Abstract
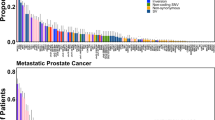
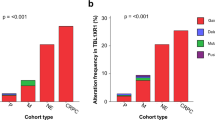
Comprehensive genomic characterization of prostate cancer has identified recurrent alterations in genes involved in androgen signaling, DNA repair, and PI3K signaling, among others. However, larger and uniform genomic analysis may identify additional recurrently mutated genes at lower frequencies. Here we aggregate and uniformly analyze exome sequencing data from 1,013 prostate cancers. We identify and validate a new class of E26 transformation-specific (ETS)-fusion-negative tumors defined by mutations in epigenetic regulators, as well as alterations in pathways not previously implicated in prostate cancer, such as the spliceosome pathway. We find that the incidence of significantly mutated genes (SMGs) follows a long-tail distribution, with many genes mutated in less than 3% of cases. We identify a total of 97 SMGs, including 70 not previously implicated in prostate cancer, such as the ubiquitin ligase CUL3 and the transcription factor SPEN. Finally, comparing primary and metastatic prostate cancer identifies a set of genomic markers that may inform risk stratification.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
31 May 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Robinson, D. et al. Integrative clinical genomics of advanced prostate cancer. Cell 161, 1215–1228 (2015).
Cancer Genome Atlas Research Network. Molecular taxonomy of primary prostate cancer. Cell 163, 1011–1025 (2015).
Tomlins, S. A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Barbieri, C. E. et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nat. Genet. 44, 685–689 (2012).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Beltran, H. et al. Divergent clonal evolution of castration-resistant neuroendocrine prostate cancer. Nat. Med. 22, 298–305 (2016).
Kumar, A. et al. Substantial interindividual and limited intraindividual genomic diversity among tumors from men with metastatic prostate cancer. Nat. Med. 22, 369–378 (2016).
Baca, S. C. et al. Punctuated evolution of prostate cancer genomes. Cell 153, 666–677 (2013).
Hieronymus, H. et al. Copy number alteration burden predicts prostate cancer relapse. Proc. Natl. Acad. Sci. USA 111, 11139–11144 (2014).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, 43–49 (2013).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
Theurillat, J.-P. P. et al. Ubiquitylome analysis identifies dysregulation of effector substrates in SPOP-mutant prostate cancer. Science 346, 85–89 (2014).
Geng, C. et al. Prostate cancer–associated mutations in speckle-type POZ protein (SPOP) regulate steroid receptor coactivator 3 protein turnover. Proc. Natl. Acad. Sci. USA 110, 6997–7002 (2013).
Yuan, W.-C. et al. A Cullin3–KLHL20 ubiquitin ligase–dependent pathway targets PML to potentiate HIF-1 signaling and prostate cancer progression. Cancer Cell 20, 214–228 (2011).
Groner, A. C. et al. TRIM24 is an oncogenic transcriptional activator in prostate cancer. Cancer Cell 29, 846–858 (2016).
Boysen, G. et al. SPOP mutation leads to genomic instability in prostate cancer. eLife 4, e09207 (2015).
Abida, W. et al. Prospective genomic profiling of prostate cancer across disease states reveals germline and somatic alterations that may affect clinical decision making. JCO Precis. Oncol. https://doi.org/10.1200/PO.17.00029 (2017).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Dolatshad, H. et al. Disruption of SF3B1 results in deregulated expression and splicing of key genes and pathways in myelodysplastic syndrome hematopoietic stem and progenitor cells. Leukemia 29, 1092–1103 (2015).
Ciriello, G. et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell 163, 506–519 (2015).
Papaemmanuil, E. et al. Somatic SF3B1 mutation in myelodysplasia with ring sideroblasts. N. Engl. J. Med. 365, 1384–1395 (2011).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Shi, Y. et al. Sharp, an inducible cofactor that integrates nuclear receptor repression and activation. Genes Dev. 15, 1140–1151 (2001).
