Abstract
Polycomb repressive complexes (PRCs) control organismic development in higher eukaryotes through epigenetic gene repression1,2,3,4. PRC proteins do not contain DNA-binding domains, thus prompting questions regarding how PRCs find their target loci5. Here we present genome-wide evidence of PRC2 recruitment by telomere-repeat-binding factors (TRBs) through telobox-related motifs in Arabidopsis. A triple trb1-2, trb2-1, and trb3-2 (trb1/2/3) mutant with a developmental phenotype and a transcriptome strikingly similar to those of strong PRC2 mutants showed redistribution of trimethyl histone H3 Lys27 (H3K27me3) marks and lower H3K27me3 levels, which were correlated with derepression of TRB1-target genes. TRB1–3 physically interacted with the PRC2 proteins CLF and SWN. A SEP3 reporter gene with a telobox mutation showed ectopic expression, which was correlated with H3K27me3 depletion, whereas tethering TRB1 to the mutated cis element partially restored repression. We propose that telobox-related motifs recruit PRC2 through the interaction between TRBs and CLF/SWN, a mechanism essential for H3K27me3 deposition at a subset of target genes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lewis, E. B. A gene complex controlling segmentation in Drosophila. Nature 276, 565–570 (1978).
Mozgova, I. & Hennig, L. The polycomb group protein regulatory network. Annu. Rev. Plant Biol. 66, 269–296 (2015).
Xiao, J. & Wagner, D. Polycomb repression in the regulation of growth and development in Arabidopsis. Curr. Opin. Plant Biol. 23, 15–24 (2015).
Simon, J. A. & Kingston, R. E. Occupying chromatin: Polycomb mechanisms for getting to genomic targets, stopping transcriptional traffic, and staying put. Mol. Cell 49, 808–824 (2013).
Kassis, J. A. & Brown, J. L. Polycomb group response elements in Drosophila and vertebrates. Adv. Genet. 81, 83–118 (2013).
Förderer, A., Zhou, Y. & Turck, F. The age of multiplexity: recruitment and interactions of Polycomb complexes in plants. Curr. Opin. Plant Biol. 29, 169–178 (2016).
Derkacheva, M. et al. Arabidopsis MSI1 connects LHP1 to PRC2 complexes. EMBO J. 32, 2073–2085 (2013).
Liang, S. C. et al. Kicking against the PRCs: a domesticated transposase antagonises silencing mediated by Polycomb group proteins and is an accessory component of Polycomb repressive complex 2. PLoS Genet. 11, e1005660 (2015).
Molitor, A. M., Bu, Z., Yu, Y. & Shen, W. H. Arabidopsis AL PHD-PRC1 complexes promote seed germination through H3K4me3-to-H3K27me3 chromatin state switch in repression of seed developmental genes. PLoS Genet. 10, e1004091 (2014).
Lanzuolo, C. & Orlando, V. Memories from the polycomb group proteins. Annu. Rev. Genet. 46, 561–589 (2012).
Qüesta, J. I., Song, J., Geraldo, N., An, H. & Dean, C. Arabidopsis transcriptional repressor VAL1 triggers Polycomb silencing at FLC during vernalization. Science 353, 485–488 (2016).
Yuan, W. et al. A cis cold memory element and a trans epigenome reader mediate Polycomb silencing of FLC by vernalization in Arabidopsis. Nat. Genet. 48, 1527–1534 (2016).
Berger, N., Dubreucq, B., Roudier, F., Dubos, C. & Lepiniec, L. Transcriptional regulation of Arabidopsis. LEAFY COTYLEDON2 involves RLE, a cis-element that regulates trimethylation of histone H3 at lysine-27. Plant Cell 23, 4065–4078 (2011).
Lodha, M., Marco, C. F. & Timmermans, M. C. P. The ASYMMETRIC LEAVES complex maintains repression of KNOX homeobox genes via direct recruitment of Polycomb-repressivecomplex2. Genes Dev. 27, 596–601 (2013).
Merini, W. et al. The Arabidopsis Polycomb repressive complex 1 (PRC1) components AtBMI1A, B, and C impact gene networks throughout all stages of plant development. Plant Physiol. 173, 627–641 (2017).
