Abstract
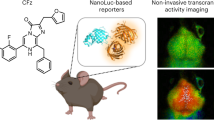
Measurement of gene expression in the brain requires invasive analysis of brain tissue or non-invasive methods that are limited by low sensitivity. Here we introduce a method for non-invasive, multiplexed, site-specific monitoring of endogenous gene or transgene expression in the brain through engineered reporters called released markers of activity (RMAs). RMAs consist of an easily detectable reporter and a receptor-binding domain that enables transcytosis across the brain endothelium. RMAs are expressed in the brain but exit into the blood, where they can be easily measured. We show that expressing RMAs at a single mouse brain site representing approximately 1% of the brain volume provides up to a 100,000-fold signal increase over the baseline. Expression of RMAs in tens to hundreds of neurons is sufficient for their reliable detection. We demonstrate that chemogenetic activation of cells expressing Fos-responsive RMA increases serum RMA levels >6-fold compared to non-activated controls. RMAs provide a non-invasive method for repeatable, multiplexed monitoring of gene expression in the intact animal brain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that all data supporting the results in this study are available within the paper, its Supplementary Information and its Source Data file. Microscopy images are available from the corresponding author upon reasonable request owing to their large size and number. The plasmids designed in this study are available on Addgene (https://www.addgene.org/browse/article/28229133/). Source data are provided with this paper.
References
Richiardi, J. et al. Correlated gene expression supports synchronous activity in brain networks. Science 348, 1241–1244 (2015).
Minatohara, K., Akiyoshi, M. & Okuno, H. Role of immediate-early genes in synaptic plasticity and neuronal ensembles underlying the memory trace. Front. Mol. Neurosci. 8, 78 (2016).
Seidlitz, J. et al. Transcriptomic and cellular decoding of regional brain vulnerability to neurogenetic disorders. Nat. Commun. 11, 3358 (2020).
Genove, G., DeMarco, U., Xu, H., Goins, W. F. & Ahrens, E. T. A new transgene reporter for in vivo magnetic resonance imaging. Nat. Med. 11, 450–454 (2005).
Shapiro, M. G. et al. Directed evolution of a magnetic resonance imaging contrast agent for noninvasive imaging of dopamine. Nat. Biotechnol. 28, 264–270 (2010).
Sigmund, F. et al. Bacterial encapsulins as orthogonal compartments for mammalian cell engineering. Nat. Commun. 9, 1990 (2018).
Schilling, F. et al. MRI measurements of reporter-mediated increases in transmembrane water exchange enable detection of a gene reporter. Nat. Biotechnol. 35, 75–80 (2017).
Mukherjee, A., Wu, D., Davis, H. C. & Shapiro, M. G. Non-invasive imaging using reporter genes altering cellular water permeability. Nat. Commun. 7, 13891 (2016).
Farhadi, A., Sigmund, F., Westmeyer, G. G. & Shapiro, M. G. Genetically encodable materials for non-invasive biological imaging. Nat. Mater. 20, 585–592 (2021).
Farhadi, A., Ho, G. H., Sawyer, D. P., Bourdeau, R. W. & Shapiro, M. G. Ultrasound imaging of gene expression in mammalian cells. Science 365, 1469–1475 (2019).
Lu, G. J. et al. Acoustically modulated magnetic resonance imaging of gas-filled protein nanostructures. Nat. Mater. 17, 456–463 (2018).
Gottschalk, S. et al. Rapid volumetric optoacoustic imaging of neural dynamics across the mouse brain. Nat. Biomed. Eng. 3, 392–401 (2019).
Marvin, J. S. et al. A genetically encoded fluorescent sensor for in vivo imaging of GABA. Nat. Methods 16, 763–770 (2019).
Ntziachristos, V. Going deeper than microscopy: the optical imaging frontier in biology. Nat. Methods 7, 603–614 (2010).
