mean and error bars standard deviation. The lower panel shows lipid droplet formation, as assessed by Oil Red-O staining at the indicated time points. Representative pictures of 3 independent experiments with similar results. (b) Pie charts illustrating the fraction of DNase I hypersentive sites with dynamic chromatin accessibility during osteoblast (blue) and adipocyte differentiation (red) of BM-hMSC-TERT4 cells ± 50 kb from the transcription start site of osteoblast- and adipocyte-selective gene cluster respectively. (c) UCSC genome browser screen shots showing ATAC-seq read density in undifferentiated BM-hMSC-TERT4 (n = 1) and AT-hMSC-TERT (n = 2, combined read density of two independent experiments) cells ± 50 kb of the transcription start site of ALPL and FABP4. DNase I hypersensitive sites in differentiating BM-hMSC-TERT4 cells are indicated. (d) Box plot (thicker band, mean; box, first and third quartiles; whiskers, 1.5 times of interquartile range) quantifying the number of ATAC-seq sites in undifferentiated AT-hMSC-TERT cells in the vicinity of osteoblast-selective (Cl. 5, n = 291) and adipocyte-selective genes (Cl. 9, n = 358). ATAC-seq sites were counted for each gene (± 50 kb from transcription start site). Statistical analysis was performed using an unpaired two-tailed Wilcoxon-Mann-Whitney test.
Supplementary Figure 3 Osteogenesis and adipogenesis involve different modes of enhancer activation.
Related to Fig. 3. (a) Model illustrating the calculation of cumulative log2 fold change of the indicated genomic mark at individual enhancers. UCSC screenshots show DNase-seq, MED1 and H3K27ac ChIP-seq read density for an osteoblast-selective locus (left) and an adipocyte-selective locus (right) during differentiation of BM-hMSC-TERT4 cells. Genome browser tracks show combined read density of two independent experiments. Enhancers are plotted according to their cumulative log2 fold change in MED1 or H3K27ac ChIP-seq signal relative to their cumulative log2 fold change in DNase-seq signal. (b) Density heat maps showing the cumulative log2 fold change in MED1 occupancy over DNase-seq signal at enhancer groups defined by dynamic MED1 occupancy during osteoblast and adipocyte differentiation. Upper left quadrant (red) indicates enhancers selectively activated during adipogenesis; upper right quadrant (purple) indicates enhancers activated during both adipogenesis and osteogenesis; lower right (blue) indicates enhancers selectively activated during osteogenesis; and lower left (white) indicates enhancers that are repressed during both osteoblast and adipocyte differentiation.
Supplementary Figure 4 Dynamic enhancer profiles are enriched for functional sequence variations associated with eQTLs and mesenchymal-tissue-related diseases.
Related to Fig. 4. (a) Heat map indicating the overlap between the different adipocyte enhancer clusters and osteoblast enhancer clusters using a Jaccard index. Number of enhancers in each cluster is indicated in Fig. 4a. (b) Heat map showing the enrichment of dynamic enhancers from the enhancer clusters (Fig. 4a) near genes (± 50 kb from the transcription start site) in the different RNA-seq clusters (Fig. 1c). Enrichment is indicated as log2 enrichment relative to genes that do not significantly change expression during differentiation of BM-hMSC-TERT4 cells. (c) Normalized RNA-seq counts (RPKM) of the osteoblast-selective gene KCNJ15 (Cl. 5) and the adipocyte-selective gene SYN2 (Cl. 9) during differentiation of BM-hMSC-TERT4 cells (n = 3). Center values represent mean and error bars standard deviation. KCNJ15, potassium voltage-gated channel subfamily J member 15; SYN2, synapsin 2. (d) UCSC screenshots visualizing overlap between putative enhancers in BM-hMSC-TERT4 cells and eQTLs for SYN2. DNase-seq, and MED1 and H3K27ac ChIP-seq signals are indicated for MSCs, and day 1 and 7 of adipocyte differentiation. Enhancers overlapping with eQTLs are highlighted with red arrows. Genome browser tracks show combined read density of two independent experiments. (e) UCSC screenshots visualizing overlap between putative enhancers in BM-hMSC-TERT4 cells and eQTLs for KCNJ15. DNase-seq, and MED1 and H3K27ac ChIP-seq signals are indicated for MSCs, and day 1 and 7 of osteoblast differentiation. Enhancers overlapping with eQTLs are highlighted with blue arrows. Genome browser tracks show combined read density of two independent experiments. (f) Heat map showing the significance of the overlap between disease-associated SNPs sampled from the GWAS catalog for the indicated disease with enhancer clusters from this study. Body mass index (n = 1500), Plasma adiponectin levels (n = 144), Subcutaneous adipose tissue mass (n = 143), Plasma triglycerides (n = 231), Bone mineral density (n = 5416), Type 2 diabetes (n = 1511), Height (n = 6971), Inflammatory bowel disease (n = 7413), Alzheimer’s (n = 757), Asthma (n = 1852), Onset of menopause (n = 862). P value was calculated using a binominal cumulative distribution function (linked to Fig. 4c).