Yang, C. et al. VAL- and AtBMI1-mediated H2Aub initiate the switch from embryonic to postgerminative growth in Arabidopsis. Curr. Biol. 23, 1324–1329 (2013).
Xiao, J. et al. Cis and trans determinants of epigenetic silencing by Polycomb repressive complex 2 in Arabidopsis. Nat. Genet. 49, 1546–1552 (2017).
Hecker, A. et al. The Arabidopsis GAGA-binding factor BASIC PENTACYSTEINE6 recruits the POLYCOMB-REPRESSIVE COMPLEX1 component LIKE HETEROCHROMATIN PROTEIN1 to GAGA DNA motifs. Plant Physiol. 168, 1013–1024 (2015).
Zhou, Y., Hartwig, B., James, G. V., Schneeberger, K. & Turck, F. Complementary activities of TELOMERE REPEAT BINDING proteins and Polycomb group complexes in transcriptional regulation of target genes. Plant Cell 28, 87–101 (2016).
Hyun, Y. et al. Site-directed mutagenesis in Arabidopsis thaliana using dividing tissue-targeted RGEN of the CRISPR/Cas system to generate heritable null alleles. Planta 241, 271–284 (2015).
Schrumpfová, P. P. et al. Telomere repeat binding proteins are functional components of Arabidopsis telomeres and interact with telomerase. Plant J. 77, 770–781 (2014).
Lee, W. K. & Cho, M. H. Telomere-binding protein regulates the chromosome ends through the interaction with histone deacetylases in Arabidopsis thaliana. Nucleic Acids Res. 44, 4610–4624 (2016).
Wang, H. et al. Arabidopsis flower and embryo developmental genes are repressed in seedlings by different combinations of Polycomb group proteins in association with distinct sets of cis-regulatory elements. PLoS Genet. 12, e1005771 (2016).
Turck, F. et al. Arabidopsis TFL2/LHP1 specifically associates with genes marked by trimethylation of histone H3 lysine 27. PLoS Genet. 3, e86 (2007).
Chanvivattana, Y. et al. Interaction of Polycomb-group proteins controlling flowering inArabidopsis. Development 131, 5263–5276 (2004).
Schrumpfová, P. P. et al. Telomere binding protein TRB1 is associated with promoters of translation machinery genes in vivo. Plant Mol. Biol. 90, 189–206 (2016).
Deng, W. et al. Arabidopsis Polycomb Repressive Complex 2 binding sites contain putative GAGA factor binding motifs within coding regions of genes. BMC Genomics 14, 593 (2013).
Trémousaygue, D. et al. Internal telomeric repeats and ‘TCP domain’ protein-binding sites co-operate to regulate gene expression in Arabidopsis thaliana cycling cells. Plant J. 33, 957–966 (2003).
Mozgová, I., Schrumpfová, P. P., Hofr, C. & Fajkus, J. Functional characterization of domains in AtTRB1, a putative telomere-binding protein in Arabidopsis thaliana. Phytochemistry 69, 1814–1819 (2008).
Kuchar, M. & Fajkus, J. Interactions of putative telomere-binding proteins in Arabidopsis thaliana: identification of functional TRF2 homolog in plants. FEBS Lett. 578, 311–315 (2004).
Liu, C., Xi, W., Shen, L., Tan, C. & Yu, H. Regulation of floral patterning by flowering time genes. Dev. Cell 16, 711–722 (2009).
Lee, W. K., Yun, J. H., Lee, W. & Cho, M. H. DNA-binding domain of AtTRB2 reveals unique features of a single Myb histone protein family that binds to both Arabidopsis- and human-type telomericDNA sequences. Mol. Plant 5, 1406–1408 (2012).
Yun, J. H. et al. Solution structure of telomere binding domain of AtTRB2 derived from Arabidopsis thaliana. Biochem. Biophys. Res. Commun. 452, 436–442 (2014).
Sander, J. D. & Joung, J. K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Shao, Z., Zhang, Y., Yuan, G. C., Orkin, S. H. & Waxman, D. J. MAnorm: a robust model for quantitative comparison of ChIP-Seq data sets. Genome Biol. 13, R16 (2012).
