Abstract
Parasitic nematodes (roundworms) and platyhelminths (flatworms) cause debilitating chronic infections of humans and animals, decimate crop production and are a major impediment to socioeconomic development. Here we report a broad comparative study of 81 genomes of parasitic and non-parasitic worms. We have identified gene family births and hundreds of expanded gene families at key nodes in the phylogeny that are relevant to parasitism. Examples include gene families that modulate host immune responses, enable parasite migration though host tissues or allow the parasite to feed. We reveal extensive lineage-specific differences in core metabolism and protein families historically targeted for drug development. From an in silico screen, we have identified and prioritized new potential drug targets and compounds for testing. This comparative genomics resource provides a much-needed boost for the research community to understand and combat parasitic worms.
Similar content being viewed by others
Main
Over a quarter of humans are infected with parasitic nematodes (roundworms) or platyhelminths (flatworms)1. Although rarely lethal, infections are typically chronic, leading to pain, malnutrition, physical disabilities, delayed development, deformity, social stigma or a burden on family members caring for the afflicted. These diseases encompass many of the most neglected tropical diseases and attract little research investment. Parasitic nematodes and platyhelminths impede economic development through human disability, and billions of dollars of lost production in the livestock2 and crop3 industries.
Few drugs are available to treat worm infections. Repeated mass administration of monotherapies is increasing the risk of resistance to human anthelmintics4 and has driven widespread resistance in farm animals5. There are no vaccines for humans, and few for animals6. The commonly used nematicides of plant parasites are environmentally toxic7, and need replacement.
The phylum Nematoda is part of the superphylum Ecdysozoa and has five major clades (I to V), four of which contain human-infective parasites and are analyzed here (Fig. 1). The phylum Platyhelminthes is part of the superphylum Lophotrochozoa and the majority of parasite species are cestodes (tapeworms) and trematodes (flukes). Comparing the genomes of parasites from these two phyla may reveal common strategies employed to subvert host defenses and drive disease processes.
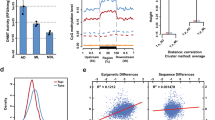
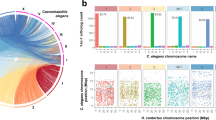
a, Maximum-likelihood phylogeny based on a partitioned analysis of a concatenated data matrix of 21,649 amino acid sites from 202 single-copy orthologous proteins present in at least 23 of the species. Values on marked nodes are bootstrap support values; all unmarked nodes were supported by 100 bootstrap replicates; nodes with solid marks were constrained in the analysis. Bar plots show genome sizes and total lengths of different genome features, and normalized gene count (Supplementary Note 1.2) for proteins with inferred functions based on sequence similarity (having an assigned protein name; Methods), or those without (named ‘hypothetical protein’). Species for which we have sequenced genomes are marked with asterisks; 33 ‘tier 1’ genomes are in black. LTR, long terminal repeat; LINE, long interspersed nuclear element. b, Assembly statistics. Blue rows indicate the 33 ‘tier 1’ genomes. Asterisks indicate the species for which we have sequenced genomes.
We have combined 36 published genomes8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34 with new assemblies for 31 nematode and 14 platyhelminth species into a large genome comparison of parasitic and non-parasitic worms. We have used these data to identify gene families and processes associated with the major parasitic groups. To accelerate the search for new interventions, we have mined the dataset of more than 1.4 million genes to predict new drug targets and drugs.
Results
Genomic diversity in parasitic nematodes and platyhelminths
We have produced draft genomes for 45 nematode and platyhelminth species and predicted 0.8 million protein-coding genes, with 9,132–17,274 genes per species (5–95% percentile range; see Methods, Supplementary Tables 1–3, Supplementary Fig. 1 and Supplementary Notes 1.1 and 1.2). We combined these new data with 36 published worm genomes—comprising 31 parasitic8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30 and five free-living18,31,32,33,34 species—and 10 outgroups35,36,37,38,39,40,41,42,43,44 from other animal phyla, into a comparative genomics resource of 91 species (Fig. 1 and Supplementary Tables 2 and 4). There was relatively little variation in gene set completeness (coefficient of variation, c.v. = 0.15) among the nematodes and platyhelminths, despite variation in assembly contiguity (c.v. = 8.5; Fig. 1b and Supplementary Table 2). Nevertheless, findings made using a subset of high-quality assemblies that were designated ‘tier 1’ (Methods and Supplementary Table 4) were corroborated against all species.
Genome size varied greatly within each phylum, from 42 to 700 Mb in nematodes, and from 104 to 1,259 Mb in platyhelminths. In a small number of cases, size estimates may have been artifactually inflated by high heterozygosity causing alternative haplotypes to be represented within the assemblies (Supplementary Note 1.3 and Supplementary Table 2a). A more important factor appeared to be repeat content that ranged widely, from 3.8 to 54.5% (5–95% percentile; Supplementary Table 5). A multiple regression model, built to rank the major factors driving genome size variation, identified long terminal repeat transposons, simple repeats, assembly quality, DNA transposable elements, total length of introns and low complexity sequence as being the most important (Supplementary Note 1.3, Methods and Supplementary Table 6). Genome size variation is thus largely due to non-coding elements, as expected45, including repetitive and non-repetitive DNA, suggesting it is either non-adaptive or responding to selection only at the level of overall genome size.
Gene family births and expansions
We inferred gene families from the predicted proteomes of the 91 species using Ensembl Compara46. Of the 1.6 million proteins, 1.4 million were placed into 108,351 families (Supplementary Note 2.1 and Supplementary Data), for which phylogenetic trees were built and orthology and paralogy inferred (Methods, Supplementary Fig. 2 and Supplementary Table 7). Species trees inferred from 202 single-copy gene families that were present in at least 25% of species (Fig. 1), or from presence/absence of gene families, largely agreed with the expected species and clade relationships, except for a couple of known contentious issues (Supplementary Fig. 3, Supplementary Note 2.2 and Methods).
The species in our dataset contained significant novelty in gene content. For example, ~28,000 parasitic nematode gene families contained members from two or more parasitic species but were absent from Caenorhabditis elegans and 47% of gene families lacked any functional annotation (Supplementary Note 2.1 and Methods). The latter families tended to be smaller than those with annotations (Supplementary Fig. 4) and, in many cases, correspond to families that are so highly diverged that ancestry cannot be traced, reflecting the huge breadth of unexplored parasite biology.
Gene families specific to particular parasite clades are likely to reflect important aspects of parasite biology and possible targets for new antiparasitic interventions. At key nodes in the phylogeny that are relevant to parasitism, we identified 5,881 families with apparent clade-specificity (synapomorphies; Supplementary Note 2.3, Methods and Supplementary Table 8), although our ability to discriminate truly parasite-specific clades was limited by the low number of free-living species. The apparent synapomorphies were either gene family births, or subfamilies that were so diverged from their homologues that they appeared as separate families. Functional annotation of these families was diverse (Fig. 2), but they were frequently associated with sensory perception (such as G-protein coupled receptors; GPCRs), parasite surfaces (platyhelminth tegument or nematode cuticle maintenance proteins) and protein degradation (proteases and protease inhibitors).
a, Rectangular matrices indicate counts of synapomorphic families grouped by 18 functional categories, detailed in the top left corner. Representative functional annotation of a family was inferred if more than 90% of the species present contained at least one gene with a particular domain. The node in the tree to which a panel refers is indicated in each matrix. ‘Other’ indicates families with functional annotation that could not be grouped into one of the 18 categories. ‘None’ indicates families that had no representative functional annotation. b, Expansions of apyrase and PUMA gene families. Families were defined using Compara. For color key and species labels, see Fig. 1. The plot for a family shows the gene count in each species, superimposed on the species tree. A scale bar beside the plot for a family shows the minimum, median and maximum gene count across the species, for that family.
Among nematodes, clade IVa (which includes Strongyloides spp.; Fig. 1) showed the highest number of clade-specific families, including a novel ferrochelatase-like family. Most nematodes lack functional ferrochelatases for the last step of haem biosynthesis47, but harbor ferrochelatase-like genes of unknown function, to which the synapomorphic clade IVa family was similar (Supplementary Fig. 5 and Methods). Exceptions are animal parasites in nematode clades III (for example ascarids and filaria) and IV that acquired a functional ferrochelatase via horizontal gene transfer48,49. Within the parasitic platyhelminths, a clade-specific inositol-pentakisphosphate 2-kinase (IP2K) was identified. In some species of Echinococcus tapeworms, IP2K produces inositol hexakisphosphate nanodeposits in the extracellular wall (the laminated layer) that protects larval metacestodes50. The deposits increase the surface area for adsorption of host proteins and may promote interactions with the host51.
Paralogous expansions of gene families, particularly those that are large or repeatedly involve related processes, can be evidence of adaptive evolution. We searched among our 10,986 highest-confidence gene families (those containing ≥10 genes from tier 1 species) for those that had expanded in parasite clades. A combination of scoring metrics (Methods) reduced the list to 995 differentially distributed families with a bias in copy number in at least one parasite clade. Twenty-five expansions have previously been observed, including 21 with possible roles in parasitism (Supplementary Fig. 6). A further 43 were placed into major functional classes that historically have been favored as drug targets (kinases, GPCRs, ion channels and proteases52; Supplementary Table 9a).
By manually inspecting the distribution of the remaining 927 families across the full species tree, we identified 176 families with striking expansions (Supplementary Table 9a and Supplementary Note 2.4). Thirty two had no functional annotation; for example, family 393312 was highly expanded in clade Va nematodes (Supplementary Fig. 7 and Supplementary Table 9a). Even when families could be functionally annotated to some extent (for example, based on a protein domain), discerning their precise biological role was a challenge. For example, a sulfotransferase family that was expanded in flukes compared with tapeworms includes the Schistosoma mansoni locus that is implicated in resistance to the drug oxamniquine53 but the endogenous substrate for this enzyme is unknown (Supplementary Fig. 7j).
Among the newly identified expansions, we focused on those with richer functional information, especially where they were related to similar biological processes. For instance, we identified several expansions of gene families involved in innate immunity of the parasites, as well as their development. These included families implicated in protection against bacterial or fungal infections in nematode clade IVa (bus-4 GT31 galactosyltransferase54, irg-355) and clades Va/Vc (lysozyme56 and the dual oxidase bli-357) (Supplementary Fig. 8a–d). In nematode clade IIIb, a family was expanded that contains orthologs of the Parascaris coiled-coiled protein PUMA, involved in kinetochore biology58 (Fig. 2b). This expansion possibly relates to the evolution of chromatin diminution in this clade, which results in an increased number of chromosomes requiring correct segregation during metaphase59. In nematode clade IVa and in Bursaphelenchus, an expansion of a steroid kinase family (Supplementary Fig. 8e) is suggestive of novelty in steroid-regulated processes in this group, such as the switch between free-living or parasitic stages in Strongyloides60.
