Abstract
Integrated in vitro models of human organogenesis are needed to elucidate the multi-systemic events underlying development and disease. Here we report the generation of human trunk-like structures that model the co-morphogenesis, patterning and differentiation of the human spine and spinal cord. We identified differentiation conditions for human pluripotent stem cells favoring the formation of an embryo-like extending antero-posterior (AP) axis. Single-cell and spatial transcriptomics show that somitic and spinal cord differentiation trajectories organize along this axis and can self-assemble into a neural tube surrounded by somites upon extracellular matrix addition. Morphogenesis is coupled with AP patterning mechanisms, which results, at later stages of organogenesis, in in vivo-like arrays of neural subtypes along a neural tube surrounded by spine and muscle progenitors contacted by neuronal projections. This integrated system of trunk development indicates that in vivo-like multi-tissue co-morphogenesis and topographic organization of terminal cell types can be achieved in human organoids, opening windows for the development of more complex models of organogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All scRNA-seq and tomo-seq data are available at the Gene Expression Omnibus: GSE215983 (SuperSeries composed of two SubSeries (single-cell and tomo-seq files))—the tomo-seq data (GSE215982) and the scRNA-seq data (GSE215981). The human embryo scRNA-seq data used in Extended Data Fig. 10 can be found at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE171892 and https://www.ebi.ac.uk/biostudies/arrayexpress/studies/E-MTAB-9388. For single-cell analysis, we used GRCh38-2020-A (from 10x Genomics) based on hg38. The human genome CRCh38 is available at https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000001405.26/. Additional data supporting the findings of this study are available in the article and the Supplementary Information. Further information for resources or technical information are available upon reasonable request from the corresponding author. Requests for transcriptomic analysis should be sent to a.vanoudenaarden@hubrecht.eu.
Code availability
The tomo-seq analysis script is available at https://github.com/anna-alemany.
References
Sahu, S. & Sharan, S. K. Translating embryogenesis to generate organoids: novel approaches to personalized medicine. iScience 23, 101485 (2020).
Shao, Y. & Fu, J. Engineering multiscale structural orders for high-fidelity embryoids and organoids. Cell Stem Cell 29, 722–743 (2022).
Moris, N., Alev, C., Pera, M. & Martinez Arias, A. Biomedical and societal impacts of in vitro embryo models of mammalian development. Stem Cell Rep. 16, 1021–1030 (2021).
Sozen, B., Conkar, D. & Veenvliet, J. V. Carnegie in 4D? Stem-cell-based models of human embryo development. Semin. Cell Dev. Biol. 131, 44–57 (2022).
Wymeersch, F. J., Wilson, V. & Tsakiridis, A. Understanding axial progenitor biology in vivo and in vitro. Development 148, dev180612 (2021).
Pourquié, O., Al Tanoury, Z. & Chal, J. The long road to making muscle in vitro. In Current Topics in Developmental Biology Vol. 129 (ed Brivanlou, A. H.) 123–142 (Academic Press, 2018).
Henrique, D., Abranches, E., Verrier, L. & Storey, K. G. Neuromesodermal progenitors and the making of the spinal cord. Development 142, 2864–2875 (2015).
Nedelec, S. & Martinez-Arias, A. In vitro models of spinal motor circuit’s development in mammals: achievements and challenges. Curr. Opin. Neurobiol. 66, 240–249 (2021).
Deschamps, J. & Duboule, D. Embryonic timing, axial stem cells, chromatin dynamics, and the Hox clock. Genes Dev. 31, 1406–1416 (2017).
Philippidou, P. & Dasen, J. S. Hox genes: choreographers in neural development, architects of circuit organization. Neuron 80, 12–34 (2013).
Abdel Fattah, A. R. et al. Actuation enhances patterning in human neural tube organoids. Nat. Commun. 12, 3192 (2021).
Budjan, C. et al. Paraxial mesoderm organoids model development of human somites. eLlife 11, e68925 (2022).
Duval, N. et al. BMP4 patterns Smad activity and generates stereotyped cell fate organization in spinal organoids. Development 146, dev175430 (2019).
Faustino Martins, J.-M. et al. Self-organizing 3D human trunk neuromuscular organoids. Cell Stem Cell 26, 172–186 (2020).