Heger, A., Webber, C., Goodson, M., Ponting, C. P. & Lunter, G. GAT: a simulation framework for testing the association of genomic intervals. Bioinformatics 29, 2046–2048 (2013).
Yilmaz, A. et al. AGRIS: the Arabidopsis Gene Regulatory Information Server, an update. Nucleic Acids Res. 39, D1118–D1122 (2011).
Mathelier, A. et al. JASPAR 2014: an extensively expanded and updated open-access database of transcription factor binding profiles. Nucleic Acids Res. 42, D142–D147 (2014).
Higo, K., Ugawa, Y., Iwamoto, M. & Korenaga, T. Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res. 27, 297–300 (1999).
Hehl, R. & Bülow, L. AthaMap web tools for the analysis of transcriptional and posttranscriptional regulation of gene expression in Arabidopsis thaliana. Methods Mol. Biol. 1158, 139–156 (2014).
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Samson, F. et al. FLAGdb/FST: a database of mapped flanking insertion sites (FSTs) of Arabidopsis thaliana T-DNA transformants. Nucleic Acids Res. 30, 94–97 (2002).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Reimer, J. J. & Turck, F. Genome-wide mapping of protein-DNA interaction by chromatin immunoprecipitation and DNA microarray hybridization (ChIP-chip). Part A: ChIP-chip molecular methods. Methods Mol. Biol. 631, 139–160 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Zang, C. et al. A clustering approach for identification of enriched domains from histone modification ChIP-Seq data. Bioinformatics 25, 1952–1958 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Anders, S., Pyl, P. T. & Huber, W. HTSeq: a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Fisher, R. A. A new test for 2 × 2 tables. Nature 156, 388 (1945).
Sheen, J. Signal transduction in maize and Arabidopsis mesophyll protoplasts. Plant Physiol. 127, 1466–1475 (2001).
Hu, C. D. & Kerppola, T. K. Simultaneous visualization of multiple protein interactions in living cells using multicolor fluorescence complementation analysis. Nat. Biotechnol. 21, 539–545 (2003).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Acknowledgements
We thank P. Taenzler for excellent technical help and A. Brazel for critical reading of the manuscript. This work was supported by core funding from the Max Planck Society (Y. Zhou, K.K., J.A.D., and F.T.), the Chinese Scholarship Council (T.Y.) and the National Natural Science Foundation of China (grant no. 31570319; Y.W. and Y. Zhang). We thank K. Riha (Gregor Mendel Institute of Molecular Plant Biology) and J. Goodrich (University of Edinburgh) for providing materials.
Author information
Authors and Affiliations
Contributions
Y. Zhou performed and supervised all experimental work; K.K. performed the functional analysis of the SEP3 gene; J.A.D. assisted in Y2H assays and coimmunoprecipitation; K.K. and J.A.D. contributed to the development of theTRB2 CRISPR–CAS9 line; K.K. and T.Y. contributed the protoplast work; Y. Zhang and Y.W. carried out all bioinformatics analysis; and Y. Zhou, K.K., and F.T. planned the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Phenotypes of trb, clf, and trb clf mutants.
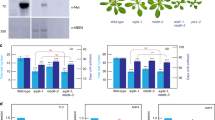
(a) Phenotype of 28-days-old Col-0, trb1-2, trb3-2, trb1-2 trb3-2, clf-28, trb1-2 clf-28, trb3-2 clf-28 and trb1-2 trb3-2 clf-2 plants grown at 22 °C in long-day (LD) conditions. Scale bar indicates 1 cm. (b) Flowering time scored for genotypes in (a) as number of leaves and average of rosette size at bolting. Error bars indicate mean ± s.e.m. (n = 9). Statistical significance was determined by one-way analysis of variance (ANOVA) and multiple comparison by the Holm–Sidak method (P < 0.05). Different letters indicate significance groups. (c) Phenotype of 28-day-old Ws-2, trb2-1, clf-50 and trb2-1 clf-50 plants grown at 22 °C in LD conditions. Scale bar indicates 1 cm. (d and e) Flowering time scored for genotypes in (c) as number of leaves and average rosette size at bolting. Significance tested as in (b). (f) Expression of TRB2 in Ws-2 and trb2-1 mutant backgrounds measured by qRT-PCR. PP2A was used as reference gene. The presence of full-length transcripts was analyzed.