Infections with parasitic worms are typified by their chronicity and a plausible involvement in host–parasite interactions is a recurring theme for many of the families. Taenia tapeworms and clade V strongylid nematodes (that is Va, Vb and Vc; Fig. 1) contained two expanded families with apyrase domains that may have a role in hydrolyzing ATP (a host danger signal) from damaged host tissue61 (Fig. 2b and Supplementary Fig. 9a). Moreover, many of the strongylid members also contained amine oxidoreductase domains, possibly to reduce production of pro-inflammatory amines, such as histamine, from host tissues62. In platyhelminths, we observed expansions of tetraspanin families that are likely components of the host/pathogen interface. Described examples show tetraspanins being part of extracellular vesicles released by helminths within hosts63; or binding the Fc domain of host antibodies64; or being highly immunogenic65 (Supplementary Fig. 9b,c). In strongylids, especially clade Vc, an expansion of the fatty acid and retinol-binding (FAR) family, implicated in host–parasite interaction of plant- and animal-parasitic nematodes66,67 (Supplementary Fig. 9d), suggests a role in immune modulation. Repertoires of glycosyl transferases have expanded in nematode clades Vc and IV, and tapeworms (Supplementary Fig. 10a–c), and may be used to evade or divert host immunity by modifying parasite surface molecules directly exposed to the immune system68; alternatively, surface glycoproteins may interact with lectin receptors on innate immune cells in an inhibitory manner69. An expanded chondroitin hydrolase family in nematode clade Vc may possibly be used either for larval migration through host connective tissue or to digest host intestinal walls (Supplementary Fig. 9e). Similarly, an expanded GH5 glycosyl hydrolase family contained schistosome members with egg-enriched expression8,70 that may be used for traversing host tissues such as bladder or intestinal walls (Supplementary Fig. 9f). In nematode clade I, we found an expansion of a family with the PAN/Apple domain, which is implicated in attachment of some protozoan parasites to host cells71, and possibly modulates host lectin-based immune activation (Supplementary Fig. 9g).
The SCP/TAPS (sperm-coating protein/Tpx/antigen 5/pathogenesis-related protein 1) genes have been associated with parasitism through their abundance, secretion and evidence of their role in immunomodulation72 but are poorly understood. This diverse superfamily appeared as eight expanded Compara families. A more comprehensive phylogenetic analysis of the full repertoire of 3,167 SCP/TAPS sequences (Supplementary Note 2.5, Supplementary Table 10 and Methods) revealed intra- and interspecific expansions and diversification over different evolutionary timescales (Fig. 3 and Supplementary Figs. 11a,b and 12). In particular, the SCP/TAPS superfamily has expanded independently in nematode clade V (18–381 copies in each species) and in clade IVa parasites (39–166 copies) (Fig. 3 and Supplementary Fig. 11c). Dracunculus medinensis (Guinea worm) was unusual in being the only member of clade III to display an expansion (66 copies), which may reflect modulation of the host immune response during the tissue migration phase of its large adult females.
A maximum-likelihood tree of SCP/TAPS genes. Colors represent different species groups. Homo sapiens GLIPR2 was used to root the tree. Blue dots show high bootstrap values (≥0.8). A clade was collapsed into a triangle if more than half its leaves were genes from the same species group. Nematode clade I had fewer counts, but was collapsed to show its relationship to other clades’ expansions. ‘Strongylid’ refers to clades Va, Vb and Vc.
Proteins historically targeted for drug development
Proteases, GPCRs, ion channels and kinases dominate the list of targets for existing drugs for human diseases52, and are attractive leads for developing new ones. We therefore explored the diversity of these superfamilies across the nematodes and platyhelminths (Supplementary Fig. 13, Supplementary Note 3 and Methods).
Proteases and protease inhibitors perform diverse functions in parasites, including immunomodulation, host tissue penetration, modification of the host environment (for example, anticoagulation) and digestion of blood73. M12 astacins have particularly expanded in nematode clade IVa (five families), as previously reported18, but there are two additional expansions in clades Vc and Vb (Fig. 4, Supplementary Fig. 14 and Supplementary Table 11). Because many of these species invade through skin (IVa, Vc; Supplementary Table 12) and migrate through the digestive system and lung (IVa, Vc, Vb; Supplementary Table 13), these expansions are consistent with evidence that astacins are involved in skin penetration and migration through connective tissue74. The cathepsin B C1-cysteine proteases are particularly expanded in species that feed on blood (two expansions in nematode clades Vc and Va30, with highest platyhelminth gene counts in schistosomatids and Fasciola12; Supplementary Fig. 14). Indeed, they are involved in blood digestion in adult nematodes75 and platyhelminths76, but some likely have different roles such as larval development77 and host invasion78.
Relative abundance profiles for 84 protease and 31 protease inhibitor families represented in at least 3 of the 81 nematode and platyhelminth species. Thirty-three protease families and 6 protease inhibitor families present in fewer than 3 species were omitted from the visualization. For each species, the gene count in a class was normalized by dividing by the total gene count for that species. Families mentioned in the Results or Supplementary Note text are labeled; complete annotations of all protease families are in Supplementary Table 11.
Different protease inhibitors may modulate activity of parasite proteases or protect parasitic nematodes and platyhelminths from degradation by host proteases, facilitate feeding or manipulate the host response to the parasite79. The I2 (Kunitz-BPTI) trypsin inhibitors are the most abundant protease inhibitors across parasitic nematodes and platyhelminths (Fig. 4). An expansion of the I17 family, which includes secretory leukocyte peptidase inhibitor, was reported previously in Trichuris muris17 but the striking confinement of this expansion to most of the parasites of clade I is now apparent (Fig. 4). We also observed a notable family of α-2-macroglobulin (I39) protease inhibitors that are present in all platyhelminths but expanded in tapeworms (Supplementary Fig. 14). The tapeworm α-2-macroglobulins may be involved in reducing blood clotting at attachment or feeding sites; alternatively, they may modulate the host immune response, since α-2-macroglobulins bind several cytokines and hormones80. Chymotrypsin/elastase inhibitors (family I8) were particularly expanded in clades Vc and IVa (consistent with upregulation of I8 genes in Strongyloides parasitic stages18) and to a lesser extent in clade IIIb (Fig. 4), consistent with evidence that they may protect Ascaris from host proteases81. We also identified protein domain combinations that were specific to either nematodes or platyhelminths (131 and 50 domain combinations, respectively). Many of these involved protease and protease inhibitor domains. In nematodes, several combinations included Kunitz protease inhibitor domains, and in platyhelminths metalloprotease families M18 and M28 were found in novel combinations (Supplementary Table 14, Supplementary Note 3.2 and Methods).
Of the 230 gene families annotated as GPCRs (Supplementary Figs. 13 and 15 and Supplementary Note 3.3), only 21 were conserved across phyla. Chemosensory GPCRs, while abundant in nematodes, were not identified in platyhelminths, although they are identifiable in other Lophotrochozoa (such as Mollusca82), suggesting that either the platyhelminths have lost this class or they are very divergent (Supplementary Table 15). GPCR families lacking sequence similarity with known receptors included the platyhelminth-specific rhodopsin-like orphan families (PROFs), which are likely to be class A receptors and peptide responsive, and several other fluke-specific non-PROF GPCR families. The massive radiation of chemoreceptors in C. elegans was unmatched in any other nematode (87% versus ≤48% of GPCRs). All parasitic nematodes possessed chemoreceptors, with the most in clade IVa, including several large families synapomorphic to this clade (Supplementary Fig. 15), perhaps related to their unusual life cycles that alternate between free-living and parasitic forms.
Independent expansion and functional divergence has differentiated the nematode and platyhelminth pentameric ligand gated ion channels (Supplementary Fig. 16, Supplementary Table 16 and Supplementary Note 3.4). For example, glutamate signaling arose independently in platyhelminths and nematodes83, and in trematodes the normal role of acetylcholine has been reversed, from activating to inhibitory84. Our analysis suggested the platyhelminth acetylcholine-gated anion channels are most related to the Acr-26/27 group of nematode nicotinic acetylcholine receptors that are the target of the anthelmintics morantel and pyrantel85, rather than to nematode acetylcholine-gated cation channels, targeted by nicotine and levamisole (Supplementary Fig. 17).
ABC transporters (Supplementary Table 17 and Supplementary Note 3.5) and kinases (Supplementary Note 3.6 and Supplementary Fig. 18) showed losses and independent expansion within nematodes and platyhelminths. The P-glycoprotein class of transporters, responsible for the transport of environmental toxins and linked with anthelmintic resistance, is expanded relative to vertebrates86, with increased numbers in nematodes (Supplementary Fig. 19).
Metabolic reconstructions of nematodes and platyhelminths
In the context of drug discovery, understanding the metabolic capabilities of parasitic worms may reveal vulnerabilities that can be exploited in target-based screens for new compounds. For each of the 81 nematode and platyhelminth species, metabolism was reconstructed based on high confidence assignment of enzyme classes (Supplementary Table 18a). The nematodes had a greater range of annotated enzymes per species than the platyhelminths (Supplementary Fig. 20a), in part reflecting the paucity of biochemical studies in platyhelminths. Because variation in assembly quality or divergence from model organisms87 could bias enzyme predictions, we identified losses of pathways and differences in pathway coverage across different clades (Supplementary Note 4, Methods, Fig. 5 and Supplementary Fig. 21). Pathways related to almost all metabolic superpathways in the Kyoto Encyclopedia of Genes and Genomes (KEGG)88 showed significantly lower coverage for platyhelminths (versus nematodes) and filaria (versus other nematodes) (Supplementary Fig. 20b).
a, Topology-based detection of KEGG metabolic modules among tier 1 species (dark green, present; light green, largely present (only one enzyme not found)). Only modules detected to be complete in at least one species are shown. The EC annotations used for this figure included those from pathway hole-filling and those based on Compara families (Supplementary Table 18a, b). b, Biochemical pathways that appear to have been completely or partially lost from certain platyhelminth and nematode clades. PRPP, phosphoribosyl pyrophosphate.
