Abstract
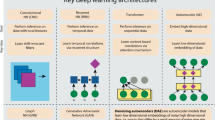
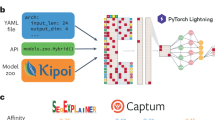
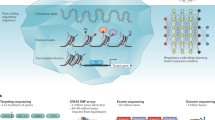
Deep learning methods are a class of machine learning techniques capable of identifying highly complex patterns in large datasets. Here, we provide a perspective and primer on deep learning applications for genome analysis. We discuss successful applications in the fields of regulatory genomics, variant calling and pathogenicity scores. We include general guidance for how to effectively use deep learning methods as well as a practical guide to tools and resources. This primer is accompanied by an interactive online tutorial.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Angermueller, C., Pärnamaa, T., Parts, L. & Stegle, O. Deep learning for computational biology. Mol. Syst. Biol. 12, 878 (2016).
Ching, T. et al. Opportunities and obstacles for deep learning in biology and medicine. J. R. Soc. Interface 15, 20170387 (2018).
Telenti, A., Lippert, C., Chang, P. C. & DePristo, M. Deep learning of genomic variation and regulatory network data. Hum. Mol. Genet. 27, R63–R71 (2018).
Yue, T. & Wang, H. Deep learning for genomics: a concise overview. Preprint at https://arxiv.org/abs/1802.00810 (2018).
Camacho, D. M., Collins, K. M., Powers, R. K., Costello, J. C. & Collins, J. J. Next-generation machine learning for biological networks. Cell 173, 1581–1592 (2018).
Libbrecht, M. W. & Noble, W. S. Machine learning applications in genetics and genomics. Nat. Rev. Genet. 16, 321–332 (2015).
Goodfellow, I., Bengio, Y., Courville, A. & Bengio, Y. Deep Learning Vol. 1 (MIT Press, Cambridge, 2016).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. Imagenet classification with deep convolutional neural networks. Adv. Neural Inf. Process. Syst. 1, 1097–1105 (2012).
Hinton, G. E. & Salakhutdinov, R. R. Reducing the dimensionality of data with neural networks. Science 313, 504–507 (2006).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Khodabandelou, G., Mozziconacci, J. & Routhier, E. Genome functional annotation using deep convolutional neural network. Preprint at https://www.biorxiv.org/content/early/2018/05/25/330308 (2018).
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 12, 931–934 (2015).
Powers, D. M. W. Evaluation: from precision, recall and F-measure to ROC, informedness, markedness and correlation. J. Mach. Learn. Technol. 2, 37–63 (2011).
Kelley, D. R., Snoek, J. & Rinn, J. L. Basset: learning the regulatory code of the accessible genome with deep convolutional neural networks. Genome Res. 26, 990–999 (2016).
Hastie, T., Tibshirani, R. & Friedman, J. H. The Elements of Statistical Learning Vol. 1 (Springer Science+Business Media, New York, 2001).
Quang, D. & Xie, X. DanQ: a hybrid convolutional and recurrent deep neural network for quantifying the function of DNA sequences. Nucleic Acids Res. 44, e107 (2016).
Sundararajan, M., Taly, A. & Yan, Q. Axiomatic attribution for deep networks. Preprint at https://arxiv.org/abs/1703.01365v2 (2017).
Shrikumar, A., Greenside, P. & Kundaje, A. Learning important features through propagating activation differences. Proc. Int. Conf. Mach. Learn. 70, 3145–3153 (2017).
Ribeiro, M. T., Singh, S. & Guestrin, C. “Why should I trust you?”: explaining the predictions of any classifier. in KDD 1135–1144 (AAAI Press, Menlo Park, CA, USA, 2016).
Park, Y. & Kellis, M. Deep learning for regulatory genomics. Nat. Biotechnol. 33, 825–826 (2015).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Lanchantin, J., Singh, R., Wang, B. & Qi, Y. Deep motif dashboard: visualizing and understanding genomic sequences using deep neural networks. Pac. Symp. Biocomput. 22, 254–265 (2017).
Zeng, H., Edwards, M. D., Liu, G. & Gifford, D. K. Convolutional neural network architectures for predicting DNA-protein binding. Bioinformatics 32, i121–i127 (2016).
Liu, F., Li, H., Ren, C., Bo, X. & Shu, W. PEDLA: predicting enhancers with a deep learning-based algorithmic framework. Sci. Rep. 6, 28517 (2016).
Kleftogiannis, D., Kalnis, P. & Bajic, V. B. DEEP: a general computational framework for predicting enhancers. Nucleic Acids Res. 43, e6 (2015).