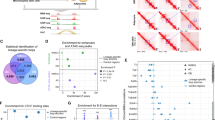
Supplementary Figure 5 Machine-learning strategies predict the repertoire of motifs that drive adipogenesis and osteogenesis.
Related to Fig. 5. (a) and (b) Motif activity during differentiation of BM-hMSC-TERT4 cells for selected transcription factors known to be associated with adipocyte or osteoblast differentiation in different model systems. CEBPA, CCAAT/enhancer binding protein alpha; KLF5, Kruppel like factor 5; CREB1, cAMP responsive element binding protein 1; EBF1, early B-cell factor 1; KLF15, Kruppel like factor 15; NR1D1, nuclear receptor subfamily 1 group D member 1 (Rev-Erbalpha/EAR1); PPARG, peroxisome proliferator activated receptor gamma; RXRA, retinoid X receptor alpha; STAT5A, signal transducer and activator of transcription 5A; ZEB1, zinc finger E-box binding homeobox 1. MSX2, msh homeobox 2; RUNX2, runt related transcription factor 2; SP3, Sp3 transcription factor; TBX3, T-box 3; HOXB7 homeobox B7; ATF4, activating transcription factor 4; DLX5, distal-less homeobox 5; LEF1, lymphoid enhancer binding factor 1; FOSL1, FOS like 1, AP-1 transcription factor subunit; FOXC2, forkhead box C2. The PWM for the factors are indicated (left). The graph shows changes in motif activity modeled based on data from DNase-seq (grey), MED1 ChIP-seq (green) or H3K27ac ChIP-seq (purple) experiments. RNA expression of the factors is indicated by a dashed line. (c) Box plot (thicker band: mean; box: first and third quartiles; whiskers: 1.5 times of interquartile range) showing the number of IMAGE-predicted target genes per transcription factor motif for different groups of motifs that gain or lose activity during differentiation of BM-hMSC-TERT4 cells as well as for motifs which don’t score a significant change in motif activity (linked to Fig. 5b, n = 126, n = 88, n = 92, n = 28, n = 171, n = 589). (d) Heat map (left) showing the enrichment of IMAGE-predicted target genes of PPARG (n = 286), RUNX2 (n = 350) and GR (n = 219) in the RNA-seq clusters over a random distribution (observed/expected). Violin plot (kernel density distribution surrounding boxplots with thicker band, mean; box, first and third quartiles; whiskers, 1.5 times of interquartile range) indicating the expression of the predicted target genes of PPARG, RUNX2 and GR in undifferentiated MSCs. (e) Visualization of subnetworks including only transcription factors as nodes and edges starting from transcription factors that gain motif activity during osteoblast (upper panel, n = 28) or during adipocyte differentiation (lower panel, n = 92). The subnetworks are part of a transcriptional network based on the IMAGE predicted target genes from each transcription factor motif (5,399 nodes and 26,5775 edges). (f) Network enrichment analysis between the RNA-seq clusters (Fig. 1c) using the IMAGE based transcriptional network (5,399 nodes and 265,775 edges) and the NEAT tool to calculate a P value based on a hypergeometric model (related to Fig. 5h). Arrows represent log odds scores of significantly (P < 0.05) overrepresented regulatory connections between the groups.
Supplementary Figure 6 A subset of MSC transcription factors drive osteogenesis and suppresses adipogenesis.