Iwano, S. et al. Single-cell bioluminescence imaging of deep tissue in freely moving animals. Science 359, 935–939 (2018).
de Wildt, R. M., Mundy, C. R., Gorick, B. D. & Tomlinson, I. M. Antibody arrays for high-throughput screening of antibody–antigen interactions. Nat. Biotechnol. 18, 989–994 (2000).
Shendure, J. & Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 26, 1135–1145 (2008).
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Naumova, O. Y., Lee, M., Rychkov, S. Y., Vlasova, N. V. & Grigorenko, E. L. Gene expression in the human brain: the current state of the study of specificity and spatiotemporal dynamics. Child Dev. 84, 76–88 (2013).
Kang, H. J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Roopenian, D. C. & Akilesh, S. FcRn: the neonatal Fc receptor comes of age. Nat. Rev. Immunol. 7, 715–725 (2007).
Rissin, D. M. et al. Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations. Nat. Biotechnol. 28, 595–599 (2010).
Swaminathan, J. et al. Highly parallel single-molecule identification of proteins in zeptomole-scale mixtures. Nat. Biotechnol. 36, 1076–1082 (2018).
Sze, J. Y., Ivanov, A. P., Cass, A. E. G. & Edel, J. B. Single molecule multiplexed nanopore protein screening in human serum using aptamer modified DNA carriers. Nat. Commun. 8, 1552 (2017).
Tannous, B. A., Kim, D.-E., Fernandez, J. L., Weissleder, R. & Breakefield, X. O. Codon-optimized Gaussia luciferase cDNA for mammalian gene expression in culture and in vivo. Mol. Ther. 11, 435–443 (2005).
Aalipour, A. et al. Engineered immune cells as highly sensitive cancer diagnostics. Nat. Biotechnol. 37, 531–539 (2019).
Deane, R. et al. IgG-assisted age-dependent clearance of Alzheimer’s amyloid β peptide by the blood–brain barrier neonatal Fc receptor. J. Neurosci. 25, 11495–11503 (2005).
Cooper, P. R. et al. Efflux of monoclonal antibodies from rat brain by neonatal Fc receptor, FcRn. Brain Res. 1534, 13–21 (2013).
Zhang, Y. & Pardridge, W. M. Mediated efflux of IgG molecules from brain to blood across the blood–brain barrier. J. Neuroimmunol. 114, 168–172 (2001).
Borrok, M. J. et al. pH-dependent binding engineering reveals an FcRn affinity threshold that governs IgG recycling. J. Biol. Chem. 290, 4282–4290 (2015).
Westerink, R. H. S. & Ewing, A. G. The PC12 cell as model for neurosecretion. Acta Physiol. 192, 273–285 (2008).
Verhaegen, M. & Christopoulos, T. K. Recombinant Gaussia luciferase. Overexpression, purification, and analytical application of a bioluminescent reporter for DNA hybridization. Anal. Chem. 74, 4378–4385 (2002).
Oganesyan, V. et al. Structural insights into neonatal Fc receptor-based recycling mechanisms. J. Biol. Chem. 289, 7812–7824 (2014).
Medesan, C., Matesoi, D., Radu, C., Ghetie, V. & Ward, E. S. Delineation of the amino acid residues involved in transcytosis and catabolism of mouse IgG1. J. Immunol. 158, 2211–2217 (1997).
Lee, C.-H. et al. An engineered human Fc domain that behaves like a pH-toggle switch for ultra-long circulation persistence. Nat. Commun. 10, 5031 (2019).
Herculano-Houzel, S., Mota, B. & Lent, R. Cellular scaling rules for rodent brains. Proc. Natl Acad. Sci. USA 103, 12138–12143 (2006).
Eixarch, H. et al. Transgene expression levels determine the immunogenicity of transduced hematopoietic grafts in partially myeloablated mice. Mol. Ther. 17, 1904–1909 (2009).
Pyzik, M., Kozicky, L. K., Gandhi, A. K. & Blumberg, R. S. The therapeutic age of the neonatal Fc receptor. Nat. Rev. Immunol. 23, 415–432 (2023).