Supplementary Figure 2 Phenotype of higher-order trb mutants and complementation of triple trb1/2/3.
(a) Phenotype of 10-day-old seedlings of Col-0, trb1/2, trb1/3 and trb2/3 mutants grown at 22 °C in LD conditions. Scale bar: 1 cm. (b) Phenotype of 10-day-old seedlings of Col-0, trb1/2/3, and gTRB2-Flag transgenic lines (L12 and L21) in trb1/2/3 background. Scale bar: 1 cm. (c) +1 insertion (2nd exon of TRB2) in trb2-2 line is identified in trb1/3 double mutant background. The predicted DNA cleavage site is marked with an arrow. (d) Phenotype of 10-day-old seedlings of Col-0, trb1/2/3 and trb2-2 trb1/3 mutants. Scale bar: 1 cm.
Supplementary Figure 3 Nuclear localization of TRB proteins and their effect on teleomere integrity.
(a) Co-localization of TRB2 and TRB1 in the nucleus via transient expression of TRB2-GFP and TRB1-RFP in Tobacco leaves. Scale bar indicates 10 μM. (b) Analysis of pixel density along the axis indicated by a white line in (a) was performed by ImageJ. GFP signal intensities are shown in green and RFP signals in red. (c,d) gTRB2-YFP (c) and gTRB3-YFP (d) fusion proteins are detected in the nuclei of almost all cells in transgenic Arabidopsis plants. Yellow fluorescence detected in the root tip (upper panel) and the first true leaves (lower panel). Scale bars: 50 µm. (e) Telomere length measured by TRF analysis of genomic DNA prepared from 10-day-old Col-0, Ws-2, ku70, fifth generation (G5) tert, second (G2), and third (G3) generation of trb1/2/3 or trb1/3 trb2-/+ seedlings with ku70 and tert being included as reference.
Supplementary Figure 4 Correlation between H3K27me3 and expression changes for genes misregulated in trb1/2/3.
(a) Boxplot shows H3K27me3 ChIP-seq read intensity across genes commonly down- or up-regulated in trb123 and clf swn compared to all annotated genes. Venn diagrams are a reproduced from Fig. 1d for better clarity. A Hypergeometric test was used for statistical analysis. (b) Scatter plot showing the negative correlation between H3K27me3 and expression changes for genes mis-regulated in trb1/2/3.
Supplementary Figure 5 Quantitative PCR validation of RNA-seq and H3K27me3 ChIP–seq data in Col-0 and trb1/2/3 seedlings.
(a) Genes up-regulated in trb1/2/3 mutants. (b) Genes down-regulated in trb1/2/3 mutants. RNA was extracted from 10 day-old seedlings grown at 22 °C in LDs. Values were normalized to the reference gene PP2A using the –ΔΔCT approach. Vertical central line and whiskers indicate mean ± s. d. of four independently grown and processed replicate experiments used for qRT-PCR, symbols indicate data points as averages of three technical qPCR replicates. Statistical significance was determined by a Mann–Whitney–Wilcoxon test (df = 6) (c) H3K27me3 level change in Col-0 biased genes. (d) H3K27me3 level change in trb1/2/3 biased genes. (e) H3K27me3 level in common genes. Vertical central line and whiskers indicate mean ± s. d. of three independently grown and processed replicate experiments used for ChIP-qPCR, symbols represent averages of three technical qPCR replicates. Statistical significance was determined by a Mann–Whitney–Wilcoxon test (df = 4).
Supplementary Figure 6 List of cis elements enriched in different H3K27me3 gene categories detected in trb1/2/3 mutants.
Sequence logos of the 15 most enriched elements detected by MEME analysis.
Supplementary Figure 7 ChIP–qPCR analysis of H3K27me3 coverage in transiently FIE-GFP- and TRB1-GFP-expressing protoplasts.
