Abstract
Crop genetic improvement requires balancing complex tradeoffs caused by gene pleiotropy and linkage drags, as exemplified by IPA1 (Ideal Plant Architecture 1), a typical pleiotropic gene in rice that increases grains per panicle but reduces tillers. In this study, we identified a 54-base pair cis-regulatory region in IPA1 via a tiling-deletion-based CRISPR–Cas9 screen that, when deleted, resolves the tradeoff between grains per panicle and tiller number, leading to substantially enhanced grain yield per plant. Mechanistic studies revealed that the deleted fragment is a target site for the transcription factor An-1 to repress IPA1 expression in panicles and roots. Targeting gene regulatory regions should help dissect tradeoff effects and provide a rich source of targets for breeding complementary beneficial traits.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data supporting the findings of this study are available in the article, supplementary information and source data files. Source data are provided with this paper.
References
Takatsuji, H. Regulating tradeoffs to improve rice production. Front. Plant Sci. 8, 171 (2017).
Nelson, R., Wiesner-Hanks, T., Wisser, R. & Balint-Kurti, P. Navigating complexity to breed disease-resistant crops. Nat. Rev. Genet. 19, 21–33 (2018).
Wang, J., Long, X., Chern, M. & Chen, X. Understanding the molecular mechanisms of trade-offs between plant growth and immunity. Sci. China Life Sci. 64, 234–241 (2021).
Wang, L. et al. Roles of FERONIA-like receptor genes in regulating grain size and quality in rice. Sci. China Life Sci. 64, 294–310 (2021).
Li, Y. et al. Chalk5 encodes a vacuolar H+-translocating pyrophosphatase influencing grain chalkiness in rice. Nat. Genet. 46, 398–404 (2014).
Zhang, S. et al. Detection of major loci associated with the variation of 18 important agronomic traits between Solanum pimpinellifolium and cultivated tomatoes. Plant J. 95, 312–323 (2018).
Wu, W. et al. A single-nucleotide polymorphism causes smaller grain size and loss of seed shattering during African rice domestication. Nat. Plants 3, 17064 (2017).
Uauy, C., Distelfeld, A., Fahima, T., Blechl, A. & Dubcovsky, J. A NAC gene regulating senescence improves grain protein, zinc, and iron content in wheat. Science 314, 1298–1301 (2006).
Guttieri, M. J., Stein, R. J. & Waters, B. M. Nutrient partitioning and grain yield of TaNAM-RNAi wheat under abiotic stress. Plant Soil 371, 573–591 (2013).
Hedden, P. The genes of the Green Revolution. Trends Genet. 19, 5–9 (2003).
Li, S. et al. Modulating plant growth-metabolism coordination for sustainable agriculture. Nature 560, 595–600 (2018).
Sasaki, A. et al. Green revolution: a mutant gibberellin-synthesis gene in rice. Nature 416, 701–702 (2002).
Visscher, P. M. & Yang, J. A plethora of pleiotropy across complex traits. Nat. Genet. 48, 707–708 (2016).
Wang, Y. & Li, J. The plant architecture of rice (Oryza sativa). Plant Mol. Biol. 59, 75–84 (2005).
Yano, K. et al. GWAS with principal component analysis identifies a gene comprehensively controlling rice architecture. Proc. Natl Acad. Sci. USA 116, 21262–21267 (2019).
Wang, B., Smith, S. M. & Li, J. Genetic regulation of shoot architecture. Annu. Rev. Plant Biol. 69, 437–468 (2018).
Jiao, Y. et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 42, 541–544 (2010).
Miura, K. et al. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 42, 545–549 (2010).
Wang, J. et al. A single transcription factor promotes both yield and immunity in rice. Science 361, 1026–1028 (2018).
Lu, Z. et al. Genome-wide binding analysis of the transcription activator ideal plant architecture1 reveals a complex network regulating rice plant architecture. Plant Cell 25, 3743–3759 (2013).
Zhang, L. et al. A natural tandem array alleviates epigenetic repression of IPA1 and leads to superior yielding rice. Nat. Commun. 8, 14789 (2017).
Huang, X. et al. Genomic architecture of heterosis for yield traits in rice. Nature 537, 629–633 (2016).