Gil, G. A. et al. c-Fos activated phospholipid synthesis is required for neurite elongation in differentiating PC12 cells. Mol. Biol. Cell 15, 1881–1894 (2004).
Alexander, G. M. et al. Remote control of neuronal activity in transgenic mice expressing evolved G protein-coupled receptors. Neuron 63, 27–39 (2009).
Roth, B. L. DREADDs for neuroscientists. Neuron 89, 683–694 (2016).
Szablowski, J. O., Lee-Gosselin, A., Lue, B., Malounda, D. & Shapiro, M. G. Acoustically targeted chemogenetics for the non-invasive control of neural circuits. Nat. Biomed. Eng. 2, 475–484 (2018).
Sørensen, A. T. et al. A robust activity marking system for exploring active neuronal ensembles. eLife 5, e13918 (2016).
Bernau, K. et al. In vivo tracking of human neural progenitor cells in the rat brain using bioluminescence imaging. J. Neurosci. Methods 228, 67–78 (2014).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Kobayashi, N. FcRn-mediated transcytosis of immunoglobulin G in human renal proximal tubular epithelial cells. Am. J. Physiol. Renal Physiol 282, F358–F365 (2002).
Pyzik, M. et al. Hepatic FcRn regulates albumin homeostasis and susceptibility to liver injury. Proc. Natl Acad. Sci. USA 114, E2862–E2871 (2017).
Zetterberg, H. & Burnham, S. C. Blood-based molecular biomarkers for Alzheimer’s disease. Mol. Brain 12, 26 (2019).
Niederkofler, V. et al. Identification of serotonergic neuronal modules that affect aggressive behavior. Cell Rep. 17, 1934–1949 (2016).
Clifton, N. E. et al. Dynamic expression of genes associated with schizophrenia and bipolar disorder across development. Transl. Psychiatry 9, 74–74 (2019).
Tiklová, K. et al. Disease duration influences gene expression in neuromelanin-positive cells from Parkinsonas disease patients. Front. Mol. Neurosci 14, 763777 (2021).
Ham, S. & Lee, S.-J. V. Advances in transcriptome analysis of human brain aging. Exp. Mol. Med. 52, 1787–1797 (2020).
Pasi, K. J. et al. Multiyear follow-up of AAV5-hFVIII-SQ gene therapy for hemophilia A. N. Engl. J. Med. 382, 29–40 (2020).
Kim, H. J. et al. Stereotactic brain injection of human umbilical cord blood mesenchymal stem cells in patients with Alzheimer’s disease dementia: a phase 1 clinical trial. Alzheimers Dement. (N Y) 1, 95–102 (2015).
Duerinck, J. et al. Intracerebral administration of CTLA-4 and PD-1 immune checkpoint blocking monoclonal antibodies in patients with recurrent glioblastoma: a phase I clinical trial. J. Immunother. Cancer 9, e002296 (2021).
Goertsen, D. et al. AAV capsid variants with brain-wide transgene expression and decreased liver targeting after intravenous delivery in mouse and marmoset. Nat. Neurosci. 25, 106–115 (2022).
Thévenot, E. et al. Targeted delivery of self-complementary adeno-associated virus serotype 9 to the brain, using magnetic resonance imaging-guided focused ultrasound. Hum. Gene Ther. 23, 1144–1155 (2012).
Nouraein, S. et al. Acoustically targeted noninvasive gene therapy in large brain volumes. Gene Ther. https://doi.org/10.1038/s41434-023-00421-1 (2023).
McMahon, D., O’Reilly, M. A. & Hynynen, K. Therapeutic agent delivery across the blood–brain barrier using focused ultrasound. Annu. Rev. Biomed. Eng. 23, 89–113 (2021).
Lipsman, N. et al. Blood–brain barrier opening in Alzheimer’s disease using MR-guided focused ultrasound. Nat. Commun. 9, 2336 (2018).