In contrast to most animals, nematodes possess the glyoxylate cycle that enables conversion of lipids to carbohydrates, to be used for biosyntheses (for example, during early development) and to avert starvation89. The glyoxylate cycle appears to have been lost independently in the filaria and Trichinella species (Fig. 5a; M00012), both of which are tissue-dwelling obligate parasites. The filaria and Trichinella have also independently lost alanine-glyoxylate transaminase that converts glyoxylate to glycine (Fig. 5b). Glycine can be converted by the glycine cleavage system (GCS) to 5,10-methylenetetrahydrofolate, a useful one-carbon pool for biosyntheses, and two key GCS proteins appear to have been lost independently from filaria and tapeworms, suggesting their GCS is non-functional (Supplementary Table 19e). In addition, filaria have lost the ability to produce and use ketone bodies, a temporary store of acetyl coenzyme A (CoA) under starvation conditions (Supplementary Table 19b). The filaria lost these features after they diverged from D. medinensis, an outgroup to the filaria in clade IIIc that has a major difference in its life cycle, namely, a free-living larval stage (Supplementary Table 12).
The absence of multiple initial steps of pyrimidine synthesis was observed in some nematodes, including all filaria (as previously reported23) and tapeworms, suggesting they obtain pyrimidines from Wolbachia endosymbionts or from their hosts, respectively (Supplementary Table 19f). Similarly, all platyhelminths and some nematodes (especially clade IVa and filaria IIIc) appear to lack key enzymes for purine synthesis (Supplementary Table 19g) and rely on salvage instead. However, despite the widespread belief that nematodes cannot synthesize purines90,91, complete or near-complete purine synthesis pathways were found in most members of clades I, IIIb and V. Nematodes are known to be unable to synthesize haem47 but the pathway was found in platyhelminths, including S. mansoni (despite conflicting biochemical data47) (Supplementary Table 19h and Supplementary Table 20i).
Genes from the β-oxidation pathway, used to break down lipids as an energy source, were not detected in schistosomes and some cyclophyllidean tapeworms (Hymenolepis, Echinococcus; Fig. 5a, M00087; Supplementary Table 19a). These species live in glucose-rich environments and may have evolved to use glucose and glycogen as principal energy sources. However, biochemical data suggest they do perform β-oxidation92, so they may have highly diverged but functional β-oxidation genes.
The lactate dehydrogenase (LDH) pathway is a major source of ATP in anaerobic but glucose-rich environments. Platyhelminths have high numbers of LDH genes, as do blood-feeding Ancylostoma hookworms (Supplementary Fig. 22g). Nematode clades Vc (including Ancylostoma) and IIIb have expansions of α-glucosidases that may break down starch and disaccharides in host food to glucose (Supplementary Fig. 22a). Many nematodes and flatworms use malate dismutation as an alternative pathway for anaerobic ATP production93. The importance of the pathway for clade IIIb nematodes was reflected in expanded families encoding two key pathway enzymes PEPCK and methylmalonyl CoA epimerase, and the intracellular trafficking chaperone for cobalamin (vitamin B-12), a cofactor for the pathway (Supplementary Fig. 22c–e and Supplementary Table 9a). A second cobalamin-related family (CobQ/CbiP) is clade IIIb-specific and appears to have been gained by horizontal gene transfer from bacteria (Supplementary Fig. 23a, Supplementary Note 2.6 and Methods). A glutamate dehydrogenase family expanded in clade IIIb (Supplementary Fig. 22h) is consistent with a GABA (γ-aminobutyric acid) shunt that helps maintain redox balance during malate dismutation. In clade Va, an expansion in the propionate breakdown pathway94 (Supplementary Fig. 22f), suggested degradation of propionate, originating from malate dismutation or fermentation in the host’s stomach95. Clade I nematodes have an acetate/succinate transporter that appeared to have been gained from bacteria (Supplementary Note 2.6 and Methods), and may participate in acetate/succinate uptake or efflux (Supplementary Fig. 23b).
Identifying new anthelmintic drug targets and drugs
As an alternative to a purely target-based approach that would require extensive compound screening, we explored drug repurposing possibilities. We developed a pipeline to identify the most promising targets from parasitic nematodes and platyhelminths. These sequences were used in searches of the ChEMBL database that contains curated activity data on defined targets in other species and their associated drugs and compounds (Supplementary Note 5 and Methods). Our pipeline identified compounds that are predicted to interact with the top 15% of highest-scoring worm targets (n = 289). These targets included 17 out of 19 known or likely targets for World Health Organization-listed anthelmintics that are represented in ChEMBL (Supplementary Table 21b). When compounds within a single chemical class were collapsed to one representative, this potential screening set contained 5,046 drug-like compounds, including 817 drugs with phase III or IV approval and 4,229 medicinal chemistry compounds (Supplementary Table 21d). We used a self-organizing map to cluster these compounds based on their molecular fingerprints (Fig. 6). This classification showed that the screening set was significantly more structurally diverse than existing anthelmintic compounds (Supplementary Fig. 24).
A self-organizing map clustering known anthelmintic compounds (Supplementary Table 21a) and our proposed screening set of 5,046 compounds. The density of red and green shows the number of screening set and known anthelmintic compounds clustered in each cell, respectively. Structures for representative known anthelmintic compounds are shown at the top, and examples from the proposed screening set along the bottom.
The 289 targets were further reduced to 40 high-priority targets, based on predicted selectivity, avoidance of side-effects (clade-specific chokepoints or lack of human homologues) and putative vulnerabilities, such as those suggested by gene family expansions in parasite lineages, or belonging to pathways containing known or likely anthelmintic targets (Supplementary Fig. 25). These 40 targets were associated with 720 drug-like compounds comprising 181 phase III/IV drugs and 539 medicinal chemistry compounds. There is independent evidence that some of these have anthelmintic activity. For example, we identified several compounds that potentially target glycogen phosphorylase, which is in the same pathway as a likely anthelmintic target (glycogen phosphorylase phosphatase, likely target of niridazole; Supplementary Fig. 25). These compounds included the phase III drug alvocidib (flavopiridol), which has anthelmintic activity against C. elegans96. Another example is the target cathepsin B, expanded in nematode clade Va (Supplementary Table 9a), for which we identified several compounds including the phase III drug odanacatib, which has been shown to have anthelmintic activity against hookworms97. Existing drugs such as these are attractive candidates for repurposing and fast-track therapy development, while the medicinal chemistry compounds provide a starting point for broader anthelmintic screening.
Discussion
The evolution of parasitism in nematodes and platyhelminths occurred independently, starting from different ancestral gene sets and physiologies. Despite this, common selective pressures of adaptation to host gut, blood or tissue environments, the need to avoid hosts’ immune systems, and the acquisition of complex life cycles to effect transmission, may have driven adaptations in common biological pathways. While previous comparative analyses of parasitic worms have been limited to a small number of species within narrow clades, we have surveyed parasitic worms spanning two phyla, with a focus on those infecting humans and livestock. A large body of draft genome data (both published and unpublished) was utilized but, by focusing on lineage-specific trends rather than individual species-specific differences, our analysis was deliberately conservative. In particular, we have focused on large gene family expansions, supported by the best-quality data and for which functional information was available. Sequencing of further free-living species, better functional characterization, and identification of remote orthologs (particularly for platyhelminths87), will undoubtedly refine the resolution of parasite-specific differences, but our gene family analyses have already revealed expansions and synapomorphies in functional classes of likely importance to parasitism, such as feeding and interaction with hosts. We have used a drug repurposing approach to predict potential new anthelmintic drug targets and drugs/drug-like compounds that we urge the community to explore. Further new potential drug targets, worthy of high-throughput compound screening, may be exposed by the losses of key metabolic pathways and horizontally acquired genes that we find in particular parasite groups. This is an unprecedented dataset of parasitic worm genomes that provides a new type of pan-species reference and a much needed stimulus to the study of parasitic worm biology.
URLs
SMALT, http://www.sanger.ac.uk/science/tools/smalt-0; RepeatModeler, http://www.repeatmasker.org/RepeatModeler.html; TransposonPSI, http://transposonpsi.sourceforge.net; RepeatMasker, http://www.repeatmasker.org; code for calculating gene family metrics, http://tinyurl.com/comparaFamiliesAnalysis-py; WormBase ParaSite, https://parasite.wormbase.org/.
Methods
Sample collection and preparation
Sources of material and sequencing approaches are summarized in Supplementary Table 1.
Wellcome Sanger Institute (WSI) data production
The genomes of 36 species (Supplementary Tables 1 and 2) were sequenced at WSI. The C. elegans N2 was also resequenced at WSI.
WSI sequencing and assembly
PCR-free 400–550 bp paired-end Illumina libraries were prepared from <0.1 ng to 5 µg genomic DNA, as described for Strongyloides stercoralis18. Where there was insufficient DNA, adapter-ligated material was subjected to ~8 PCR cycles.
We used 1–10 μg gDNA or whole genome amplification DNA to generate 3 kb mate-pair libraries, as described for S. stercoralis18. If there was insufficient gDNA, whole genome amplification was performed using GenomiPhi v2. Each library was run on ≥1 Illumina HiSeq 2000 lane.
Short insert paired-end reads were corrected and assembled with SGA v0.9.798 (Supplementary Fig. 26a). This assembly was used to calculate the k-mer distribution for all odd k of 41–81, using GenomeTools v.1.3.799. The k-mer length for which the maximum number of unique k-mers was present was used as the k-mer setting in a second assembly, using Velvet v1.2.03100 with SGA-corrected reads. For species with 3 kb mate-pair data, the Velvet assembly was scaffolded using SSPACE101. Contigs were extended, and gaps closed and shortened, using Gapfiller102 and IMAGE103. Short fragment reads were remapped to the assembly using SMALT (see URLs), and unaligned reads assembled using Velvet100 and this merged with the main assembly. The assembly was re-scaffolded using SSPACE101, and consensus base quality improved with iCORN104. REAPR105 was used to break incorrectly assembled scaffolds/contigs. We carried out manual improvement for Wuchereria bancrofti and D. medinensis using Gap5106 and Illumina read-pairs.
WSI assembly quality control
Contamination screening. Assemblies were screened for contamination using BLAST107 against vertebrate and invertebrate sequences (see ref. 108). For Anisakis simplex, the assembly contained minor laboratory contamination with S. mansoni, which we removed using BLASTN against S. mansoni.
Assembly completeness. CEGMA v2.4109 was used to assess completeness. Consistent sets of CEGMA genes were missing from some phylogenetic groups (Supplementary Table 2); these were discounted from the completeness calculation for those species (‘CEGMA’ in Supplementary Table 2).