Gouti, M. et al. In vitro generation of neuromesodermal progenitors reveals distinct roles for Wnt signalling in the specification of spinal cord and paraxial mesoderm identity. PLoS Biol. 12, e1001937 (2014).
Zheng, Y. et al. Dorsal-ventral patterned neural cyst from human pluripotent stem cells in a neurogenic niche. Sci. Adv. 5, eaax5933 (2019).
Lee, J.-H. et al. Production of human spinal-cord organoids recapitulating neural-tube morphogenesis. Nat. Biomed. Eng. 6, 435–448 (2022).
Libby, A. R. G. et al. Axial elongation of caudalized human organoids mimics aspects of neural tube development. Development 148, dev198275 (2021).
Maury, Y. et al. Combinatorial analysis of developmental cues efficiently converts human pluripotent stem cells into multiple neuronal subtypes. Nat. Biotechnol. 33, 89–96 (2015).
Miao, Y. et al. Reconstruction and deconstruction of human somitogenesis in vitro. Nature 614, 500–508 (2023).
Mouilleau, V. et al. Dynamic extrinsic pacing of the HOX clock in human axial progenitors controls motor neuron subtype specification. Development 148, dev194514 (2021).
Ogura, T., Sakaguchi, H., Miyamoto, S. & Takahashi, J. Three-dimensional induction of dorsal, intermediate and ventral spinal cord tissues from human pluripotent stem cells. Development 145, dev162214 (2018).
Olmsted, Z. T. & Paluh, J. L. Co-development of central and peripheral neurons with trunk mesendoderm in human elongating multi-lineage organized gastruloids. Nat. Commun. 12, 3020 (2021).
Sanaki-Matsumiya, M. et al. Periodic formation of epithelial somites from human pluripotent stem cells. Nat. Commun. 13, 2325 (2022).
van den Brink, S. C. et al. Single-cell and spatial transcriptomics reveal somitogenesis in gastruloids. Nature 582, 405–409 (2020).
Veenvliet, J. V. et al. Mouse embryonic stem cells self-organize into trunk-like structures with neural tube and somites. Science 370, eaba4937 (2020).
Xu, P.-F. et al. Construction of a mammalian embryo model from stem cells organized by a morphogen signalling centre. Nat. Commun. 12, 3277 (2021).
Xue, W. et al. Generation of dorsoventral human spinal cord organoids via functionalizing composite scaffold for drug testing. iScience 26, 105898 (2022).
Yaman, Y. I. & Ramanathan, S. Controlling human organoid symmetry breaking reveals signaling gradients drive segmentation clock waves. Cell 186, 513–527 (2023).
Yamanaka, Y. et al. Reconstituting human somitogenesis in vitro. Nature 614, 509–520 (2023).
Diaz-Cuadros, M. et al. In vitro characterization of the human segmentation clock. Nature 580, 113–118 (2020).
Park, J., Hsiung, H.-A., Khven, I., La Manno, G. & Lutolf, M. P. Self-organizing in vitro mouse neural tube organoids mimic embryonic development. Development 149, dev201052 (2022).
Beccari, L. et al. Multi-axial self-organization properties of mouse embryonic stem cells into gastruloids. Nature 562, 272–276 (2018).
Moris, N. et al. An in vitro model of early anteroposterior organization during human development. Nature 582, 410–415 (2020).
Rito, T., Libby, A. R. G., Demuth, M. & Briscoe, J. Notochord and axial progenitor generation by timely BMP and NODAL inhibition during vertebrate trunk formation. Preprint at bioRxiv https://doi.org/10.1101/2023.02.27.530267 (2023).
Van Den Brink, S. C. et al. Symmetry breaking, germ layer specification and axial organisation in aggregates of mouse embryonic stem cells. Development 141, 4231–4242 (2014).
Olmsted, Z. T. & Paluh, J. L. Stem cell neurodevelopmental solutions for restorative treatments of the human trunk and spine. Front. Cell. Neurosci. 15, 667590 (2021).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Hemmati-Brivanlou, A., Kelly, O. G. & Melton, D. A. Follistatin, an antagonist of activin, is expressed in the Spemann organizer and displays direct neuralizing activity. Cell 77, 283–295 (1994).