Alfaro, J. A. et al. The emerging landscape of single-molecule protein sequencing technologies. Nat. Methods 18, 604–617 (2021).
Ying, T., Feng, Y., Wang, Y., Chen, W. & Dimitrov, D. S. Monomeric IgG1 Fc molecules displaying unique Fc receptor interactions that are exploitable to treat inflammation-mediated diseases. MAbs 6, 1201–1210 (2014).
Challis, R. C. et al. Systemic AAV vectors for widespread and targeted gene delivery in rodents. Nat. Protoc. 14, 379–414 (2019).
Lawlor, P. A., Bland, R. J., Mouravlev, A., Young, D. & During, M. J. Efficient gene delivery and selective transduction of glial cells in the mammalian brain by AAV serotypes isolated from nonhuman primates. Mol. Ther. 17, 1692–1702 (2009).
Acknowledgements
We thank J. J. Tabor (Rice University) for providing the pET28a bacterial expression plasmid and G. Bao (Rice University) for providing use of the ultracentrifuge. We thank V. Gradinaru (Caltech) and the Caltech CLOVER Center for providing the pUCmini-iCAP-PHP.eB and pHelper plasmids. This research was supported by the David and Lucile Packard Foundation 2021-73005 (J.O.S.), National Institute of Biomedical Imaging and Bioengineering Trailblazer Award R21EB033059 (J.O.S.), National Institute of General Medical Sciences DP2GM140923 (G.C.), National Institute on Drug Abuse R00DA043609 (G.C.) and National Institute of Neurological Disorders and Stroke F31NS125927 (J.A.W.).
Author information
Authors and Affiliations
Contributions
J.O.S. and S.L. conceived and planned the research. S.L. and J.O.S. designed the experiments and wrote the paper, with input from all other authors. S.L. performed and participated in all experiments described in the study. S.N. performed AAV production. S.N. and J.A.W. performed stereotaxic injection and retro-orbital blood collection. S.N. and J.J.K. performed the histological experiments. J.J.K. maintained PC-12 and conducted the in vitro Fos activation. S.N., J.A.W., J.J.K., Z.H. and B.X. analyzed histological images. S.N. and Z.H. assisted with IVIS imaging and drug administration. Z.H. constructed plasmids for the chemogenetic experiment. B.X. maintained and transfected astrocytes and performed protein purification. G.C. provided advice on the chemogenetic experiment and experimental guidance to J.A.W.
Corresponding author
Ethics declarations
Competing interests
J.O.S. and S.L. are co-inventors on a US patent application that incorporates discoveries described in this paper. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11 and Sequence 1.
Supplementary Data 2
Statistical source data for Supplementary Fig. 1.
Supplementary Data 3
Statistical source data for Supplementary Fig. 3.
Supplementary Data 4
Statistical source data for Supplementary Fig. 4.
Supplementary Data 5
Statistical source data for Supplementary Fig. 5.
Supplementary Data 6
Statistical source data for Supplementary Fig. 6.
Supplementary Data 7
Statistical source data for Supplementary Fig. 7.
Supplementary Data 8
Statistical source data for Supplementary Fig. 8.
Supplementary Data 9
Statistical source data for Supplementary Fig. 9.
Supplementary Data 10
Statistical source data for Supplementary Fig. 10.
Supplementary Data 11
Statistical source data for Supplementary Fig. 11.
Source data
Source Data Fig. 2
Statistical source data for Fig. 2.
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Fig. 4
Statistical source data for Fig. 4.
Source Data Fig. 5
Statistical source data for Fig. 5.
Source Data Fig. 6
Statistical source data for Fig. 6.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, S., Nouraein, S., Kwon, J.J. et al. Engineered serum markers for non-invasive monitoring of gene expression in the brain. Nat Biotechnol (2024). https://doi.org/10.1038/s41587-023-02087-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41587-023-02087-x