Effect of repeats. We re-mapped the short-insert library’s reads to the appropriate assembly using SMALT (see URLs; indexing -k13 -s4 and mapping -y 0.9 -x -r 1). For each scaffold of ≥8 kb, median (meds) and mean (ms) per-base read-depth were calculated using BEDTools110, and genome-wide depth (medg) calculated as the median meds (ref. 17). For a ls bp scaffold, the extra sequence that would be gained by ‘uncollapsing’ repeats was estimated as es = (ms − medg) × ls/medg (Supplementary Table 5).
WSI gene prediction
Our pipeline111 had four steps (Supplementary Fig. 27a). First, repeats were masked. Second, preliminary gene predictions, to use as input for MAKER v2.2.28112 were generated using Augustus 2.5.5113, SNAP 2013-02-16114, GeneMark-ES 2.3a115, genBlastG116 and RATT117. Third, species-specific ESTs and complementary DNAs from INSDC118, and proteins from related species, were aligned to the genome using BLAST107. Last, EST/protein alignments and gene models were used by MAKER to produce a gene set.
McDonnell Genome Institute (MGI) data production
The genomes of six species were sequenced at MGI (Supplementary Tables 1 and 2).
MGI sequencing, assembly and quality control
Genome sequencing was carried out on Illumina and 454 instruments (see ref. 119). The workflow for each assembly is in Supplementary Table 1.
Three kilobase, 8 kb and fragment 454 reads (or Illumina reads) were subject to adapter removal, quality trimming and length filtering (Supplementary Fig. 26b). Cleaned 454 reads were assembled using Newbler120 before being scaffolded with an in-house tool CIGA, which links contigs based on cDNA evidence. Cleaned Illumina reads were assembled using AllPaths-LG121. The assembly was scaffolded further using an in-house tool Pygap, using Illumina short paired-end sequences; and L_RNA_scaffolder122, using 454 cDNA data.
An assisted assembly approach was used for Trichinella nativa, whereby ‘cleaned’ Illumina 3 kb paired-end sequence data were mapped against the T. spiralis genome using bwa123 (Supplementary Fig. 26b), and the T. nativa residues were substituted at aligned positions (see ref. 119).
Adaptor sequences and contaminants were identified by comparison to a database of vectors and contaminants, using Megablast124.
MGI transcriptome sequencing and gene prediction
Transcriptome libraries (Supplementary Table 22) were generated with the Illumina TS stranded protocol, and reads assembled using Trinity125 (see ref. 119).
Genes were predicted using MAKER112, based on input gene models from SNAP114, FGENESH (Softberry), Augustus113, and aligned messenger RNA, EST, transcriptome and protein data from the same or related species (Supplementary Fig. 27b; see ref. 119).
Blaxter Nematode and Neglected Genomics (BaNG) data production
The genomes of three species were sequenced by BaNG (Supplementary Tables 1 and 2).
Sequencing was performed on Illumina HiSeq 2000 and HiSeq 2500 instruments, using 100 or 125 base, paired-end protocols. Paired-end libraries were generated using the Illumina TruSeq protocol.
Sequence data were filtered of contaminating host reads using blobtools126. Cleaned reads were normalized with the khmer software127 using a k-mer of 41, and then assembled with ABySS (v1.3.3)128, with a minimum of three pairs needed to connect contigs during scaffolding (n = 3) (Supplementary Fig. 26c). Assemblies were assessed using blobtools and CEGMA109.
Augustus113 was used to predict gene models, trained using annotations from MAKER112. As hints for MAKER, we used Litomosoides sigmodontis 454 RNA sequencing data assembled with MIRA129 and Newbler120, and Onchocerca ochengi Illumina RNA sequencing data130 assembled using Trinity131 (Supplementary Fig. 27c).
Defining high-quality ‘tier 1’ species
A subset of nematode and platyhelminth genomes, termed ‘tier 1’, was selected that had better-quality assemblies and spanned the major clades (Supplementary Table 4). To choose these, species were selected that (1) had contiguous assemblies (usually N50/scaffold-count >5), and complete proteomes (usually CEGMA partial >85%), or (2) that helped to ensure ~50% of the genera in each species group (‘Analysis group’ in Supplementary Table 4) were represented.
Analysis of repeat content and genome size
For each species, repeat libraries were built using RepeatModeler (see URLs), TransposonPSI (see URLs) and LTRharvest132, and the three libraries merged (see ref. 133). The merged library was used to mask repeats in a species’ genome using RepeatMasker (see URLs; –s).
The initial standard regression model and stepwise model fitting used ‘lm’ and ‘step’ in R v3.2.2. The Bayesian mixed-effect model used MCMCglmm134 (v2.24). To create a mixed-effect model, the species tree (see Methods) was transformed into an ultrametric tree using PATHd8135, with a small constant added to short branches to ensure no zero-length branches were reconstructed; and outgroup species were removed.
Compara database
An in-house Ensembl Compara46 database was constructed containing the 81 platyhelminths and nematodes, and 10 additional outgroups (Supplementary Table 2). All parasitic nematode/platyhelminth species with gene sets available at the time (April 2014) were included.
The species tree used to construct the initial version of our database used an edited version of the National Center for Biotechnology Information (NCBI) taxonomy136 with several controversial speciation nodes represented as multifurcations. For our final database, the input species tree was derived by building a tree based on the previous database version, based on one-to-one orthologs present in ≥20 species. To do this, proteins in each ortholog group were aligned using MAFFT v6.857137; alignments trimmed using GBlocks v0.91b138, concatenated and used to build a maximum likelihood tree using a partitioned analysis in RAxML v7.8.6139, using the minimum Akaike’s information criterion (minAIC) model for each ortholog group.
The database was queried to identify gene families, orthologs and paralogs.
Species tree and tree based on gene family presence
We identified 202 gene families present in ≥25% of the 91 species (81 helminths and 10 outgroups) in our Compara database (Methods) and always single-copy. For each family, amino acid sequences were aligned using MAFFT v7.205137 (-auto). Each alignment was trimmed using GBlocks v0.91b138 (-b4 = 4 -b3 = 4 -b5 = h), and its likelihood calculated on a maximum-parsimony guide tree for all relatively simple (single-matrix) amino acid substitution models in RAxML v8.0.24139, and the minAIC model identified. Alignments were concatenated and a maximum-likelihood tree built, under a partitioned model in which sites from a gene were assigned the minAIC model for that gene, with a discrete gamma distribution of rates across sites. Relationships within outgroup lineages were constrained to match the standard view of metazoan relationships (for example, Dunn et al.140). The final tree was the highest likelihood one from five search replicates with different random number seeds. One hundred bootstrap resampling replicates were performed, each based on a single rapid search.
We also constructed a maximum-likelihood phylogeny based on gene family presence/absence for families not shared by all 81 nematode/platyhelminth species, using RAxML v8.2.8139, with a two-state model and the Lewis method to correct for absence of constant-state observations.
Functional annotation
InterProScan141 v5.0.7 was used to identify conserved domains from all predicted proteins. A name was assigned to each predicted protein based on curated information in UniProt142 for orthologs identified from our Compara database (Methods), or based on InterPro143 domains (see ref. 144). Gene ontology (GO) terms were assigned by transferring GO terms from orthologs144, and using InterProScan.
Signal peptides and transmembrane domains were predicted using Phobius145 v1.01 and SecretomeP146 v1.0. A protein predicted by Phobius to have a transmembrane domain was categorized as ‘membrane-bound’, and non-membrane-bound proteins as ‘classically secreted’ if Phobius predicted a signal peptide within 70 amino acids of their start. Remaining proteins in which SecretomeP predicted a signal peptide were classified as ‘non-classically secreted’ (Supplementary Table 7).
Pairwise combinations of Pfam domains were identified in proteins of the 81 nematodes and platyhelminths. After excluding those present in complete genomes of other phyla in UniProt (June 2016), we classified a combination as ‘nematode-specific’ (or ‘flatworm-specific’) if it was present in >30% of nematodes (platyhelminths) and no platyhelminths (nematodes) (Supplementary Table 14).
Synapomorphic gene families
Families in our Compara database (Methods) were analyzed using KinFin v0.8.3147, by providing InterPro IDs (Methods) and a species tree that had clades III, IV and V as a polytomy (Fig. 2). Synapomorphic families were identified at 25 nodes of interest (Supplementary Table 8), by using Dollo parsimony and requiring a family must contain genes from ≥1 descendant species from each child node of the node of interest, and must not contain other species. Families were filtered to retain those that (1) contained ≥90% of descendant species of the node of interest, and (2) in which >90% of species contained ≥1 gene with a particular InterPro domain.
Candidate lateral gene transfers
Ferrochelatase families in our Compara database (Methods) were extracted by screening for a Ferrochelatase (IPR001015) domain. Additional ferrochelatases were retrieved from NCBI for 17 bacterial taxa (Supplementary Table 8c). Sequences were aligned using MAFFT v7.267 (E-INS-i algorithm)137 and the alignment trimmed using trimAl v1.4148. Phylogenetic analysis was carried out using RAxML139 under the PROTGAMMAGTR model, and 20 alternative runs on distinct starting trees. Non-parametric bootstrap analysis was carried out for 100 replicates.
For cobyric acid synthase and acetate/succinate transporter, the top BLAST hits from GenBank, and representative sequences from other taxonomic groups, were aligned with MAFFT v7.205137 (-auto), and alignments trimmed with trimal v1.4148. Phylogenetic analyses were performed using RAxML v8.2.8139 under the model that minimized the AIC (LG4X for cobyric acid synthase, LG4M for acetate transporter), based on 5 random-addition-sequence replicates, and 100 non-parametric bootstrap replicates.
Gene family expansions
We used three metrics to identify families in our Compara database (Methods) that varied greatly in gene count across species (see ref. 149). To control for fragmented assemblies, we used summed protein length per species (in a family) as a proxy for gene count in these metrics:
1. Coefficient of variation:
where s is the standard deviation in summed protein length per species, and \(\bar x\) its mean.
2. Maximum Z-score:
where T is the set of non-overlapping species groups (‘Analysis group’ in Supplementary Table 4), c a group in T, index i refers to a particular species, \(\bar x_{i, i \in c}\) the mean of the summed protein length (per species) in c, and \(s_{i, i \notin c}\) the standard deviation in summed protein length per species in species outside c.
3. Maximum enrichment coefficient:
To increase reliability, these metrics were calculated by only considering tier 1 species (those with high-quality assemblies; Methods). Our code for calculating metrics is available (see URLs).