Xue, J. et al. The genetic arms race between plant and Xanthomonas: lessons learned from TALE biology. Sci. China Life Sci. 64, 51–65 (2021).
Song, X. et al. IPA1 functions as a downstream transcription factor repressed by D53 in strigolactone signaling in rice. Cell Res. 27, 1128–1141 (2017).
Wang, J. et al. Tissue-specific ubiquitination by IPA1 INTERACTING PROTEIN1 modulates IPA1 protein levels to regulate plant architecture in rice. Plant Cell 29, 697–707 (2017).
Li, X., Xie, Y., Zhu, Q. & Liu, Y. G. Targeted genome editing in genes and cis-regulatory regions improves qualitative and quantitative traits in crops. Mol. Plant 10, 1368–1370 (2017).
Rodríguez-Leal, D., Lemmon, Z. H., Man, J., Bartlett, M. E. & Lippman, Z. B. Engineering quantitative trait variation for crop improvement by genome editing. Cell 171, 470–480 (2017).
Hendelman, A. et al. Conserved pleiotropy of an ancient plant homeobox gene uncovered by cis-regulatory dissection. Cell 184, 1724–1739 (2021).
Wittkopp, P. J. & Kalay, G. Cis-regulatory elements: molecular mechanisms and evolutionary processes underlying divergence. Nat. Rev. Genet. 13, 59–69 (2011).
Liu, L. et al. Enhancing grain-yield-related traits by CRISPR–Cas9 promoter editing of maize CLE genes. Nat. Plants 7, 287–294 (2021).
Chow, C. N. et al. PlantPAN3.0: a new and updated resource for reconstructing transcriptional regulatory networks from ChIP-seq experiments in plants. Nucleic Acids Res. 47, D1155–D1163 (2019).
Luo, J. et al. An-1 encodes a basic helix-loop-helix protein that regulates awn development, grain size, and grain number in rice. Plant Cell 25, 3360–3376 (2013).
Jin, J. et al. GAD1 encodes a secreted peptide that regulates grain number, grain length, and awn development in rice domestication. Plant Cell 28, 2453–2463 (2016).
Meyer, R. S. & Purugganan, M. D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Lemmon, Z. H., Bukowski, R., Sun, Q. & Doebley, J. F. The role of cis regulatory evolution in maize domestication. PLoS Genet. 10, e1004745 (2014).
Swinnen, G., Goossens, A. & Pauwels, L. Lessons from domestication: targeting cis-regulatory elements for crop improvement. Trends Plant Sci. 21, 506–515 (2016).
Chen, R. et al. Rice functional genomics: decades’ efforts and roads ahead. Sci. China Life Sci. 65, 33–92 (2022).
Schwarzer, W. & Spitz, F. The architecture of gene expression: integrating dispersed cis-regulatory modules into coherent regulatory domains. Curr. Opin. Genet. Dev. 27, 74–82 (2014).
Priest, H. D., Filichkin, S. A. & Mockler, T. C. Cis-regulatory elements in plant cell signaling. Curr. Opin. Plant Biol. 12, 643–649 (2009).
Hernandez-Garcia, C. M. & Finer, J. J. Identification and validation of promoters and cis-acting regulatory elements. Plant Sci. 217-218, 109–119 (2014).
Naito, Y., Hino, K., Bono, H. & Ui-Tei, K. CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 31, 1120–1123 (2015).
Yu, H. et al. A route to de novo domestication of wild allotetraploid rice. Cell 184, 1156–1170 (2021).
Meng, X. et al. Construction of a genome-wide mutant library in rice using CRISPR/Cas9. Mol. Plant 10, 1238–1241 (2017).
Saleh, A., Alvarez-Venegas, R. & Avramova, Z. An efficient chromatin immunoprecipitation (ChIP) protocol for studying histone modifications in Arabidopsis plants. Nat. Protoc. 3, 1018–1025 (2008).
Hellens, R. P. et al. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 1, 13 (2005).
Zhang, H. et al. Genome editing of upstream open reading frames enables translational control in plants. Nat. Biotechnol. 36, 894–898 (2018).