Camus, A., Perea-Gomez, A., Moreau, A. & Collignon, J. Absence of Nodal signaling promotes precocious neural differentiation in the mouse embryo. Dev. Biol. 295, 743–755 (2006).
Sagner, A. & Briscoe, J. Establishing neuronal diversity in the spinal cord: a time and a place. Development 146, dev182154 (2019).
Tani, S., Chung, U.-I., Ohba, S. & Hojo, H. Understanding paraxial mesoderm development and sclerotome specification for skeletal repair. Exp. Mol. Med. 52, 1166–1177 (2020).
Goto, H., Kimmey, S. C., Row, R. H., Matus, D. Q. & Martin, B. L. FGF and canonical Wnt signaling cooperate to induce paraxial mesoderm from tailbud neuromesodermal progenitors through regulation of a two-step epithelial to mesenchymal transition. Development 144, 1412–1424 (2017).
Araya, C., Carmona-Fontaine, C. & Clarke, J. D. W. Extracellular matrix couples the convergence movements of mesoderm and neural plate during the early stages of neurulation. Dev. Dyn. 245, 580–589 (2016).
Buckley, C. E. & St Johnston, D. Apical–basal polarity and the control of epithelial form and function. Nat. Rev. Mol. Cell Biol. 23, 559–577 (2022).
Long, K. R. & Huttner, W. B. How the extracellular matrix shapes neural development. Open Biol. 9, 180216 (2019).
Mongera, A., Michaut, A., Guillot, C., Xiong, F. & Pourquié, O. Mechanics of anteroposterior axis formation in vertebrates. Annu. Rev. Cell Dev. Biol. 35, 259–283 (2019).
Boulet, A. M. & Capecchi, M. R. Signaling by FGF4 and FGF8 is required for axial elongation of the mouse embryo. Dev. Biol. 371, 235–245 (2012).
Cunningham, T. J., Kumar, S., Yamaguchi, T. P. & Duester, G. Wnt8a and Wnt3a cooperate in the axial stem cell niche to promote mammalian body axis extension. Dev. Dyn. 244, 797–807 (2015).
Ribes, V., Le Roux, I., Rhinn, M., Schuhbaur, B. & Dollé, P. Early mouse caudal development relies on crosstalk between retinoic acid, Shh and Fgf signalling pathways. Development 136, 665–676 (2009).
Anand, G. M. et al. Controlling organoid symmetry breaking uncovers an excitable system underlying human axial elongation. Cell 186, 497–512 (2023).
Padmanabhan, R. Retinoic acid-induced caudal regression syndrome in the mouse fetus. Reprod. Toxicol. 12, 139–151 (1998).
Tan, Y. & Cahan, P. SingleCellNet: a computational tool to classify single cell RNA-seq data across platforms and across species. Cell Syst. 9, 207–213 (2019).
Tyser, R. C. V. et al. Single-cell transcriptomic characterization of a gastrulating human embryo. Nature 600, 285–289 (2021).
Rayon, T., Maizels, R. J., Barrington, C. & Briscoe, J. Single-cell transcriptome profiling of the human developing spinal cord reveals a conserved genetic programme with human-specific features. Development 148, dev199711 (2021).
Mayeuf-Louchart, A. et al. Endothelial cell specification in the somite is compromised in Pax3-positive progenitors of Foxc1/2 conditional mutants, with loss of forelimb myogenesis. Development 143, 872–879 (2016).
Mayeuf-Louchart, A. et al. Notch regulation of myogenic versus endothelial fates of cells that migrate from the somite to the limb. Proc. Natl Acad. Sci. USA 111, 8844–8849 (2014).
Prummel, K. D., Nieuwenhuize, S. & Mosimann, C. The lateral plate mesoderm. Development 147, dev175059 (2020).
Junker, J. P. et al. Genome-wide RNA tomography in the zebrafish embryo. Cell 159, 662–675 (2014).
Rossi, G. et al. Capturing cardiogenesis in gastruloids. Cell Stem Cell 28, 230–240 (2021).
Rayon, T. et al. Species-specific pace of development is associated with differences in protein stability. Science 369, eaba7667 (2020).
Marklund, U. et al. Detailed expression analysis of regulatory genes in the early developing human neural tube. Stem Cells Dev. 23, 5–15 (2014).