Légaré, S. et al. The estrogen receptor cofactor SPEN functions as a tumor suppressor and candidate biomarker of drug responsiveness in hormone-dependent breast cancers. Cancer Res. 75, 4351–4363 (2015).
Kuchay, S. et al. FBXL2- and PTPL1-mediated degradation of p110-free p85β regulatory subunit controls the PI(3)K signalling cascade. Nat. Cell. Biol. 15, 472–480 (2013).
Fraser, M. et al. Genomic hallmarks of localized, non-indolent prostate cancer. Nature 541, 359–364 (2017).
Cibulskis, K. et al. ContEst: estimating cross-contamination of human samples in next-generation sequencing data. Bioinformatics 27, 2601–2602 (2011).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Costello, M. et al. Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res. 41, e67 (2013).
Van Allen, E. M. et al. Whole-exome sequencing and clinical interpretation of formalin-fixed, paraffin-embedded tumor samples to guide precision cancer medicine. Nat. Med. 20, 682–688 (2014).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor–normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Chang, M. T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131 (2016).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Cheng, D. T. et al. Memorial Sloan Kettering–Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J. Mol. Diagn. 17, 251–264 (2015).
Chakravarty, D. OncoKB: a precision oncology knowledge base. Precis. Oncol. https://doi.org/10.1200/PO.17.00011 (2017).
McGranahan, N. et al. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci. Transl. Med. 7, 283ra54 (2015).
Hartmaier, R. J. et al. High-throughput genomic profiling of adult solid tumors reveals novel insights into cancer pathogenesis. Cancer. Res. 77, 2464–2475 (2017).
Acknowledgements
We thank the patients for participating in this study. We also thank the Broad Cancer Genome Analysis and Data Sciences groups for analysis methodology and computational support. This work was supported by the SU2C-PCF Prostate Cancer International Dream Team, Prostate Cancer Foundation Young Investigator Awards (B.S.T., C.P., N.S., and E.M.V.A.), Prostate Cancer Foundation–V Foundation Challenge Award (E.M.V.A., P.S.N., and J.S.d.B.), NIH K08CA188615 (E.M.V.A.), NCI P50-CA097186 and NCI P50-CA92629 SPOREs in Prostate Cancer, the Marie-Josée and Henry R. Kravis Center for Molecular Oncology, a National Cancer Institute Cancer Center Core Grant (P30-CA008748), and the Robertson Foundation (B.S.T. and N.S.).
Author information
Authors and Affiliations
Consortia
Contributions
J.A., S.A.M.W., N.S., E.M.V.A., D.L., J.G., R.K., E.R., W.K.C., D.C., G.C.H., C.E.B., A.S., C.M.B., A.V.P., C.T., F.D., M.A.R., and B.S.T. contributed with algorithm development and analysis of genomic data. R.L., L.A.G., I.C., B.M., C.P., C.M., H.B., Z.Z., S.M., F.W.H., D.R., Y.M.W., P.W.K., M.-E.T., W.A., H.I.S., P.S.N., J.S.d.B., M.A.R., C.L.S., and A.M.C. developed the patient cohort, obtained tumor biopsies, performed molecular testing for metastatic cases, and carried out data interpretation of the overall cohort. J.A., S.A.M.W., D.L., N.S., and E.M.V.A. performed final aggregate cohort assembly, mutation review, interpretation, and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
E.M.V.A. is a consultant for Tango Therapeutics and Genome Medical.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1
Workflow of sample inclusion and quality control.
Supplementary Figure 2 Correlates of genomic burden.
Metastatic tumors have increased mutational and copy number burden as compared to primary tumors, adjusted for differences in purity and coverage. In primary tumors, increased age at diagnosis and higher Gleason score are associated with higher mutational and copy number burden, adjusting for purity and coverage. Reported P values for each predictor (metastatic versus primary disease; age; Gleason score) are from the multivariate regression adjusting for purity and coverage. The center values represent the median value of each group and the error bars below or above the median line define the first and third quartile respectively.