SCP/TAPS
SCP/TAPS genes were identified as having Pfam PF00188, or being in a SCP/TAPS family in our Compara database (Methods). Those between 146 aa (shortest C. elegans SCP/TAPS) and 1,000 aa were included in the phylogenetic analysis (Supplementary Table 10). Clusters were detected among sequences from a species group (‘analysis group’ in Supplementary Table 4) using USEARCH150 (UCLUST, aa identity cut-off = 0.70), and a consensus sequence generated for each cluster. The consensus sequences were aligned using MAFFT137 (v7.271, –localpair –maxiterate 2 –retree 1 –bl 45); the alignment trimmed with trimAl148 (-gt 0.006); and a maximum likelihood tree built using FastTreeMP151 (v2.1.7 SSE3, -wag -gamma).
Proteins historically targeted for drug development
Each nematode/platyhelminth proteome was searched against candidate proteases using MEROPS batch-BLAST152 (E < 0.001), and PfamScan153 was used to identify additional homologues in some species (Supplementary Table 11).
Putative GPCRs, identified from the literature and GO:0004930 annotations in WormBase154, were used to identify families in our Compara database (Methods). For each family, HHSuite155 was used to search Uniprot, SCOPUS, Pfam, and PDB; 200 families hitting ≥2 databases were deemed actual GPCR families (see ref. 156). Additional families were identified from synapomorphies (Methods) and curation, giving 230 GPCR families (Supplementary Table 15).
To build a phylogenetic tree of ion channels, known genes from C. elegans157, Brugia malayi158, Haemonchus contortus159, Oesophagostomum dentatum159 and S. mansoni84 were gathered, and their homologues in Compara families in WormBase ParaSite160. Genes with <3 or >8 transmembrane domains (predicted by HMMTOP161) were discarded. Genes were aligned with MAFFT137, and the alignment trimmed with trimAl148. The phylogeny was inferred with MrBayes3.2162. Posterior probabilities were calculated from eight reversible jump Markov chain Monte Carlo chains over 20,000,000 generations.
Kinase models were taken from Kinomer163, and thresholds optimized to detect known C. elegans kinases (see ref. 164). The final thresholds were used to filter HMMER search results (against Kinomer) for nematode and platyhelminth species (Supplementary Table 23).
C. elegans ABC transporter and cys-loop receptor subunit genes were collated from WormBase154, to which we added H. contortus acr-26 and acr-27 (absent from C. elegans85). Homologs in nematodes and platyhelminths were identified using BLASTP (Supplementary Tables 16 and 17).
GO and InterPro/Pfam annotation enrichment
Counts of proteins annotated with each GO term (or InterPro/Pfam domain) per species were normalized by dividing by the total GO annotations in a particular species. To test for enrichment of a particular GO term in a species group (‘analysis group’ in Supplementary Table 4), we used a Mann-Whitney U test to compare normalized counts in that species group, to those in all other species (Supplementary Table 24).
Metabolism
EC (Enzyme Commission number) predictions for nematodes and platyhelminths were derived by combining DETECT v2.0165, PRIAM166, KAAS167 and BRENDA168 (see ref. 169, Supplementary Fig. 28 and Supplementary Table 18), and supplemented for the 33 tier 1 species (Methods) by pathway hole-filling using Pathway Tools170 (v18.5). Comparisons of all 81 species (Supplementary Fig. 20a and Supplementary Table 20) did not include ECs from hole-filling. Lower confidence ECs were inferred using families from our Compara database (Methods). Auxotrophies were predicted using Pathway Tools and BioCyc171. To predict carbohydrate-active enzymes, HMMER3 was used to search dbCAN172 (Supplementary Table 25).
Pathway coverage was the fraction of ECs in a reference pathway that were annotated in a species (see ref. 173). We included pathways for which KEGG had a reference pathway for a nematode/platyhelminth (Supplementary Table 18e). Presence of KEGG modules was predicted using modDFS174, and species clustered based on module presence using Ward-linkage, based on Jaccard similarity index175.
Chokepoint enzymes were predicted following Taylor et al.176, using subnetworks of KEGG networks formed by just the enzymes (ECs) we had annotated in each particular species.
Potential anthelmintic drug targets and drugs
Potential drug targets
Nematode and platyhelminth proteins from tier 1 species (with high-quality assemblies; Methods) were searched against single-protein targets from ChEMBL v21177 using BLASTP (E ≤ 1 × 10−10). After collapsing by gene family, 1,925 worm genes remained.
To assign a ‘target score’ to each worm gene, the main factors considered were similarity to known drug targets; lack of human homologues; and whether C. elegans/Drosophila melanogaster homologues had lethal phenotypes (see ref. 178).
Potential new anthelmintic drugs
ChEMBL v21177 was used to identify 827,889 compounds with activities against ChEMBL targets to which worm proteins had BLAST matches. To calculate ‘compound scores’, we prioritized compounds in high clinical development phases, oral/topical administration, crystal structures, properties consistent with oral drugs and lacking toxicity (see ref. 178).
Our top 15% (249) of highest-scoring worm targets had 292,499 compounds. These were filtered by selecting compounds that (1) co-appeared in a PDBe179 (Protein Data Bank in Europe) structure with the ChEMBL target; or (2) had median pChEMBL > 5; leaving 131,452 ‘top drug candidates’.
A ‘diverse screening set’
The 131,452 candidates were placed into 27,944 chemical classes, based on ECFP4 fingerprints (see ref. 178). They were filtered by (1) discarding medicinal chemistry compounds that did not co-appear in a PDBe structure with the ChEMBL target, or have median pCHEMBL > 7; (2) checking availability for purchase in ZINC 15180; and (3) for each worm target, taking the highest-scoring compound from each class; this gave 5,046 compounds.
Self-organizing map
We constructed a self-organizing map of our diverse screening set plus known anthelmintic compounds (Supplementary Table 21a; see ref. 178), using Kohonen v3.02181 in R v3.3.0, using a 20 × 20 cell hexagonal, non-toroidal grid. The self-organizing map was trained for 4,000 steps, where training optimized Tanimoto distances between ECFP4 fingerprints.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Sequence data have been deposited in the European Nucleotide Archive (ENA). Assemblies and annotation are available at WormBase and WormBase-ParaSite (https://parasite.wormbase.org/). All have been submitted to GenBank under the BioProject IDs listed in Supplementary Table 1.
References
G.B.D. 2015 Disease and Injury Incidence and Prevalence Collaborators. Global, regional, and national incidence, prevalence, and years lived with disability for 310 diseases and injuries, 1990-2015: a systematic analysis for the Global Burden of Disease Study 2015. Lancet 388, 1545–1602 (2016).
Charlier, J., van der Voort, M., Kenyon, F., Skuce, P. & Vercruysse, J. Chasing helminths and their economic impact on farmed ruminants. Trends Parasitol. 30, 361–367 (2014).
Jones, J. T. et al. Top 10 plant-parasitic nematodes in molecular plant pathology. Mol. Plant Pathol. 14, 946–961 (2013).
Furtado, L. F., de Paiva Bello, A. C. & Rabelo, E. M. Benzimidazole resistance in helminths: From problem to diagnosis. Acta Trop. 162, 95–102 (2016).
Kaplan, R. M. & Vidyashankar, A. N. An inconvenient truth: global worming and anthelmintic resistance. Vet. Parasitol. 186, 70–78 (2012).
Hewitson, J. P. & Maizels, R. M. Vaccination against helminth parasite infections. Expert Rev. Vaccines 13, 473–487 (2014).
Ntalli, N. G. & Caboni, P. Botanical nematicides: a review. J. Agric. Food. Chem. 60, 9929–9940 (2012).
Young, N. D. et al. Whole-genome sequence of Schistosoma haematobium. Nat. Genet. 44, 221–225 (2012).
The Schistosoma japonicum Genome Sequencing and Functional Analysis Consortium. The Schistosoma japonicum genome reveals features of host–parasite interplay. Nature 460, 345–351 (2009).
Protasio, A. V. et al. A systematically improved high quality genome and transcriptome of the human blood fluke Schistosoma mansoni. PLoS Negl. Trop. Dis. 6, e1455 (2012).
Wang, X. et al. The draft genome of the carcinogenic human liver fluke Clonorchis sinensis. Genome. Biol. 12, R107 (2011).
McNulty, S. N. et al. Genomes of Fasciola hepatica from the Americas reveal colonization with Neorickettsia endobacteria related to the agents of potomac horse and human sennetsu fevers. PLoS Genet. 13, e1006537 (2017).
Tsai, I. J. et al. The genomes of four tapeworm species reveal adaptations to parasitism. Nature 496, 57–63 (2013).
Bennett, H. M. et al. The genome of the sparganosis tapeworm Spirometra erinaceieuropaei isolated from the biopsy of a migrating brain lesion. Genome. Biol. 15, 510 (2014).
Schiffer, P. H. et al. The genome of Romanomermis culicivorax: revealing fundamental changes in the core developmental genetic toolkit in Nematoda. BMC Genomics 14, 923 (2013).
Mitreva, M. et al. The draft genome of the parasitic nematode Trichinella spiralis. Nat. Genet. 43, 228–235 (2011).
Foth, B. J. et al. Whipworm genome and dual-species transcriptome analyses provide molecular insights into an intimate host–parasite interaction. Nat. Genet. 46, 693–700 (2014).
Hunt, V. L. et al. The genomic basis of parasitism in the Strongyloides clade of nematodes. Nat. Genet. 48, 299–307 (2016).
Kikuchi, T. et al. Genomic insights into the origin of parasitism in the emerging plant pathogen Bursaphelenchus xylophilus. PLoS Pathog. 7, e1002219 (2011).
Cotton, J. A. et al. The genome and life-stage specific transcriptomes of Globodera pallida elucidate key aspects of plant parasitism by a cyst nematode. Genome Biol. 15, R43 (2014).
Opperman, C. H. et al. Sequence and genetic map of Meloidogyne hapla: a compact nematode genome for plant parasitism. Proc. Natl Acad. Sci. USA 105, 14802–14807 (2008).
Ghedin, E. et al. Draft genome of the filarial nematode parasite Brugia malayi. Science 317, 1756–1760 (2007).
Godel, C. et al. The genome of the heartworm, Dirofilaria immitis, reveals drug and vaccine targets. FASEB J. 26, 4650–4661 (2012).
Desjardins, C. A. et al. Genomics of Loa loa, a Wolbachia-free filarial parasite of humans. Nat. Genet. 45, 495–500 (2013).