Matsuda, M. et al. Recapitulating the human segmentation clock with pluripotent stem cells. Nature 580, 124–129 (2020).
Blauwkamp, T. A., Nigam, S., Ardehali, R., Weissman, I. L. & Nusse, R. Endogenous Wnt signalling in human embryonic stem cells generates an equilibrium of distinct lineage-specified progenitors. Nat. Commun. 3, 1070 (2012).
Kyprianou, C. et al. Basement membrane remodelling regulates mouse embryogenesis. Nature 582, 253–258 (2020).
Rifes, P. et al. Redefining the role of ectoderm in somitogenesis: a player in the formation of the fibronectin matrix of presomitic mesoderm. Development 134, 3155–3165 (2007).
Radice, G. L. et al. Developmental defects in mouse embryos lacking N-cadherin. Dev. Biol. 181, 64–78 (1997).
Nikolopoulou, E., Galea, G. L., Rolo, A., Greene, N. D. E. & Copp, A. J. Neural tube closure: cellular, molecular and biomechanical mechanisms. Development 144, 552–566 (2017).
Ensini, M., Tsuchida, T. N., Belting, H. G. & Jessell, T. M. The control of rostrocaudal pattern in the developing spinal cord: specification of motor neuron subtype identity is initiated by signals from paraxial mesoderm. Development 125, 969–982 (1998).
Wolujewicz, P., Steele, J. W., Kaltschmidt, J. A., Finnell, R. H. & Ross, M. E. Unraveling the complex genetics of neural tube defects: From biological models to human genomics and back. Genesis 59, e23459 (2021).
Andersen, J. et al. Generation of functional human 3D cortico-motor assembloids. Cell 183, 1913–1929 (2020).
Barbeau, S., Tahraoui-Bories, J., Legay, C. & Martinat, C. Building neuromuscular junctions in vitro. Development 147, dev193920 (2020).
Cederquist, G. Y. et al. Specification of positional identity in forebrain organoids. Nat. Biotechnol. 37, 436–444 (2019).
Renier, N. et al. Mapping of brain activity by automated volume analysis of immediate early genes. Cell 165, 1789–1802 (2016).
Acknowledgements
We thank M. Russeau for help and assistance on pluripotent stem cell culture; P. Lacour, F. Luciani, T. Rashed and H. Malek for their help during the project; J. Dasen for providing guinea pig anti-HOXC9 antibody; C. Metin for sharing ARL13B antibody; and L. Goutebroze for providing the Phalloidin probes. We thank H. Wichterle and the Ribes laboratory, in particular V. Ribes and P. Gilardi, as well as F. Causeret for helpful discussions along the project and critical reading of the manuscript. We thank J.-A. Girault and I. Caillé for their comments on the manuscripts. Imaging experiments and iPSC cultures were carried out, respectively, at the Imaging Platform and the Cell and Tissue Engineering Facility of the Institut du Fer à Moulin. This project was supported by two grants from the ATIP/Avenir program (S.N. and J.F.), Laboratoire d’Excellence (LabEx) Biopsy (ANR-10-LABX-73, S.N.), ANR SYMASYM (ANR-18-CE16-0021-03, S.N. and X.M.) as well as the NWO Gravitation Project: BRAINSCAPES: A Roadmap from Neurogenetics to Neurobiology (NWO: 024.004.012, B.v.S. and A.v.O.) and a ZonMW PSIDER grant (GREAT, 40-46800-98-015, A.L. and A.v.O.).
Author information
Authors and Affiliations
Contributions
S.N. and S.G. conceived the project. S.N., S.G., X.M., A.v.O. and R.R. designed experiments. S.G., S.N., R.R. and C.M. performed most wet lab experiments. K.B., R.R., S.N. and X.M. performed time-lapse imaging. J.F. performed clearing and 3D imaging. A.v.O. and A.L. performed scRNA-seq and tomo-seq experiments. A.v.O. and B.v.S. performed scRNA-seq and tomo-seq analysis. Project administration: S.N., X.M. and A.v.O. Supervision: S.N., A.v.O. and X.M. Writing—original draft: S.G. and S.N. Writing—review and editing: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Alexandre Mayran and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Morphological evolution of hiPSC-derived aggregates over time.