Min, X. et al. Predicting enhancers with deep convolutional neural networks. BMC Bioinformatics 18 (Suppl. 13), 478 (2017).
Eser, U. & Stirling Churchman, L. FIDDLE: an integrative deep learning framework for functional genomic data inference. Preprint at https://www.biorxiv.org/content/early/2016/10/17/081380 (2016).
Li, Y., Shi, W. & Wasserman, W. W. Genome-wide prediction of cis-regulatory regions using supervised deep learning methods. BMC Bioinformatics 19, 202 (2018).
Wang, Y. et al. Predicting DNA methylation state of CpG dinucleotide using genome topological features and deep networks. Sci. Rep. 6, 19598 (2016).
Schreiber, J., Libbrecht, M., Bilmes, J. & Noble, W. Nucleotide sequence and DNaseI sensitivity are predictive of 3D chromatin architecture. Preprint at https://www.biorxiv.org/content/early/2017/01/30/103614 (2017).
Zeng, W., Wu, M. & Jiang, R. Prediction of enhancer-promoter interactions via natural language processing. BMC Genomics 19 (Suppl. 2), 84 (2018).
Shrikumar, A., Greenside, P. & Kundaje, A. Reverse-complement parameter sharing improves deep learning models for genomics. Preprint at https://www.biorxiv.org/content/early/2017/01/27/103663 (2017).
Tan, J., Hammond, J. H., Hogan, D. A. & Greene, C. S. ADAGE-based integration of publicly available Pseudomonas aeruginosa gene expression data with denoising autoencoders illuminates microbe-host interactions. mSystems 1, e00025-15 (2016).
Chen, Y., Li, Y., Narayan, R., Subramanian, A. & Xie, X. Gene expression inference with deep learning. Bioinformatics 32, 1832–1839 (2016).
Chen, L., Cai, C., Chen, V. & Lu, X. Learning a hierarchical representation of the yeast transcriptomic machinery using an autoencoder model. BMC Bioinformatics 17 (Suppl. 1), 9 (2016).
Cui, H. et al. Boosting gene expression clustering with system-wide biological information: a robust autoencoder approach. Preprint at https://www.biorxiv.org/content/early/2017/11/05/214122 (2017).
Xie, R., Wen, J., Quitadamo, A., Cheng, J. & Shi, X. A deep auto-encoder model for gene expression prediction. BMC Genomics 18 (Suppl. 9), 845 (2017).
Jha, A., Gazzara, M. R. & Barash, Y. Integrative deep models for alternative splicing. Bioinformatics 33, i274–i282 (2017).
Tripathi, R., Patel, S., Kumari, V., Chakraborty, P. & Varadwaj, P. K. DeepLNC, a long non-coding RNA prediction tool using deep neural network. Netw. Model. Anal. Health Inform. Bioinform. 5, 21 (2016).
Yu, N., Yu, Z. & Pan, Y. A deep learning method for lincRNA detection using auto-encoder algorithm. BMC Bioinformatics 18 (Suppl. 15), 511 (2017).
Hill, S. T. et al. A deep recurrent neural network discovers complex biological rules to decipher RNA protein-coding potential. Nucleic Acids Res. 46, 8105–8113 (2018).
Angermueller, C., Lee, H. J., Reik, W. & Stegle, O. DeepCpG: accurate prediction of single-cell DNA methylation states using deep learning. Genome Biol. 18, 67 (2017).
Shaham, U. et al. Removal of batch effects using distribution-matching residual networks. Bioinformatics 33, 2539–2546 (2017).
Lin, C., Jain, S., Kim, H. & Bar-Joseph, Z. Using neural networks for reducing the dimensions of single-cell RNA-Seq data. Nucleic Acids Res. 45, e156 (2017).
Poplin, R. et al. Creating a universal SNP and small indel variant caller with deep neural networks. Preprint at https://www.biorxiv.org/content/early/2018/03/20/092890 (2017).
Luo, R., Sedlazeck, F.J., Lam, T.-W. & Schatz, M. Clairvoyante: a multi-task convolutional deep neural network for variant calling in single molecule sequencing. Preprint at https://www.biorxiv.org/content/early/2018/09/26/310458 (2018).
Luo, R., Lam, T.-W. & Schatz, M. Skyhawk: an artificial neural network-based discriminator for reviewing clinically significant genomic variants. Preprint at https://www.biorxiv.org/content/early/2018/05/01/311985 (2018).
Torracinta, R. et al. Adaptive somatic mutations calls with deep learning and semi-simulated data. Preprint at https://www.biorxiv.org/content/early/2016/10/04/079087 (2016).