Cotton, J. A. et al. The genome of Onchocerca volvulus, agent of river blindness. Nat. Microbiol. 2, 16216 (2016).
Wang, J. et al. Silencing of germline-expressed genes by DNA elimination in somatic cells. Dev. Cell 23, 1072–1080 (2012).
Tang, Y. T. et al. Genome of the human hookworm Necator americanus. Nat. Genet. 46, 261–269 (2014).
Tyagi, R. et al. Cracking the nodule worm code advances knowledge of parasite biology and biotechnology to tackle major diseases of livestock. Biotechnol. Adv. 33, 980–991 (2015).
McNulty, S. N. et al. Dictyocaulus viviparus genome, variome and transcriptome elucidate lungworm biology and support future intervention. Sci. Rep. 6, 20316 (2016).
Laing, R. et al. The genome and transcriptome of Haemonchus contortus, a key model parasite for drug and vaccine discovery. Genome. Biol. 14, R88 (2013).
C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Dieterich, C. et al. The Pristionchus pacificus genome provides a unique perspective on nematode lifestyle and parasitism. Nat. Genet. 40, 1193–1198 (2008).
Robb, S. M., Ross, E. & Sanchez Alvarado, A. SmedGD: the Schmidtea mediterranea genome database. Nucleic Acids Res. 36, D599–D606 (2008).
Srinivasan, J. et al. The draft genome and transcriptome of Panagrellus redivivus are shaped by the harsh demands of a free-living lifestyle. Genetics 193, 1279–1295 (2013).
Srivastava, M. et al. The Amphimedon queenslandica genome and the evolution of animal complexity. Nature 466, 720–726 (2010).
Simakov, O. et al. Insights into bilaterian evolution from three spiralian genomes. Nature 493, 526–531 (2013).
Satou, Y. et al. Improved genome assembly and evidence-based global gene model set for the chordate Ciona intestinalis: new insight into intron and operon populations. Genome Biol. 9, R152 (2008).
Zhang, G. et al. The oyster genome reveals stress adaptation and complexity of shell formation. Nature 490, 49–54 (2012).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Adams, M. D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Pagel Van Zee, J. et al. Tick genomics: the Ixodes genome project and beyond. Int. J. Parasitol. 37, 1297–1305 (2007).
Putnam, N. H. et al. Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317, 86–94 (2007).
Srivastava, M. et al. The Trichoplax genome and the nature of placozoans. Nature 454, 955–960 (2008).
Lynch, M., Bobay, L. M., Catania, F., Gout, J. F. & Rho, M. The repatterning of eukaryotic genomes by random genetic drift. Annu. Rev. Genomics Hum. Genet. 12, 347–366 (2011).
Vilella, A. J. et al. EnsemblCompara GeneTrees: complete, duplication-aware phylogenetic trees in vertebrates. Genome Res. 19, 327–335 (2009).
Rao, A. U., Carta, L. K., Lesuisse, E. & Hamza, I. Lack of heme synthesis in a free-living eukaryote. Proc. Natl Acad. Sci. USA 102, 4270–4275 (2005).
Wu, B. et al. Interdomain lateral gene transfer of an essential ferrochelatase gene in human parasitic nematodes. Proc. Natl Acad. Sci. USA 110, 7748–7753 (2013).
Nagayasu, E. et al. Identification of a bacteria-like ferrochelatase in Strongyloides venezuelensis, an animal parasitic nematode. PLoS ONE 8, e58458 (2013).
Casaravilla, C. et al. Characterization of myo-inositol hexakisphosphate deposits from larval Echinococcus granulosus. FEBS J. 273, 3192–3203 (2006).
Diaz, A., Casaravilla, C., Barrios, A. A. & Ferreira, A. M. Parasite molecules and host responses in cystic echinococcosis. Parasite Immunol. 38, 193–205 (2016).
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Valentim, C. L. et al. Genetic and molecular basis of drug resistance and species-specific drug action in schistosome parasites. Science 342, 1385–1389 (2013).
Parsons, L. M. et al. Caenorhabditis elegans bacterial pathogen resistant bus-4 mutants produce altered mucins. PLoS ONE 9, e107250 (2014).
Shapira, M. et al. A conserved role for a GATA transcription factor in regulating epithelial innate immune responses. Proc. Natl Acad. Sci. USA 103, 14086–14091 (2006).
Hewitson, J. P. et al. Proteomic analysis of secretory products from the model gastrointestinal nematode Heligmosomoides polygyrus reveals dominance of venom allergen-like (VAL) proteins. J. Proteomics 74, 1573–1594 (2011).
van der Hoeven, R., Cruz, M. R., Chavez, V. & Garsin, D. A. Localization of the dual oxidase BLI-3 and characterization of its NADPH oxidase domain during infection of Caenorhabditis elegans. PLoS ONE 10, e0124091 (2015).
Esteban, M. R., Giovinazzo, G., de la Hera, A. & Goday, C. PUMA1: a novel protein that associates with the centrosomes, spindle and centromeres in the nematode Parascaris. J. Cell Sci. 111, 723–735 (1998).
Tobler, H., Etter, A. & Muller, F. Chromatin diminution in nematode development. Trends Genet. 8, 427–432 (1992).
Albarqi, M. M. et al. Regulation of life cycle checkpoints and developmental activation of infective larvae in Strongyloides stercoralis by dafachronic acid. PLoS Pathog. 12, e1005358 (2016).
Zarlenga, D. S., Nisbet, A. J., Gasbarre, L. C. & Garrett, W. M. A calcium-activated nucleotidase secreted from Ostertagia ostertagi 4th-stage larvae is a member of the novel salivary apyrases present in blood-feeding arthropods. Parasitology 138, 333–343 (2011).
Cathcart, M. K. & Bhattacharjee, A. Monoamine oxidase A (MAO-A): a signature marker of alternatively activated monocytes/macrophages. Inflamm. Cell Signal. 1, e161 (2014).
Coakley, G., Maizels, R. M. & Buck, A. H. Exosomes and other extracellular vesicles: the new communicators in parasite infections. Trends Parasitol. 31, 477–489 (2015).
Wu, C. et al. Mapping the binding between the tetraspanin molecule (Sjc23) of Schistosoma japonicum and human non-immune IgG. PLoS ONE 6, e19112 (2011).
Krautz-Peterson, G. et al. Schistosoma mansoni infection of mice, rats and humans elicits a strong antibody response to a limited number of reduction-sensitive epitopes on five major tegumental membrane proteins. PLoS Negl. Trop. Dis. 11, e0005306 (2017).
Prior, A. et al. A surface-associated retinol- and fatty acid-binding protein (Gp-FAR-1) from the potato cyst nematode Globodera pallida: lipid binding activities, structural analysis and expression pattern. Biochem. J. 356, 387–394 (2001).
Rey-Burusco, M. F. et al. Diversity in the structures and ligand-binding sites of nematode fatty acid and retinol-binding proteins revealed by Na-FAR-1 from Necator americanus. Biochem. J. 471, 403–414 (2015).
Dell, A., Haslam, S. M. & Morris, H. R. in Parasitic Nematodes: Molecular Biology, Biochemistry and Immunology (eds. Kennedy, M. W. & Harnett, W.) 285–307 (Cabi Publishing, Oxfordshire, UK, 2013).
Rodrigues, J. A. et al. Parasite glycobiology: a bittersweet symphony. PLoS Pathog. 11, e1005169 (2015).
Anderson, L. et al. Schistosoma mansoni egg, adult male and female comparative gene expression analysis and identification of novel genes by RNA-Seq. PLoS Negl. Trop. Dis. 9, e0004334 (2015).
Gong, H. et al. A novel PAN/apple domain-containing protein from Toxoplasma gondii: characterization and receptor identification. PLoS ONE 7, e30169 (2012).
Cantacessi, C. et al. A portrait of the “SCP/TAPS” proteins of eukaryotes—developing a framework for fundamental research and biotechnological outcomes. Biotechnol. Adv. 27, 376–388 (2009).
McKerrow, J. H., Caffrey, C., Kelly, B., Loke, P. & Sajid, M. Proteases in parasitic diseases. Annu. Rev. Pathol. 1, 497–536 (2006).
Williamson, A. L. et al. Ancylostoma caninum MTP-1, an astacin-like metalloprotease secreted by infective hookworm larvae, is involved in tissue migration. Infect. Immun. 74, 961–967 (2006).
Williamson, A. L. et al. A multi-enzyme cascade of hemoglobin proteolysis in the intestine of blood-feeding hookworms. J. Biol. Chem. 279, 35950–35957 (2004).
Delcroix, M. et al. A multienzyme network functions in intestinal protein digestion by a platyhelminth parasite. J. Biol. Chem. 281, 39316–39329 (2006).
Duffy, M. S., Cevasco, D. K., Zarlenga, D. S., Sukhumavasi, W. & Appleton, J. A. Cathepsin B homologue at the interface between a parasitic nematode and its intermediate host. Infect. Immun. 74, 1297–1304 (2006).
Cancela, M. et al. A distinctive repertoire of cathepsins is expressed by juvenile invasive Fasciola hepatica. Biochimie 90, 1461–1475 (2008).
Knox, D. P. Proteinase inhibitors and helminth parasite infection. Parasite Immunol. 29, 57–71 (2007).
Rehman, A. A., Ahsan, H. & Khan, F. H. Alpha-2-macroglobulin: a physiological guardian. J. Cell. Physiol. 228, 1665–1675 (2013).
Martzen, M. R., Geise, G. L., Hogan, B. J. & Peanasky, R. J. Ascaris suum: localization by immunochemical and fluorescent probes of host proteases and parasite proteinase inhibitors in cross-sections. Exp. Parasitol. 60, 139–149 (1985).
Nei, M., Niimura, Y. & Nozawa, M. The evolution of animal chemosensory receptor gene repertoires: roles of chance and necessity. Nat. Rev. Genet. 9, 951–963 (2008).
Lynagh, T. et al. Molecular basis for convergent evolution of glutamate recognition by pentameric ligand-gated ion channels. Sci. Rep. 5, 8558 (2015).
MacDonald, K. et al. Functional characterization of a novel family of acetylcholine-gated chloride channels in Schistosoma mansoni. PLoS Pathog. 10, e1004181 (2014).
Courtot, E. et al. Functional characterization of a novel class of morantel-sensitive acetylcholine receptors in nematodes. PLoS Pathog. 11, e1005267 (2015).