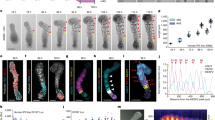
a, Representative bright field images of day 7 structures derived from WTS2 or WTC11 cell lines and from independent experiments, n = 13 for WTS2 and n = 7 for WTC11. b, Aspect ratio (Length (L)/Width (W)) of WTS2-derived structures over time. At day 7, the mean aspect ratio is 1.7 ± 0.76. c, Distribution of (Lmax-Wmax) of WTS2-derived structures over time: bin center = 50 µm. d, Box-plot of WTC11-derived structures width and length over time. At day 7, the mean length is 516 ± 285 µm. e, WTC11-derived structures aspect ratio over time. At day 7, the mean aspect ratio is 1.7 ± 0.75. f, Distribution of (Lmax-Wmax) of WTC11-derived structures over time: bin center = 50 µm. For box plots, the central line indicates the median; box limits the 1st and 3rd quartiles and whiskers the range. Each dot corresponds to a single structure and different colors indicate independent experiments. For WTS2, n = 11 for day 3; n = 12 for day 4; n = 13 for day 7. For WTC11, n = 2 for day 3; n = 3 for day 4; n = 3 for day 7. Scale bar, 200 µm (a).
Extended Data Fig. 2 Cell fate specification and topographical organization.
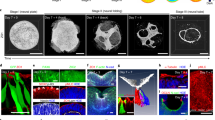
a, Schematic of the caudal region of the mammalian embryo. Axial progenitors generate both spinal neural and somitic lineages. n.t: neural tube. pPSM, aPSM: posterior and anterior PSM. b-b’, Maximum-intensity projection of whole-mount immunostaining on day 4 for markers of axial (SOX2/TBXT/CDX2), neural (SOX2) and mesodermal (TBX6) progenitors. WTS2 (b) and WTC11 (b’) derived structures. c, Percentage of structures with regionalized SOX2/TBXT/CDX2 progenitors in the entire population (day 4, n = 7, 159 structures) or among the elongated structures (at day 6, n = 3, 74 structures; day 7, n = 4, 48 structures and day 8, n = 3, 35 structures). Error bars represents the standard deviation. d-g, Immunostaining of sections at day 7 for mitotic cells (P-H3), F-Actin (Phalloidin) and neural progenitors (SOX1/3) markers (d, n = 2), or for the ventral neural progenitor marker (NKX6.1) (e, n = 4), or for the somitic markers FOXC2, SOX9, PAX3 (f, n = 2) and MEOX1 (g, n = 5). h, Low magnification view of sections of structures at day 7, immunostained for SOX9, PAX3 and SOX1/3, n = 4 i,j, 3D maximum-intensity projections of whole mount immunostaining at day 7 for the neuronal (SOX1/3, PAX3 in (i) or SOX1/3, OLIG2 in (j)) and somitic (FOXC2, SOX9, PAX3) markers. Optical plane in j. Scale bars, 50 µm (b, d-f), 100 µm (g, i, j), 500 µm (h).
Extended Data Fig. 3 Analysis of structures derived from other hiPSC lines and factors impacting neuro/mesodermal differentiation.
a, Representative bright field images of day 7 structures generated from SOX2-GFP, WTC11 and KUTE4 cell lines (n = 3, n = 3, n = 2). b-d, Immunostaining of sections of day 7 structures for neural progenitors (SOX1/3) and somitic (PAX3) markers, n = 3 for SOX2-GFP and WTC11, n = 2 for KUTE4. e-h, Representative bright field images (e, f; n = 3 and n = 4) and sections immunostained for neural progenitors (SOX2) and somitic (FOXC2) markers (g, h; n = 3 and n = 4) of day 7 structures generated by aggregating different cell numbers at day 0 (120, 160 and 300 cells). i, Representative bright field images (left panel) and sections immunostained for SOX2 and FOXC2 markers (right panel) of day 7 structures generated by aggregating different cell numbers at day 0 (300 or 600 cells) in presence of CHIR or with the addition of LDN and SB (n = 3). j, Representative bright field images (left panel) and sections immunostained for SOX2 and FOXC2 markers (right panel) of day 7 structures in presence of CHIR or with the addition of bFGF (20 ng/ml) for the first 2 days of differentiation (n = 3). k, Immunostaining of plated day 3 aggregates for neural progenitor (SOX2) and PSM (TBX6) markers. Aggregates obtained with different initial cell numbers were cultured using different B27 supplement (percentage of aggregates with TBX6 staining at day 3: B27 control, 50 ± 13% (120 cells) versus 83 ± 8% (300 cells), n = 2; B27 batch #2450484, 14 ± 9% (120 cells) versus 32 ± 21% (300 cells), n = 2; B27 batch #2450486, 2% (120 cells) versus 8% (300 cells), n = 1). l, Representative bright field images of day 7 structures obtained with the different initial cell numbers and different B27 supplements (n as in k). Scale bars, 200 µm (a, e, f, g, h, i, j, l), 100 µm (b, c, d), 50 µm (k).
