Abstract
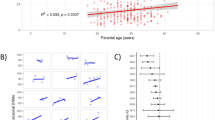
De novo mutations (DNMs) cause a large proportion of severe rare diseases of childhood. DNMs that occur early may result in mosaicism of both somatic and germ cells. Such early mutations can cause recurrence of disease. We scanned 1,007 sibling pairs from 251 families and identified 878 DNMs shared by siblings (ssDNMs) at 448 genomic sites. We estimated DNM recurrence probability based on parental mosaicism, sharing of DNMs among siblings, parent-of-origin, mutation type and genomic position. We detected 57.2% of ssDNMs in the parental blood. The recurrence probability of a DNM decreases by 2.27% per year for paternal DNMs and 1.78% per year for maternal DNMs. Maternal ssDNMs are more likely to be T>C mutations than paternal ssDNMs, and less likely to be C>T mutations. Depending on the properties of the DNM, the recurrence probability ranges from 0.011% to 28.5%. We have launched an online calculator to allow estimation of DNM recurrence probability for research purposes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Access to these data is controlled, the DNMs are a subset of the sequence variants deposited to the European Nucleotide Archive (PRJEB15197).
References
Tang, W. W. C., Kobayashi, T., Irie, N., Dietmann, S. & Surani, M. A. Specification and epigenetic programming of the human germ line. Nat. Rev. Genet. 17, 585–600 (2016).
De Felici M. in Oogenesis, 19–38 (Coticchio, G., Albertini, D. F. & de Santis, L. (Eds.); Springer Verlag, London; 2013).
Biesecker, L. G. & Spinner, N. B. A genomic view of mosaicism and human disease. Nat. Rev. Genet. 14, 307–320 (2013).
Samuels, M. E. & Friedman, J. M. Genetic mosaics and the germ line lineage. Genes (Basel) 6, 216–237 (2015).
Ju, Y. S. et al. Somatic mutations reveal asymmetric cellular dynamics in the early human embryo. Nature 543, 714–718 (2017).
Woodworth, M. B., Cirskis, K. M. & Walsh, C. A. Building a lineage from single cells: genetic techniques for cell lineage tracking. Nat. Rev. Genet. 18, 230–244 (2017).
Lodato, M. A. et al. Somatic mutation in single human neurons tracks developmental and transcriptional history. Science 350, 94–98 (2015).
Rahbari, R. et al. Timing, rates and spectra of human germline mutation. Nat. Genet. 48, 126–133 (2015).
Acuna-Hidalgo, R. et al. Post-zygotic point mutations are an underrecognized source of de novo genomic variation. Am. J. Hum. Genet. 97, 67–74 (2015).
Campbell, I. M. et al. Parent of origin, mosaicism, and recurrence risk: probabilistic modeling explains the broken symmetry of transmission genetics. Am. J. Hum. Genet. 95, 345–359 (2014).
Campbell, I. M. et al. Parental somatic mosaicism is underrecognized and influences recurrence risk of genomic disorders. Am. J. Hum. Genet. 95, 173–182 (2014).
Helderman-van Den Enden, A. T. J. M. et al. Recurrence risk due to germ line mosaicism: Duchenne and Becker muscular dystrophy. Clin. Genet. 75, 465–472 (2009).
Bakker, E. et al. Germinal mosaicism increases the recurrence risk for new Duchenne muscular dystrophy mutations. J. Med. Genet. 26, 553–559 (1989).
Mari, F. et al. Germline mosaicism in Rett syndrome identified by prenatal diagnosis. Clin. Genet. 67, 258–260 (2005).
Rose, V. M. et al. Germ-line mosaicism in tuberous sclerosis: how common? Am. J. Hum. Genet. 64, 986–992 (1999).
Verhoef, S. et al. High rate of mosaicism in tuberous sclerosis complex. Am. J. Hum. Genet. 64, 1632–1637 (1999).
Pyott, S. M. et al. Recurrence of perinatal lethal osteogenesis imperfecta in sibships: parsing the risk between parental mosaicism for dominant mutations and autosomal recessive inheritance. Genet. Med. 13, 125–130 (2011).
Jónsson, H. et al. Parental influence on human germline de novo mutations in 1,548 trios from Iceland. Nature 549, 519–522 (2017).
Kong, A. et al. Rate of de novo mutations and the importance of father’s age to disease risk. Nature 488, 471–475 (2012).
Scally, A. Mutation rates and the evolution of germline structure. Philos. Trans. R. Soc. Lond. B 371, 20150137 (2016).
Wong, W. S. W. et al. New observations on maternal age effect on germline de novo mutations. Nat. Commun. 7, 10486 (2016).
Maher, G. J. et al. Visualizing the origins of selfish de novo mutations in individual seminiferous tubules of human testes. Proc. Natl Acad. Sci. USA 113, 2454–2459 (2016).
Saitsu, H. et al. De novo mutations in the gene encoding STXBP1 (MUNC18-1) cause early infantile epileptic encephalopathy. Nat. Genet. 40, 782–788 (2008).
Allen, A. S. et al. De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013).
Scally, A. et al. Insights into hominid evolution from the gorilla genome sequence. Nature 483, 169–175 (2012).
Jónsson, H. et al. Whole genome characterization of sequence diversity of 15,220 Icelanders. Sci. Data 4, 170115 (2017).