Acknowledgements
We thank C. Sun (China Agricultural University) for providing the NIL-An-1 and NIL-an-1 seeds. This work was supported by the National Natural Science Foundation of China (31788103 to J.L. and H.Y. and 32122064 to H.Y.), the Chinese Academy of Sciences (XDA24030504 to J.L.) and the Hainan Excellent Talent Team (to J.L.).
Author information
Authors and Affiliations
Contributions
X.S. performed most of the experiments. X.S., X.M. and H.G. designed the CRISPR target and constructed the plasmid library. H.G. and X.M. transformed rice. X.S., X.M., H.G. and Q.C. characterized the genotypes and phenotypes of the edited lines. X.M., Y.J., M.C. and G.L. contributed to the rice materials. X.S., H.G., B.W., Y.W. and H.Y. analyzed the data. X.S., J.L. and H.Y. conceived and designed experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Xiangdong Fu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Design of gRNAs in the tiling-deletion screen.
Red bars, CRISPR-Cas9 target sites in each vector. Blue bars, regions between CRISPR-Cas9 target sites on same vector. TSS, transcription start site. TTS, transcription termination site. The positions in promoter and 5’ UTR were relative to the transcription start site. The positions in 3’ UTR and downstream region were relative to the transcription termination site.
Extended Data Fig. 2 Morphological phenotypes of edited lines.
Upper panel, plants at the mature stage; lower panel, their corresponding panicles. Bars = 10 cm.
Extended Data Fig. 3 Correlations between the panicle weight and tiller number in two years.
a,b, Scatter plot of panicle weight (a) and tiller number (b) of 21 edited lines in 2019 and 2020. R2 values were calculated using linear regression.
Extended Data Fig. 4 Stem diameter of ZH11, IPA1-Pro10, ipa1-5D, and ipa1-12 plants.
a, Cross-sections of internodes of ZH11, IPA1-Pro10, ipa1-5D, and ipa1-12. Bars = 1 mm. b-d, Statistical analysis of internode diameters in a. Values indicate means ± s.d. (n = 10 plants). Exact P values are shown; Tukey’s HSD test (b-d).
Extended Data Fig. 5 IPA1 expression levels in different tissues of ZH11 and IPA1-Pro10.
a, IPA1 expression levels in panicles at different stages of ZH11 and IPA1-Pro10. b,c, IPA1 expression levels in tiller bud (b) and root (c) of ZH11 and IPA1-Pro10. d, IPA1 expression profile in different tissues. Shoot base, young leaf, and root were sampled from 2-week-old seedings. Old leaf and stem were sampled from plants after heading stage. IPA1 expression levels in 0.5-cm panicle (a), tiller bud (b), root (c), and shoot base (d) of ZH11 were set to one. Values indicate means ± s.d. (n = 3 biological replicates in a-c and n = 3 technical replicates in d). Exact P values are shown; two-sided Student’s t-test.
Extended Data Fig. 6 Thousand-grain weight and days to heading of ZH11 and IPA1-Pro10 in paddy field.
a, Statistical analysis of thousand-grain weight. b, Statistical analysis of days to heading. Values indicate means ± s.d. (n = 3 biological replicates). Exact P values are shown; two-sided Student’s t-test.
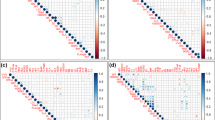
Extended Data Fig. 7 Predicted An-1 binding sites in the promoters of SPL14 orthologs.
The An-1 binding sites were predicted in the promoters of IPA1 homologous genes, including Aegilops tauschii SPL17 (LOC109731696), Brachypodium distachyon SPL14 (LOC100836973), Sorghum bicolor SPL14 (LOC8064025), Zea mays SBP8 (GRMZM2G160917) and SPL15 (GRMZM2G460544), Setaria italica SPL14 (LOC101773844), Arabidopsis thaliana SPL9 (AT2G42200) and SPL15 (AT3G57920), and Nicotiana tabacum SPL9 (LOC107809759).