Extended Data Fig. 4 Morphometric analysis of day 7 matrigel embedded structures.
a, Percentage of elongation for structures derived from 3 different cells lines (WTS2, n = 9, 892 structures total; WTC11, n = 3, 161 structures total; and SOX2-GFP, n = 2, 76 structures total). b, Box-plot of aspect ratio (length/width) of elongated structures at day 7 (WTS2, n = 8, 100 structures total; WTC11, n = 4, 57 structures total; SOX2-GFP, n = 3, 34 structures total). In the box plots, the central line indicates the median; box limits the 1st and 3rd quartiles and whiskers the range. Each dot corresponds to a single aggregate and different colors indicate independent differentiations. c-e, Gallery of bright field images of day 7 structures derived from 3 different hiPSC lines. Numbers indicate series from independent experiments. Scale bars, 200 µm (c, d, e).
Extended Data Fig. 5 Live imaging of matrigel embedded structures.
a, Time-lapse imaging showing dynamic elongation of matrigel-embedded SOX2-GFP derived structures between day 4 and 5.5. Red arrows pinpoint SOX2 negative cells. b, Time-lapse of matrigel-embedded WTS2 derived structures between day 4 and 7. Post-imaging whole mount immunostaining at day 7 for axial progenitors (SOX2, CDX2) and somitic (FOXC2) markers for the corresponding structure. #3 and #4, examples of non-elongating aggregates. Scale bars, 100 µm (a, b).
Extended Data Fig. 6 WNT, FGF and RA pathways control neural tube formation.
a, Experimental design. b, Representative images of SOX2-GFP whole mount structures in the different culture conditions at day 7, n = 3. c-d, Quantification of the aspect ratio (length/width) in the different conditions for day 7 organoids derived from SOX2-GFP (c) (n = 3, 2 to 6 wells/n/condition) and WTS2 (d) (n = 1, 2 wells/n/condition) hiPSC lines. ERKi and FGFRi = FGF pathways inhibitors. IWP2 = Wnt pathway inhibitor. In the box plots, the central line indicates the median; box limits the 1st and 3rd quartiles and whiskers the range. Each dot represents a single organoid. Statistical test in c: Kruskal-Wallis followed by Dunn’s multiple comparison test. *** P < 0.001. Scale bar, 500 µm.
Extended Data Fig. 7 Organization of the neural and somitic compartments in matrigel-embedded structures.
a, Confocal virtual slice and 3D visualization of whole mount immunostaining for F-Actin (Phalloidin) and mitotic cells (P-H3) indicating the apical side of the neuroepithelium. b,c, Representative images of longitudinal (b, n = 3) and coronal (c, n = 2) sections of day 7 structures immunostained for neural progenitors (SOX1/3, PAX6) and mesodermal (PAX3, SOX9, FOXC2) markers. d, Immunostaining of a section consecutive to the one in Fig. 3e for the neural progenitor and PSM/somitic marker PAX3. e-g, Sections of day 7 structures derived from WTC11 (e, n = 2) or SOX2-GFP (f, g, n = 4 and n = 2) cell lines immunostained for markers of neural progenitors (SOX1/3, PAX3, OLIG2, NKX6.1) and somitic (SOX9, PAX3, FOXC2). Scale bars, 50 µm (a, c), 100 µm (b, d-g).
Extended Data Fig. 8 ScRNAseq analysis of day 7 matrigel-embedded structures.