Busing, F., Meijer, E. & Van Der Leeden, R. Delete-m jackknife for unequal m. Stat Comput 9, 2–7 (1999).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Eggertsson, H. P. et al. Graphtyper enables population-scale genotyping using pangenome graphs. Nat. Genet. 49, 1654–1660 (2017).
Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Acknowledgements
We thank all of the participants in this study. The study was performed in collaboration with Illumina.
Author information
Authors and Affiliations
Contributions
H.J., G.A.A., F.Z., B.K., K.E.H., B.Ö.J. and D.F.G. analyzed the data. H.J., G.P., H.P.E., S.K., F.Z., B.K., A.G. and D.F.G. conceived methods for analyzing data. Ad.J., As.J. and O.Th.M. performed experiments. P.S., G.A.A., B.Ö.J., I.J., S.E.M. and S.A.G. collected samples and information. H.J., P.S., O.Th.M., U.T., G.M., B.V.H., D.F.G. and K.S. designed the study. H.J., P.S., D.F.G. and K.S. wrote the manuscript with input from G.A.A., G.P., K.E.H., S.N.S., U.T., A.K., B.V.H. and A.H.
Corresponding authors
Ethics declarations
Competing interests
All of the authors except S.E.M. are employees of deCODE Genetics/Amgen, Inc.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
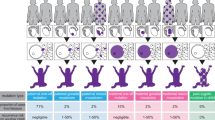
Supplementary Figure 1 Germ cell lineages of parents with nine offspring.
a,b, The male and female germ cell lineages of a nine-offspring family. c,d, The paternal and maternal germ cell lineages of a nine-offspring family. Purple and green text correspond to DNMs determined by the trio and sibling approach, respectively. Percentages are paternal and maternal imbalances, respectively. The paternal imbalance serves as a background rate for the maternal DNMs, and vice versa. In addition to ssDNMs, we incorporated DNMs that are somatic mosaic in the parent (≥4 reads supporting the alternative allele in the parent). DNM sites are ordered according to carrier frequency, from high (top) to low (bottom) frequency.
Supplementary Figure 2 Germ cell lineages of nine-offspring parents.
a,b, The male and female germ cell lineages of a nine-offspring family. c,d, The paternal and maternal germ cell lineages of a nine-offspring family. Purple and green text correspond to DNMs determined by the trio and sibling approach, respectively. Percentages are paternal and maternal allelic imbalances, respectively. The paternal allelic imbalance serves as a background rate for the maternal DNMs, and vice versa. In addition to ssDNMs, we incorporated DNMs that are somatic mosaic in the parent (≥4 reads supporting the alternative allele in the parent). DNM sites are ordered according to carrier frequency, from high (top) to low (bottom) frequency.
Supplementary Figure 3 ssDNM rate against allelic imbalance in parents for various subsets.
a, Restricting to DNMs determined by the trio approach and DNMs from the sibling approach in which the allelic balance difference between carriers and parents was rejected. A two-sided Fisher test was used for the allelic balance test difference. The P values were adjusted with Holm multiple-testing correction using the number of DNMs found solely by the sibling approach. b, In addition to rejecting the null hypothesis as in a, we required DNMs determined by the sibling approach to have at least two siblings who were non-carriers despite having the same haplotype background as the DNM carriers (absent). For a and b, the analysis was restricted to deep-sequenced parents (13 parent pairs and 518 sibling pairs). c, Same as in a except that all sibling pairs were used (238 parent pairs and 518 sibling pairs). d, Same as in b except that all sibling pairs were used (238 parent pairs and 518 sibling pairs). The center values are means and error bars are 95% CIs.
Supplementary Figure 4 The power of the sibling method.
The power of detecting a DNM segregating in the parental germline conditional on DNM transmission.
Supplementary Figure 5 Targeted resequencing of DNM sites from the 10- and 17-sibling families.
a, The allelic imbalance from targeted resequencing against carrier status determined from whole-genome sequencing. In a, sites with over 100× coverage were used, resulting in 657 genotypes from 27 offspring from two parent pairs. In a, the hinges are the first and third quantiles, and notches are ±1.58 × IQR/√n, where IQR is the interquartile range and n is the number in each category. b–e, Targeted resequenced imbalance of the parents against the imbalance from whole-genome sequencing.
Supplementary Figure 6 A schematic overview of the recurrence calculator.
The green and orange states represent the intermediate analysis steps and datasets, respectively.
Supplementary Figure 7 SMPP as a function of reads supporting the alternative allele.
The vertical error bars are the 95% CIs assessed by the jackknife procedure. The center values are the means. 13 parent pairs and 1,517 sibling trios were used in the 200× subset, and 54 parent pairs and 241 sibling trios were used in the 30× subset.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–5 and Supplementary Note
Rights and permissions
About this article
Cite this article
Jónsson, H., Sulem, P., Arnadottir, G.A. et al. Multiple transmissions of de novo mutations in families. Nat Genet 50, 1674–1680 (2018). https://doi.org/10.1038/s41588-018-0259-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0259-9
This article is cited by
-
Meta-analysis of 46,000 germline de novo mutations linked to human inherited disease
Human Genomics (2024)
-
Assessment of parental mosaicism rates in neurodevelopmental disorders caused by apparent de novo pathogenic variants using deep sequencing
Scientific Reports (2024)
-
Human embryonic genetic mosaicism and its effects on development and disease
Nature Reviews Genetics (2024)
-
Evolution of the germline mutation rate across vertebrates
Nature (2023)
-
Genomic Mosaicism of the Brain: Origin, Impact, and Utility
Neuroscience Bulletin (2023)