Supplementary Figure 3 Schematic workflow of the combination of statistical, quality and biological filters used to identify SMGs.
Flowchart describing how we applied a combination of statistical, quality, and biological filters to the results to identify SMGs. Approach to defining significantly mutated genes using both statistical and biological filters.
Supplementary Figure 4 Allele frequencies and mRNA expression of significantly mutated genes.
(a) Allele frequencies of significantly mutated genes. Boxplot showing the distribution of allele frequencies across SMGs sorted by decreasing median allele frequency. (b) mRNA expression of the SMGs across the TCGA cohort. mRNA expression of the SMGs across the PCF-SU2C cohort. The red center line in the boxplots indicates the median value.
Supplementary Figure 5 Supervised hierarchical clustering on arm-level deletions.
Samples (rows) are ordered according to six copy number clusters derived from hierarchical clustering on arm-level deletions (2q, 5q, 6q, 8p, 13q, and 16q), with SPOP and CUL3 mutations indicated in green. Chromosomes are shown from left to right, samples from top to bottom. Regions of loss are indicated by shades of blue, and gains are indicated by shades of red.
Supplementary Figure 6 Additional mutation characteristics of individual SMGs.
(a) CUL3 mutations observed in our cohort. (b) Mutations observed in PIK3R2. Hotspot mutations observed in PIK3R2 are paralogous to the oncogenic D560 mutation in PIK3R1. (c) Mutation distribution of CDK12 variants, showing that the majority of mutations are truncating and that missense variants cluster in the kinase domain.
Supplementary Figure 7 The overlap of the three bait sets used in this analysis (Agilent, Ilumina, NimbleSeq).
(a) Overlap of all bases. (b) Overlap of coding regions.
Supplementary Figure 8 Segmented copy-number profiles of 303 primary tumors from TCGA.
(a) Whole exome sequencing (top) vs Affymetrix SNP6 data (bottom). 303 tumors are shown for data set, sorted in identical order. Regions of gain are shown in shades of red, losses as shades of blue. The center summary plots show the fraction of samples with a log2(cn/2) value >0.1 (red) or <-0.1 (blue) at a given position. (b) Scatter plot to compare the segment means of matched segments >200KB from the SNP6 and the ReCapSeg data, resulting in a pearson correlation of 0.92 (C.I. 0.916-0.918). The graph shows the correlation between the inferred copy number of 29,290 microarray segments with size > 200 KB with the corresponding inferred copy number from the WES. The segments with size > 200KB represent 99.94% of the covered genome from the SNP array.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 5 and 7, and Supplementary Note
Supplementary Table 1
Source of cohorts used for this analysis
Supplementary Table 2
Complete list of all somatic mutations in this cohort
Supplementary Table 3
Patient-level information
Supplementary Table 4
Mutational significance analysis
Supplementary Table 6
Genes in cancer pathways
Supplementary Table 8
Intersected BED file
Supplementary Table 9
Unfiltered MAF file
Supplementary Table 10
Matrix of copy number calls
Rights and permissions
About this article
Cite this article
Armenia, J., Wankowicz, S.A.M., Liu, D. et al. The long tail of oncogenic drivers in prostate cancer. Nat Genet 50, 645–651 (2018). https://doi.org/10.1038/s41588-018-0078-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0078-z
This article is cited by
-
EXO1/P53/SREBP1 axis-regulated lipid metabolism promotes prostate cancer progression
Journal of Translational Medicine (2024)
-
CRISPR/Cas9 model of prostate cancer identifies Kmt2c deficiency as a metastatic driver by Odam/Cabs1 gene cluster expression
Nature Communications (2024)
-
In vivo genome-wide CRISPR screening identifies CITED2 as a driver of prostate cancer bone metastasis
Oncogene (2024)
-
The yin and yang of chromosomal instability in prostate cancer
Nature Reviews Urology (2024)
-
Mechanism-centric regulatory network identifies NME2 and MYC programs as markers of Enzalutamide resistance in CRPC
Nature Communications (2024)