Extended Data Fig. 8 An-1 expression levels in different tissues of ZH11 and IPA1-Pro10.
a, An-1 expression levels in panicles at different stages of ZH11 and IPA1-Pro10. b,c, An-1 expression levels in tiller bud (b) and root (c) of ZH11 and IPA1-Pro10. d, An-1 expression profile in different tissues. Shoot base, young leaf, and root were sampled from 2-week-old seedings. Old leaf and stem were sampled from plants after heading stage. An-1 expression levels in 0.5-cm panicle (a), tiller bud (b), root (c), and shoot base (d) of ZH11 were set to one. Values indicate means ± s.d. (n = 3 biological replicates in a-c and n = 3 technical replicates in d). Exact P values are shown; two-sided Student’s t-test.
Extended Data Fig. 9 IPA1 expression levels in different tissues of NIL-An-1 and NIL-an-1.
a, IPA1 expression levels in panicles at different stages of NIL-An-1 and NIL-an-1. b, An-1 expression in the tiller buds of NIL-An-1 and NIL-an-1. c, IPA1 expression profile in different tissues of NIL-An-1 and NIL-an-1. Shoot base, young leaf, and root were sampled from 2-week-old seedings. Old leaf and stem were sampled from plants after heading stage. Gene expression levels in 0.5-cm panicle (a), tiller bud (b), and shoot base (c) of NIL-An-1 were set to one. Values indicate means ± s.d. (n = 3 biological replicates in a and b, n = 3 technical replicates in c). Exact P values are shown; two-sided Student’s t-test.
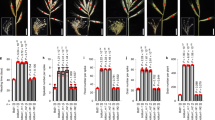
Extended Data Fig. 10 Panicle phenotypes of IPA1-Pro11, IPA1-Pro12 and IPA1-Pro13.
a-h, Panicle morphology (a), panicle weight (b), grain number per primary branch (c), secondary branch per primary branch (d), primary branch number (e), secondary branch number (f), thousand-grain weight (g), and grain setting rate (h) of plants with wild-type (–/–), heterozygous (+/–), and homozygous (+/+) ipa1-pro11 alleles. i-p, Panicle morphology (i), panicle weight (j), grain number per primary branch (k), secondary branch per primary branch (l), primary branch number (m), secondary branch number (n), thousand-grain weight (o), and grain setting rate (p) of plants with wild-type, heterozygous, and homozygous ipa1-pro12 alleles. q-x, Panicle morphology (q), panicle weight (r), grain number per primary branch (s), secondary branch per primary branch (t), primary branch number (u), secondary branch number (v), thousand-grain weight (w), and grain setting rate (x) of plants with wild-type, heterozygous and homozygous ipa1-pro13 alleles. Bars = 10 cm. Values indicate means ± s.d. (n = 15 plants). Exact P values are shown; two-sided Student’s t-test.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 and Supplementary Tables 1 and 6
Supplementary Tables 2–5
Supplementary Table 2: 39 vectors used in the tiling-deletion screen. Supplementary Table 3: Genotyping result of 792 T0 plants. Supplementary Table 4: Agronomic traits of 21 edited lines. Supplementary Table 5: Predicted transcription factor binding sites in the promoter of IPA1.
Supplementary Data 1
Statistical Source Data for Supplementary Figs. 5b,c and 6b,c
Supplementary Data 2
Unprocessed gel for Supplementary Fig. 5a
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 4
Unprocessed EMSA blots
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 8
Statistical Source Data
Source Data Extended Data Fig. 9
Statistical Source Data
Source Data Extended Data Fig. 10
Statistical Source Data
Rights and permissions
About this article
Cite this article
Song, X., Meng, X., Guo, H. et al. Targeting a gene regulatory element enhances rice grain yield by decoupling panicle number and size. Nat Biotechnol 40, 1403–1411 (2022). https://doi.org/10.1038/s41587-022-01281-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-022-01281-7
This article is cited by
-
Precise fine-turning of GhTFL1 by base editing tools defines ideal cotton plant architecture
Genome Biology (2024)
-
Application of genome editing techniques to regulate gene expression in crops
BMC Plant Biology (2024)
-
Targeted genome-modification tools and their advanced applications in crop breeding
Nature Reviews Genetics (2024)
-
The role and pathway of VQ family in plant growth, immunity, and stress response
Planta (2024)
-
Developing a CRISPR/FrCas9 system for core promoter editing in rice
aBIOTECH (2024)