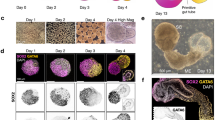
a-b, ScRNA-seq clusters generated from day 7 structures compared with annotated scRNA-seq datasets from primary human gastrulating embryo53, and human embryonic spinal cord tissue54. Statistical significance was calculated using a binomial test, where probabilities were determined by randomizing the primary tissue cluster marker gene list, based on the full list of primary tissue marker genes (n = 10.000). c, UMAPs colored by the expression of specific marker genes of each of the detected cell types and the expression of HOX genes differentially expressed along the spinal cord and somites.
Extended Data Fig. 9 Cardiac progenitor and HOX mRNA patterns in trunk organoids.
a, Representative images of day 7 sections immunostained for markers of cardiac (GATA6) and intermediate (LHX1) mesoderm progenitors and the caudal progenitor marker (CDX2), n = 2. GATA6/LHX1 clusters are in 38,1% of the structures (n = 3, 222 non-Matrigel-embedded structures). b, Expression pattern of Hoxa/c and Cdx2 genes in mouse embryo. c-e, Tomo-seq data analysis showing the expression patterns of HOXA and HOXC clusters along structure #2 (c) and for HOXB and HOXD clusters along the structure #1 (d) and #2 (e). f-g, Related to Fig. 3j, FISH for HOXC6 and HOXC9 mRNAs on additional neural tubes at day 7. f, Example of neural tube spanning from CDX2-high region (posterior) to CDX2-low/negative region (anterior) with segregated HOXC6 and HOXC9 mRNA expression domains. g, example of CDX2 highly expressed throughout the neural tube, with co-expressed HOXC6 and HOXC9 mRNA domains (5/10 organoids with co-expression, n = 3) (g). Scale bars, 100 µm (a), 50 µm (f-g).
Extended Data Fig. 10 Characterization of trunk organoids at later stages.
a, Sections of day 14 non-embedded organoids immunostained for neuronal (NEFL), motor neurons (ISL1) and neuronal progenitors (SOX1/3) markers, n = 3. b,c, Immunostaining for NEFL, ISL1 and the apical marker F-Actin (Phalloidin) at day 14 in the control condition (b, n = 3), and at day 10 after DAPT addition on day 8 (c, n = 2). d, Related to Fig. 4h, other example of collinear expression patterns of HOXC6, HOXC8 and HOXC9 along the neural tube (longitudinal cryostat sections at day 14). Scale bars, 50 µm (a-c), 200 µm (d).
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Supplementary Table
List of genes characterizing the scRNA-seq clusters. Differentially expressed genes for each cluster were determined using a Wilcoxon rank-sum test on genes showing at least 0.25 log2(fold change). List of genes characterizing the tomo-seq clusters. Comparison of protocols generating trunk-like structures, human neuromesodermal organoids or human somite-only elongating models. Recombinant proteins, small molecules, reagents and consumables used in this study. Antibodies and Phalloidin probes.
Supplementary Video 1
Live imaging (×10 magnification) from day 4 to day 6 of a SOX2-GFP structure.
Supplementary Video 2
Confocal z-stack (×63 magnification) through a day 7 SOX2-GFP structure shows the formation of a neural tube.
Supplementary Video 3
Live imaging (×63 magnification) of a day 7 SOX2-GFP organoid.
Supplementary Video 4
Live imaging (×10 magnification) from day 4 to day 7 of a WTS2 structure.
Supplementary Video 5
3D reconstructions of whole-mount light-sheet imaging of structures at day 17 showing motor neurons (ISL1) and neurons (β-III-tubulin).
Supplementary Video 6
3D reconstructions of whole-mount light-sheet imaging of structures at day 17 showing motor neurons (ISL1) and neurons (β-III-tubulin).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gribaudo, S., Robert, R., van Sambeek, B. et al. Self-organizing models of human trunk organogenesis recapitulate spinal cord and spine co-morphogenesis. Nat Biotechnol (2023). https://doi.org/10.1038/s41587-023-01956-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41587-023-01956-9
This article is cited by
-
Cellular and molecular control of vertebrate somitogenesis
Nature Reviews Molecular Cell Biology (2024)
-
Modeling human trunk development
Nature Biotechnology (2023)