(a) Western blot of GFP tagged FIE or TRB1 proteins expressed in protoplasts prepared from Col-0 and trb1/2/3-2 or Col-0 and clf-28. Anti-H3 was used as a loading control. Displayed is a representative blot of three replicated experiments. See Supplementary Fig. 14 for full scan of cropped image. (b) H3K27me3 level at SEP3, EMF1, AG and ACT2 in protoplast prepared from Col-0 and clf-28 seedlings. ACT2 was used as negative control for H3K27me3 distribution. Vertical central line and whiskers indicate mean ± s. d. of ChIP-qPCR of three independently transfected and processed batches of protoplasts. Statistical significance was determined by Mann–Whitney–Wilcoxon test (df = 4).
Supplementary Figure 8 Interaction of TRBs with CLF and SWN.
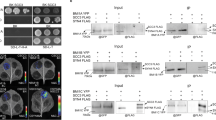
(a) TRB1-nYFP co-infiltrated with cYFP-CLF (1) or ATJ3 (2) and ATJ3 co-infiltrated with cYFP-CLF (3) in Nicotiana tabacum leaves. LHP1-RFP was used as reference for co-infiltration and ATJ3 was used as control for interactions. Scale bar = 10 μM. (b) Percentage of RFP expressing nuclei as in (a) expressing YFP and RFP signal. Total cells = 100, counted in 5 independently infiltrated leaves; Statistical analysis was a one-tailed Mann–Whitney U-test (df = 8). Vertical central line and whiskers indicate mean ± s. d. (c) Co-immunoprecipitation of CLF and (b) SWN with TRB1-3 expressed in Arabidopsis mesophyll protoplasts. TRB1-GFP, TRB2-GFP or TRB3-GFP were immunoprecipitated with anti-GFP trap beads from protoplasts co-transfected with either HA-CLF or HA-SWN. The precipitates were analyzed by western blotting with anti-GFP and anti-HA antibodies. See Supplementary Fig. 14 for full scan of cropped images.
Supplementary Figure 9 Interaction of TRBs with CLF and SWN in yeast cells.
(a) GAL4 DNA-binding domain (BD) or BD-CLF and BD-SWN fusion proteins were tested for their ability to bind to an activation domain (AD) fused to different TRB proteins. (b) Mutual interactions between TRB1-3 were tested as in (a). (c) Schematic presentation of the TRB protein domain structure. (d-e) Interaction between the MYB, H1/5, or Coiled-coil domain of TRB3 and CLF (d) and SWN (e). (f) Interaction between the MYB, H1/5, or Coiled-coil domain of TRB1 with SWN. Growth shown for three dilutions (10−1, 10−2 and 10−3) of the yeast culture on SD medium lacking LW (-LW) and LWH (-LWH) in all panels.
Supplementary Figure 10 Histochemical detection of GUS activity in transgenic SEP3pro-GUS lines with or without an intact telobox.
(a) 10-day-old seedlings carrying wild type pSEP3pro-WT-GUS. (b) 10-day-old seedlings carrying mutated SEP3pro-M-GUS. Picture in (a-b) show representative pattern observed in four single copy insertion lines transformed in the Col-0 background per construct and (c) an untransformed Col-0 control. (d) GUS activity at the inflorescence of representative SEP3pro-WT-GUS and (e) SEP3pro-M-GUS line. Scale bars in all panels indicate 1 mm.
Supplementary Figure 11 ChIP–qPCR analysis of TRB2 or TRB3 binding to the SEP3 promoter.
Distal promoter region of FT was used to as negative region for TRB2 or TRB3 binding. Bars display the mean of three biological replicates used for ChIP-qPCR, Vertical central line and whiskers indicate mean ± s. d. Statistical significance was determined by Mann–Whitney–Wilcoxon test (df = 4).
Supplementary Figure 12 TRB binding is decreased at a SEP3 promoter carrying a mutated telobox element.