Vasiliou, V., Vasiliou, K. & Nebert, D. W. Human ATP-binding cassette (ABC) transporter family. Hum. Genomics 3, 281–290 (2009).
Martin-Duran, J. M., Ryan, J. F., Vellutini, B. C., Pang, K. & Hejnol, A. Increased taxon sampling reveals thousands of hidden orthologs in flatworms. Genome Res. 27, 1263–1272 (2017).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Kondrashov, F. A., Koonin, E. V., Morgunov, I. G., Finogenova, T. V. & Kondrashova, M. N. Evolution of glyoxylate cycle enzymes in Metazoa: evidence of multiple horizontal transfer events and pseudogene formation. Biol. Direct. 1, 31 (2006).
Harder, A. The biochemistry of Haemonchus contortus and other parasitic nematodes. Adv. Parasitol. 93, 69–94 (2016).
Marr, J. J. & Müller, M. Biochemistry and Molecular Biology of Parasites (Academic Press, San Diego, CA, USA, 1995).
Pearce, E. J. & Huang, S. C. The metabolic control of schistosome egg production. Cell Microbiol. 17, 796–801 (2015).
Mehlhorn, H. (ed.) Encyclopedia of Parasitology (Springer, New York, NY, USA, 2008).
Watson, E. et al. Metabolic network rewiring of propionate flux compensates vitamin B12 deficiency in C. elegans. eLife 5, e17670 (2016).
Van Soest, P. J. Nutritional Ecology of the Ruminant (Cornell Univ. Press, Ithaca, NY, USA, 1994).
Taylor, C. M. et al. Using existing drugs as leads for broad spectrum anthelmintics targeting protein kinases. PLoS Pathog. 9, e1003149 (2013).
Vermeire, J. J., Suzuki, B. M. & Caffrey, C. R. Odanacatib, a cathepsin K cysteine protease inhibitor, kills hookworm in vivo. Pharmaceuticals 9, 39 (2016).
Simpson, J. T. & Durbin, R. Efficient de novo assembly of large genomes using compressed data structures. Genome Res. 22, 549–556 (2012).
Gremme, G., Steinbiss, S. & Kurtz, S. GenomeTools: a comprehensive software library for efficient processing of structured genome annotations. IEEE/ACM Trans. Comput. Biol. Bioinform. 10, 645–656 (2013).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Boetzer, M., Henkel, C. V., Jansen, H. J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011).
Boetzer, M. & Pirovano, W. Toward almost closed genomes with GapFiller. Genome Biol. 13, R56 (2012).
Tsai, I. J., Otto, T. D. & Berriman, M. Improving draft assemblies by iterative mapping and assembly of short reads to eliminate gaps. Genome Biol. 11, R41 (2010).
Otto, T. D., Sanders, M., Berriman, M. & Newbold, C. Iterative correction of reference nucleotides (iCORN) using second generation sequencing technology. Bioinformatics 26, 1704–1707 (2010).
Hunt, M. et al. REAPR: a universal tool for genome assembly evaluation. Genome Biol. 14, R47 (2013).
Bonfield, J. K. & Whitwham, A. Gap5—editing the billion fragment sequence assembly. Bioinformatics 26, 1699–1703 (2010).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Coghlan, A. L., Gordon, D. & Berriman, M. Contamination screening of parasitic worm genome assemblies. Protoc. Exch. https://doi.org/10.1038/protex.2018.038 (2018).
Parra, G., Bradnam, K., Ning, Z., Keane, T. & Korf, I. Assessing the gene space in draft genomes. Nucleic Acids Res. 37, 289–297 (2009).
Quinlan, A. R. BEDTools: the Swiss-army tool for genome feature analysis. Curr. Protoc. Bioinformatics 47, 11.12.1–11.12.34 (2014).
Stanley, E., Coghlan, A. L. & Berriman, M. A MAKER pipeline for prediction of protein-coding genes in parasitic worm genomes. Protoc. Exch. https://doi.org/10.1038/protex.2018.056 (2018).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491 (2011).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Ter-Hovhannisyan, V., Lomsadze, A., Chernoff, Y. O. & Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res. 18, 1979–1990 (2008).
She, R. et al. genBlastG: using BLAST searches to build homologous gene models. Bioinformatics 27, 2141–2143 (2011).
Otto, T. D., Dillon, G. P., Degrave, W. S. & Berriman, M. RATT: rapid annotation transfer tool. Nucleic Acids Res. 39, e57 (2011).
Cochrane, G., Karsch-Mizrachi, I. & Takagi, T., International Nucleotide Sequence Database Collbaoration. The International Nucleotide Sequence Database Collaboration. Nucleic Acids Res. 44, D48–D50 (2016).
Martin, J. & Mitreva, M. Genomic and transcriptomic data production for helminths. Protoc. Exch. https://doi.org/10.1038/protex.2018.044 (2018).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Butler, J. et al. ALLPATHS: de novo assembly of whole-genome shotgun microreads. Genome Res. 18, 810–820 (2008).
Xue, W. et al. L_RNA_scaffolder: scaffolding genomes with transcripts. BMC Genomics 14, 604 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Morgulis, A. et al. Database indexing for production MegaBLAST searches. Bioinformatics 24, 1757–1764 (2008).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Kumar, S. & Blaxter, M. L. Simultaneous genome sequencing of symbionts and their hosts. Symbiosis 55, 119–126 (2011).
Crusoe, M. R. et al. The khmer software package: enabling efficient nucleotide sequence analysis. F1000Res. 4, 900 (2015).
Simpson, J. T. et al. ABySS: a parallel assembler for short read sequence data. Genome Res. 19, 1117–1123 (2009).
Chevreux, B. et al. Using the miraEST assembler for reliable and automated mRNA transcript assembly and SNP detection in sequenced ESTs. Genome Res. 14, 1147–1159 (2004).
Darby, A. C. et al. Analysis of gene expression from the Wolbachia genome of a filarial nematode supports both metabolic and defensive roles within the symbiosis. Genome Res. 22, 2467–2477 (2012).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinformatics 9, 18 (2008).
Coghlan, A. L., Tsai, I. J. & Berriman, M. Creation of a comprehensive repeat library for a newly sequenced parasitic worm genome. Protoc. Exch. https://doi.org/10.1038/protex.2018.054 (2018).
Hadfield, J. D. MCMC methods for multi-response generalized linear mixed models: the MCMCglmm R package. J. Stat. Softw. 33, 1–22 (2010).
Britton, T., Anderson, C. L., Jacquet, D., Lundqvist, S. & Bremer, K. Estimating divergence times in large phylogenetic trees. Syst. Biol. 56, 741–752 (2007).
Tatusova, T. Update on genomic databases and resources at the National Center for Biotechnology Information. Methods Mol. Biol. 1415, 3–30 (2016).
Katoh, K. & Standley, D. M. MAFFT: iterative refinement and additional methods. Methods Mol. Biol. 1079, 131–146 (2014).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Dunn, C. W. et al. Broad phylogenomic sampling improves resolution of the animal tree of life. Nature 452, 745–749 (2008).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Bairoch, A. & Apweiler, R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 28, 4–48 (2000).
Hunter, S. et al. InterPro in 2011: new developments in the family and domain prediction database. Nucleic Acids Res. 40, D306–D312 (2012).
Coghlan, A. L. & Berriman, M. Functional annotation of parasitic worm genomes, by assigning protein names and GO terms. Protoc. Exch. https://doi.org/10.1038/protex.2018.055 (2018).
Kall, L., Krogh, A. & Sonnhammer, E. L. Advantages of combined transmembrane topology and signal peptide prediction—the Phobius web server. Nucleic Acids Res. 35, W429–W432 (2007).
Bendtsen, J. D., Jensen, L. J., Blom, N., Von Heijne, G. & Brunak, S. Feature-based prediction of non-classical and leaderless protein secretion. Protein Eng. Des. Sel. 17, 349–356 (2004).
Laetsch, D. R. & Blaxter, M. L. KinFin: software for taxon-aware analysis of clustered protein sequences. G3 7, 3349–3357 (2017).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Ribeiro, D., Coghlan, A. L., Harsha, B. & Berriman, M. Identification of lineage-specific gene family expansions in a database of gene families. Protoc. Exch. https://doi.org/10.1038/protex.2018.057 (2018).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Rawlings, N. D. & Morton, F. R. The MEROPS batch BLAST: a tool to detect peptidases and their non-peptidase homologues in a genome. Biochimie 90, 243–259 (2008).
Finn, R. D. et al. The Pfam protein families database. Nucleic Acids Res. 38, D211–D222 (2010).
Howe, K. L. et al. WormBase 2016: expanding to enable helminth genomic research. Nucleic Acids Res. 44, D774–D780 (2016).
Soding, J. Protein homology detection by HMM-HMM comparison. Bioinformatics 21, 951–960 (2005).
Wheeler, N., Day, T., Zamanian, M. & Kimber, M. GPCR identification in parasitic worm genome assemblies. Protoc. Exch. https://doi.org/10.1038/protex.2018.061 (2018).
Jones, A. K., Davis, P., Hodgkin, J. & Sattelle, D. B. The nicotinic acetylcholine receptor gene family of the nematode Caenorhabditis elegans: an update on nomenclature. Invert. Neurosci. 7, 129–131 (2007).
Li, B. W., Rush, A. C. & Weil, G. J. Expression of five acetylcholine receptor subunit genes in Brugia malayi adult worms. Int. J. Parasitol. Drugs Drug Resist. 5, 100–109 (2015).
Buxton, S. K. et al. Investigation of acetylcholine receptor diversity in a nematode parasite leads to characterization of tribendimidine- and derquantel-sensitive nAChRs. PLoS Pathog. 10, e1003870 (2014).
Howe, K. L., Bolt, B. J., Shafie, M., Kersey, P. & Berriman, M. WormBase ParaSite—a comprehensive resource for helminth genomics. Mol. Biochem. Parasitol. 215, 2–10 (2017).
Tusnady, G. E. & Simon, I. The HMMTOP transmembrane topology prediction server. Bioinformatics 17, 849–850 (2001).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Miranda-Saavedra, D. & Barton, G. J. Classification and functional annotation of eukaryotic protein kinases. Proteins 68, 893–914 (2007).
Martin, J. & Mitreva, M. Kinase annotation for helminths. Protoc. Exch. https://doi.org/10.1038/protex.2018.042 (2018).
Hung, S. S., Wasmuth, J., Sanford, C. & Parkinson, J. DETECT—a density estimation tool for enzyme classification and its application to Plasmodium falciparum. Bioinformatics 26, 1690–1698 (2010).