Boža, V., Brejová, B. & Vinař, T. DeepNano: deep recurrent neural networks for base calling in MinION nanopore reads. PLoS One 12, e0178751 (2017).
Teng, H., Hall, M.B., Duarte, T., Cao, M.D. & Coin, L. Chiron: translating nanopore raw signal directly into nucleotide sequence using deep learning. Preprint at https://www.biorxiv.org/content/early/2017/08/23/179531 (2017).
Qi, H. et al. MVP: predicting pathogenicity of missense variants by deep neural networks. Preprint at https://www.biorxiv.org/content/early/2018/02/02/259390 (2018).
Quang, D., Chen, Y. & Xie, X. DANN: a deep learning approach for annotating the pathogenicity of genetic variants. Bioinformatics 31, 761–763 (2015).
Korvigo, I., Afanasyev, A., Romashchenko, N. & Skoblov, M. Generalising better: applying deep learning to integrate deleteriousness prediction scores for whole-exome SNV studies. PLoS One 13, e0192829 (2018).
Yuan, Y. et al. DeepGene: an advanced cancer type classifier based on deep learning and somatic point mutations. BMC Bioinformatics 17, 476 (2016).
Yousefi, S. et al. Predicting clinical outcomes from large scale cancer genomic profiles with deep survival models. Sci. Rep. 7, 11707 (2017).
Ma, W., Qiu, Z., Song, J., Cheng, Q. & Ma, C. DeepGS: predicting phenotypes from genotypes using deep learning. Preprint at https://www.biorxiv.org/content/early/2017/12/31/241414 (2017).
Zhou, J. et al. Whole-genome deep learning analysis reveals causal role of noncoding mutations in autism. Preprint at https://www.biorxiv.org/content/early/2018/05/11/319681 (2018).
Zhou, J. et al. Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk. Nat. Genet. 50, 1171–1179 (2018).
Sundaram, L. et al. Predicting the clinical impact of human mutation with deep neural networks. Nat. Genet. 50, 1161–1170 (2018).
Avsec, Z. et al. Kipoi: accelerating the community exchange and reuse of predictive models for genomics. Preprint at https://www.biorxiv.org/content/early/2018/07/24/375345 (2018).
Webb, S. Deep learning for biology. Nature 554, 555–557 (2018).
Ghorbani, A., Abid, A. & Zou, J. Interpretation of neural networks is fragile. Preprint at https://arxiv.org/abs/1710.10547 (2017).
Gupta, A. & Zou, J. Feedback GAN (FBGAN) for DNA: a novel feedback-loop architecture for optimizing protein functions. Preprint at https://arxiv.org/abs/1804.01694 (2018).
Stranger, B. et al.; eGTEx Project. Enhancing GTEx by bridging the gaps between genotype, gene expression, and disease. Nat. Genet. 49, 1664–1670 (2017).
Acknowledgements
We thank N. Wineinger, R. Dias, J. di Iulio and D. Evans for comments on the paper. The work of A. Telenti, A. Torkamani and P.M. is supported by the Qualcomm Foundation and the NIH Center for Translational Science Award (CTSA, grant UL1TR002550). Further support to A. Torkamani is from U54GM114833 and U24TR002306. J.Z. is supported by a Chan–Zuckerberg Biohub Investigator grant and National Science Foundation (NSF) grant CRII 1657155.
Author information
Authors and Affiliations
Contributions
All authors conceived and designed the project. J.Z., M.H., P.M., A. Torkamani and A. Telenti wrote the manuscript. J.Z. and A.A. wrote the online tutorial.
Corresponding authors
Ethics declarations
Competing interests
M.H. is an employee of Peltarion.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zou, J., Huss, M., Abid, A. et al. A primer on deep learning in genomics. Nat Genet 51, 12–18 (2019). https://doi.org/10.1038/s41588-018-0295-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0295-5
This article is cited by
-
MSCAN: multi-scale self- and cross-attention network for RNA methylation site prediction
BMC Bioinformatics (2024)
-
Comprehensive analysis of mitochondria-related genes indicates that PPP2R2B is a novel biomarker and promotes the progression of bladder cancer via Wnt signaling pathway
Biology Direct (2024)
-
Genomic prediction using machine learning: a comparison of the performance of regularized regression, ensemble, instance-based and deep learning methods on synthetic and empirical data
BMC Genomics (2024)
-
A multimodal graph neural network framework for cancer molecular subtype classification
BMC Bioinformatics (2024)
-
Antimicrobial resistance crisis: could artificial intelligence be the solution?
Military Medical Research (2024)