(a) ChIP-PCR of TRB1-GFP binding to pSEP3pro-WT-GUS or pSEP3pro-M-GUS transgenic lines; empty plasmids were used as a negative transfection control. Three independent biological experiments are shown. Note that PCR fragment necessary to distinguish the endogenous and transgenic SEP3 promoter is 800 bp long and therefore not compatible with qPCR. (b) Coomassie brilliant blue staining of recombinant His-TRB3 protein purified from E.coli. (c) Illustration depicting the SEP3 promoter region. Blue line indicates telobox; yellow lines indicate four telobox-related motifs; black line indicates fragment used for EMSA. (d) TRB3 binding to Cy5 labelled PCR fragments of SEP3 promoter with WT or mutated telobox.
Supplementary Figure 13 AZF1- and TRB1-target genes do not show significant overlap.
Numbers of significant AZF1 and TRB1 target genes overlapping with Col-0 biased genes. The p-value of the only significant overlap is indicated (Hypergeometric test).
Supplementary Figure 14
Original uncropped scans of representative western blots displayed in this manuscript.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Table 1
Supplementary Data Set 1: Summary table of gene expression and H3K27me3 changes in trb1/2/3 for all Arabidopsis genes annotated in TAIR10
Sheet 1: Columns indicate 1: Presence or absence of teloboxes, 2: differential expression in trb1/2/3, 3: differential expression in clf/swn, 4: TRB1 binding according to Zhou et al. (2016)5, 5: TRB1 binding according to the present analysis, 6: H3K27me3 change in trb1/2/3 seedlings. Sheet 2: Gene expression fold-changes in trb1/2/3, clf-29swn-21 and siFIE mutants.
Supplementary Data Set 2: Summary table for GO enrichment analysis of genes differentially expressed in trb1/2/3 seedlings
Columns indicate 1: Cluster type, 2: enriched GO-term, 3: GO-term description, 4: GO-term count, 5: %, 6: p-value, 7: AGI codes, 8: length of list, 9: hits in list, 10: total gene list, 11: enrichment, 12: Bonferroni corrected p-value, 13: Benjamini corrected p-value, 14: FDR corrected p-value.
Supplementary Data Set 3: Summary table of MAnorm analysis for regions differentially enriched in H3K27me3
Columns indicate 1: Chromosome, 2: start of peak, 3: end of peak, 4: position of peak summit, 5: M-value, 6: A-value, 7: pvalue, 8: peak category, 9: normalized reads, 10: normalized read density in trb1/2/3.
Supplementary Data Set 4: Summary table for GO enrichment analysis of Col-0 biased and trb1/2/3 biased genes
Columns indicate 1: Cluster type, 2: enriched GO-term, 3: GO-term description, 4: GO-term count, 5: %, 6: p-value, 7: AGI codes, 8: length of list, 9: hits in list, 10: total gene list, 11: enrichment, 12: Bonferroni corrected p-value, 13: Benjamini corrected p-value, 14: FDR corrected p-value.
Supplementary Data Set 5: Genomic coordinates of FIE and TRB1 enriched peaks
Columns indicate 1: Chromosome, 2: start of peak, 3: end of peak, 4: length, 5: position of peak summit, 6: read tags, 7: corrected p-Value, 8: enrichment.
Rights and permissions
About this article
Cite this article
Zhou, Y., Wang, Y., Krause, K. et al. Telobox motifs recruit CLF/SWN–PRC2 for H3K27me3 deposition via TRB factors in Arabidopsis. Nat Genet 50, 638–644 (2018). https://doi.org/10.1038/s41588-018-0109-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0109-9
This article is cited by
-
The UBP5 histone H2A deubiquitinase counteracts PRCs-mediated repression to regulate Arabidopsis development
Nature Communications (2024)
-
Identification of vernalization-related genes and cold memory element (CME) required for vernalization response in radish (Raphanus sativus L.)
Plant Molecular Biology (2024)
-
A VEL3 histone deacetylase complex establishes a maternal epigenetic state controlling progeny seed dormancy
Nature Communications (2023)
-
Epigenetic silencing of callose synthase by VIL1 promotes bud-growth transition in lily bulbs
Nature Plants (2023)
-
LHP1-mediated epigenetic buffering of subgenome diversity and defense responses confers genome plasticity and adaptability in allopolyploid wheat
Nature Communications (2023)