Claudel-Renard, C., Chevalet, C., Faraut, T. & Kahn, D. Enzyme-specific profiles for genome annotation: PRIAM. Nucleic Acids Res. 31, 6633–6639 (2003).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, W182–W185 (2007).
Chang, A. et al. BRENDA in 2015: exciting developments in its 25th year of existence. Nucleic Acids Res. 43, D439–D446 (2015).
Swapna, S., Tyagi, R., Mitreva, M. & Parkinson, J. Annotating metabolic enzymes in parasitic worm proteomes. Protoc. Exch. https://doi.org/10.1038/protex.2018.047 (2018).
Karp, P. D. et al. Pathway Tools version 19.0 update: software for pathway/genome informatics and systems biology. Brief. Bioinformatics 17, 877–890 (2016).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 40, D742–D753 (2012).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Tyagi, R., Swapna, S., Parkinson, J. & Mitreva, M. Comparative analysis of metabolism in parasitic worms. Protoc. Exch. https://doi.org/10.1038/protex.2018.048 (2018).
Tyagi, R., Rosa, B. A., Lewis, W. G. & Mitreva, M. Pan-phylum comparison of nematode metabolic potential. PLoS Negl. Trop. Dis. 9, e0003788 (2015).
Real, R. & Vargas, J. M. The probabilistic basis of Jaccard’s index of similarity. Syst. Biol. 45, 380–385 (1996).
Taylor, C. M. et al. Discovery of anthelmintic drug targets and drugs using chokepoints in nematode metabolic pathways. PLoS Pathog. 9, e1003505 (2013).
Gaulton, A. et al. The ChEMBL database in 2017. Nucleic Acids Res. 45, D945–D954 (2017).
Coghlan, A. L. et al. Creating a screening set of potential anthelmintic compounds using ChEMBL. Protoc. Exch. https://doi.org/10.1038/protex.2018.053 (2018).
Velankar, S. et al. PDBe: improved accessibility of macromolecular structure data from PDB and EMDB. Nucleic Acids Res. 44, D385–D395 (2016).
Sterling, T. & Irwin, J. J. ZINC 15—ligand discovery for everyone. J. Chem. Inf. Model. 55, 2324–2337 (2015).
Wehrens, R. & Buydens, L. M. C. Self- and super-organizing maps in R: the kohonen package. J. Stat. Softw. 21, 1–19 (2007).
Acknowledgements
We thank: the WSI DNA Pipeline teams, particularly C. Griffiths, N. Park, L. Shirley, M. Quail, D. Willey and M. Jones; WSI Pathogen Informatics, especially J. Keane; T.D. Otto for bioinformatics advice; MGI faculty and staff, especially M. Schmidt, C. Fronick, M. Cordes, T. Miner, R. Fulton and other members of the Project Management, Resource Bank, Library Construction and Data Production teams; D. Hughes, M. Muffato at the European Bioinformatics Institute, for support running Maker and Ensembl Compara; and K. Gharbi and his staff at Edinburgh Genomics for support; V. Gelmedin, R. Fujiwara, F. Brazil, the late Purnomo (University of Indonesia, Jakarta), J. Ahringer, E.S. Hernández Redondo, F. Jackson, E. Redman, A. Ito, J. Saldaña, M. Fernanda Dominguez, W. Gause, M. Badets, I.E. Samonte, A. Koehler, M. Nielsen, L.S. Mansfield, T. Sonstegard for sample preparation. The work was supported by funding from Wellcome (206194), Medical Research Council (MR/L001020/1), and Biotechnology and Biological Sciences Research Council (BB/K020048/1) to M.B., and US National Institutes of Health (NIH)–National Human Genome Research Institute grant number U54HG003079, National Institute of Allergy and Infectious Diseases grant number AI081803, and National Institute of General Medical Sciences grant number GM097435 to M.M. Genome sequencing and analysis in Edinburgh was supported by EU SICA award 242131 ‘Enhanced Protective Immunity Against Filariasis’ (EPIAF) (to D.W.T.). S.A. Babayan. was also supported by the EU project EPIAF. G. Koutsovoulos was supported by a BBSRC/Edinburgh University PhD scholarship and D.R.L. by a joint Edinburgh University/James Hutton Institute PhD scholarship. J.E.A. was supported by MRC grant MR/K01207X/1. J.P. and S.S. were supported by the National Institutes of Health/NIAID (R21 AI126466) and the Natural Sciences and Engineering Research Council of Canada (RGPIN-2014-06664). Additional computing resources were provided through Compute Canada by the University of Toronto SciNet HPC Consortium. R.M.M. was supported by a Wellcome Investigator Award (ref. 106122) and Wellcome core funding to the Wellcome Centre for Molecular Parasitology (Ref 104111). J.B.M. was supported by the Scottish Government RESAS. T.S. was supported by the Institute of Parasitology, BC CAS (RVO: 60077344). A.R.L. and P.M. were supported by a Strategic Award from Wellcome (WT104104/Z/14/Z) and the Member States of the European Molecular Biology Laboratory (EMBL). Schistosome samples were obtained from the SCAN repository (Wellcome grant 104958).
Author information
Authors and Affiliations
Consortia
Contributions
Project leadership and conception: M.B. and M.M. Writing the manuscript: M.B., M.L.B., A.C., J.A.C., N.H., D.R.L., R.M.M., M.M. and R.Tyagi. Project planning or management: M.B., M.L.B., J.A.C., N.H., M.M. and A.M. Genome sequencing: H.B., K.H.P., N.H., J.M., P.O., D.W.T., A.T. and Z.X. Preparation or provision of parasite material or nucleic acid: F.A., J.E.A., K.A., S.A.Bisset, G.B., H.M.B., S.A.Babayan, T.C.B., E.C., J.C., P.J.C., C.C., E.D., M.L.E., A.E., K.S.E., P.F., J.S.G., Y.H., J.M.H., D.E.H., J.H., P.H., T.H., M.K., T.K., R.M.M., B.M., J.B.M., P.D.O., A.O., F.P., K.P., D.R., M.G.S., H.S., M.Schnyder, T.S., V.N.T., D.W.T., R.Toledo, J.F.U., L.C.W. and D.Z. Production bioinformatics: A.C., J.A.C., D.G., B.H., J.L., P.O., D.M.R., B.A.R., E.S. and A.T. Annotation of genomes: A.C., B.H., K.L.H., G.Kaur, G.Koutsovoulos, S.K., D.R.L., J.M., P.O., K.H.P., B.A.R., E.S. and Z.X. Assembly of genomes at WSI: I.J.T. Assembly of genomes at MGI: J.M., P.O., K.H.P. and Z.X. Assembly of genomes at BaNG: G.Kaur, G.Koutsovoulos, S.K. and D.R.L. Assembly and annotation of mitochondrial genomes: T.K. Analysis of variation in genome size: A.C. and J.A.C. Analysis of GO terms and domains: A.C., D.R.L., J.L., A.J.R., B.A.R. and M.Shafie. Development of visualisation software and production of the species tree: J.A.C. Analysis of synapomorphies: M.L.B. and D.R.L. Analysis of expanded gene families: M.B., A.C., J.A.C., S.R.D., N.H., A.J.R., D.M.R., G.R., B.A.R. and R.Tyagi. Analysis of hypothetical genes: A.C. and A.J.R. Analysis of SCP/TAPS: H.M.K., T.H.K., T.J.L. and I.J.T. Analysis of proteases: N.D.R. Analysis of GPCRs: T.A.D., N.J.W. and M.Z. Analysis of ion channels: R.N.B. and M.Z. Analysis of kinases: J.M. and B.A.R. Analysis of metabolic pathways: A.C., J.A.C., T.K., J.P., S.S. and R.Tyagi. Chemogenomics analyses: A.C., J.A.C., A.R.L., J.L., P.M. and N.M.O’B.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–32 and Supplementary Note
Supplementary Table 1
Sample and sequencing information
Supplementary Table 2
Assembly statistics
Supplementary Table 3
Gene set properties
Supplementary Table 4
Rationale, classification and taxonomy of species analyzed
Supplementary Table 5
Repeat content
Supplementary Table 6
Results of Bayesian mixed effect model for genome size
Supplementary Table 7
GO annotation, predicted secretion data and transmembrane domains, and per-species gene counts for Compara gene families
Supplementary Table 8
Synapomorphic gene families
Supplementary Table 9
Gene families in expanded family analysis
Supplementary Table 10
SCP/TAPS genes
Supplementary Table 11
Protease and protease inhibitor genes
Supplementary Table 12
Species life history traits
Supplementary Table 13
Parasite tissue tropism in definitive host
Supplementary Table 14
Nematode-specific and platyhelminth-specific domain combinations
Supplementary Table 15
GPCR annotation
Supplementary Table 16
Ligand gated ion channel genes
Supplementary Table 17
ABC transporter predictions
Supplementary Table 18
EC annotations, KEGG metabolic module completion, KEGG pathway conservation, auxotrophy prediction, and metabolic chokepoints
Supplementary Table 19
Annotation for metabolic pathways of particular interest
Supplementary Table 20
KEGG pathways with the most unique ECs (present in flatworms but absent in nematodes, or vice versa)
Supplementary Table 21
Known anthelmintic compounds, known or likely anthelmintic drug targets, and potential new anthelmintic drug targets and compounds
Supplementary Table 22
RNA-seq used for gene-finding quality control
Supplementary Table 23
Kinase genes
Supplementary Table 24
InterPro, Pfam and GO enrichment in different species groups
Supplementary Table 25
CAZYME predictions for the 81 nematodes and platyhelminths
Supplementary Data
Compara gene families and the genes that they contain.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
International Helminth Genomes Consortium. Comparative genomics of the major parasitic worms. Nat Genet 51, 163–174 (2019). https://doi.org/10.1038/s41588-018-0262-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0262-1
This article is cited by
-
The mrp-3 gene is involved in haem efflux and detoxification in a blood-feeding nematode
BMC Biology (2024)
-
Diversity and evolution of transposable elements in the plant-parasitic nematodes
BMC Genomics (2024)
-
A comparative analysis of planarian genomes reveals regulatory conservation in the face of rapid structural divergence
Nature Communications (2024)
-
Repurposing mebendazole against triple-negative breast cancer CNS metastasis
Journal of Neuro-Oncology (2024)
-
Nuclear genome annotation of wheel animals and thorny-headed worms: inferences about the last common ancestor of Syndermata (Rotifera s.l.)
Hydrobiologia (2024)