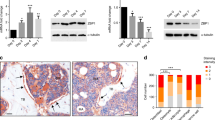
Related to Fig. 3. Motif activity during differentiation of BM-hMSC-TERT4 cells for selected transcription factors. ELK4, ETS transcription factor; SNAI2, snail family transcriptional repressor 2; MEF2A, myocyte enhancer factor 2A; NKX3-1, NK3 homeobox 1; ARNT, aryl hydrocarbon receptor nuclear translocator; SMAD3, SMAD family member 3; TEAD1, TEA domain transcription factor 1; TEAD4, TEA domain transcription factor 4; JUNB, JunB proto-oncogene, AP-1 transcription factor subunit; SMAD2, SMAD family member 2; PITX1, paired like homeodomain 1; FLI1, Fli-1 proto-oncogene, ETS transcription factor; HIF1A, hypoxia inducible factor 1 alpha subunit. The PWM for the factors are indicated (top). The graph shows changes in motif activity modeled based on data from DNase-seq (grey), MED1 ChIP-seq (green) or H3K27ac ChIP-seq (purple) experiments. RNA expression of the factors is indicated by a dashed line. (a) Heat map showing the average mRNA expression of the IMAGE-predicted target genes for the candidate transcription factors during differentiation of BM-hMSC-TERT4 cells. (b) Knockdown efficiency of the candidate factors as estimated by real-time PCR-based mRNA expression levels normalized to TBP three days after transfection, corresponding to day 0 of differentiation using two different siRNAs (n = 3). Center values represent mean, and error bars indicate s.e.m. Scramble siRNA (siCTR) or untreated cells (UT) are used as controls, and values are normalized for each gene to scrambled siRNA (siCTR). Statistical analysis was performed using an unpaired two-tailed t test. (c) Relative mRNA expression of TUBG1 normalized to TBP in MSCs (grey) and at day 7 of osteoblast (blue) or adipocyte (red) differentiation of BM-hMSC-TERT4 cells treated with two different siRNA against the indicated targets (n = 3). Scramble siRNA (siCTR) or untreated cells (UT) are used as controls. Dashed line indicates expression levels of knockdown control cells (siCTR). (d, f) Effect of knockdown of the indicated transcription factor on osteoblast differentiation as determined by ALP activity staining and Alizarin Red staining. Two different siRNAs were used for each target. Cells were fixed and stained after 5 and 14 days of osteoblast differentiation, respectively. Scramble siRNA (siCTR) or untreated cells (UT) are used as controls. Representative pictures of 2 independent experiments with similar results. (g) Effect of knockdown of the indicated transcription factor on adipocyte differentiation as determined by Oil red-O staining. Two different siRNAs were used for each target. Cells were fixed and stained after 14 days of adipocyte differentiation. Scramble siRNA (siCTR) or untreated cells (UT) are used as controls. Representative pictures of 3 independent experiments with similar results. (h) Bar plot showing the enrichment of factors with decreased motif activity during adipocyte differentiation (n = 202) over all transcription factors (n = 933) for being transcriptionally regulated in the indicated studies (P value < 0.05 when correlating expression levels between groups or a range of values such as age and BMI). (i) Bar plot showing the enrichment of factors with decreased motif activity during adipocyte differentiation (n = 202) over all transcription factors expressed above background (n = 933) that significantly overlap with eQTLs of differentially expressed genes belonging to indicated RNA-seq clusters (permutation P value < 0.1 from Fig. 6i). (j) Bar plot showing the enrichment of factors with decreased motif activity during adipocyte differentiation (n = 202) over all transcription factors expressed above background (n = 933) that significantly overlap with all SNPs associated to Body mass index and Bone mineral density (permutation P value < 0.1 from Fig. 6j).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–3
Rights and permissions
About this article
Cite this article
Rauch, A., Haakonsson, A.K., Madsen, J.G.S. et al. Osteogenesis depends on commissioning of a network of stem cell transcription factors that act as repressors of adipogenesis. Nat Genet 51, 716–727 (2019). https://doi.org/10.1038/s41588-019-0359-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0359-1
This article is cited by
-
Cytoskeletal rearrangement precedes nucleolar remodeling during adipogenesis
Communications Biology (2024)
-
DNA methylation restricts coordinated germline and neural fates in embryonic stem cell differentiation
Nature Structural & Molecular Biology (2024)
-
Screening for genes, miRNAs and transcription factors of adipogenic differentiation and dedifferentiation of mesenchymal stem cells
Journal of Orthopaedic Surgery and Research (2023)
-
Adipogenic and SWAT cells separate from a common progenitor in human brown and white adipose depots
Nature Metabolism (2023)
-
Super enhancers targeting ZBTB16 in osteogenesis protect against osteoporosis
Bone Research (2023)