Abstract
Comprehensive phenotypic characterization of the many mutations found in cancer tissues is one of the biggest challenges in cancer genomics. In this study, we evaluated the functional effects of 29,060 cancer-related transition mutations that result in protein variants on the survival and proliferation of non-tumorigenic lung cells using cytosine and adenine base editors and single guide RNA (sgRNA) libraries. By monitoring base editing efficiencies and outcomes using surrogate target sequences paired with sgRNA-encoding sequences on the lentiviral delivery construct, we identified sgRNAs that induced a single primary protein variant per sgRNA, enabling linking those mutations to the cellular phenotypes caused by base editing. The functions of the vast majority of the protein variants (28,458 variants, 98%) were classified as neutral or likely neutral; only 18 (0.06%) and 157 (0.5%) variants caused outgrowing and likely outgrowing phenotypes, respectively. We expect that our approach can be extended to more variants of unknown significance and other tumor types.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
We have submitted the deep sequencing data from this study to the National Center of Biotechnology Information’s Sequence Read Archive under accession number PRJNA667758. We have provided the datasets used in this study as Supplementary Tables 2–4 and deepcrispr.info/BEvariants.
Code availability
The custom Python scripts used for the generation of the MAGeCK input file using UMIs are available on GitHub (https://github.com/oreolic/CancerLibrary).
References
McLendon, R. et al. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Hudson, T. J. et al. International network of cancer genome projects. Nature 464, 993–998 (2010).
Campbell, P. J. et al. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385 (2018).
Sondka, Z. et al. The COSMIC Cancer Gene Census: describing genetic dysfunction across all human cancers. Nat. Rev. Cancer 18, 696–705 (2018).
Rheinbay, E. et al. Analyses of non-coding somatic drivers in 2,658 cancer whole genomes. Nature 578, 102–111 (2020).
Stratton, M. R., Campbell, P. J. & Futreal, P. A. The cancer genome. Nature 458, 719 (2009).
Giacomelli, A. O. et al. Mutational processes shape the landscape of TP53 mutations in human cancer. Nat. Genet. 50, 1381–1387 (2018).
Kotler, E. et al. A systematic p53 mutation library links differential functional impact to cancer mutation pattern and evolutionary conservation. Mol. Cell 71, 178–190 (2018).
Majithia, A. R. et al. Prospective functional classification of all possible missense variants in PPARG. Nat. Genet. 48, 1570–1575 (2016).
Brenan, L. et al. Phenotypic characterization of a comprehensive set of MAPK1/ERK2 missense mutants. Cell Rep. 17, 1171–1183 (2016).
Ahler, E. et al. A combined approach reveals a regulatory mechanism coupling Src’s kinase activity, localization, and phosphotransferase-independent functions. Mol. Cell 74, 393–408 (2019).
Matreyek, K. A. et al. Multiplex assessment of protein variant abundance by massively parallel sequencing. Nat. Genet. 50, 874–882 (2018).
Starita, L. M. et al. A multiplex homology-directed DNA repair assay reveals the impact of more than 1,000 BRCA1 missense substitution variants on protein function. Am. J. Hum. Genet. 103, 498–508 (2018).
Chiasson, M. A. et al. Multiplexed measurement of variant abundance and activity reveals VKOR topology, active site and human variant impact. eLife 9, e58026 (2020).
Kim, H. & Kim, J. S. A guide to genome engineering with programmable nucleases. Nat. Rev. Genet. 15, 321–334 (2014).
Findlay, G. M. et al. Accurate classification of BRCA1 variants with saturation genome editing. Nature 562, 217–222 (2018).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, aaf8729 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Kim, H. S. et al. Systematic identification of molecular subtype-selective vulnerabilities in non-small-cell lung cancer. Cell 155, 552–566 (2013).
Bamford, S. et al. The COSMIC (Catalogue of Somatic Mutations in Cancer) database and website. Br. J. Cancer 91, 355–358 (2004).
Hanna, R. E. et al. Massively parallel assessment of human variants with base editor screens. Cell 184, 1064–1080 (2021).
Kuscu, C. et al. CRISPR-STOP: gene silencing through base-editing-induced nonsense mutations. Nat. Methods 14, 710–712 (2017).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Song, M. et al. Sequence-specific prediction of the efficiencies of adenine and cytosine base editors. Nat. Biotechnol. 38, 1037–1043 (2020).
Kim, H. K. et al. In vivo high-throughput profiling of CRISPR–Cpf1 activity. Nat. Methods 14, 153–159 (2017).
Kim, H. K. et al. Deep learning improves prediction of CRISPR–Cpf1 guide RNA activity. Nat. Biotechnol. 36, 239–241 (2018).
Kim, H. K. et al. SpCas9 activity prediction by DeepSpCas9, a deep learning-based model with high generalization performance. Sci. Adv. 5, eaax9249 (2019).
Kim, H. K. et al. High-throughput analysis of the activities of xCas9, SpCas9-NG and SpCas9 at matched and mismatched target sequences in human cells. Nat. Biomed. Eng. 4, 111–124 (2020).
Kim, N. et al. Prediction of the sequence-specific cleavage activity of Cas9 variants. Nat. Biotechnol. 38, 1328–1336 (2020).
Kim, H. K. et al. Predicting the efficiency of prime editing guide RNAs in human cells. Nat. Biotechnol. 39, 198–206 (2021).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 15, 271–274 (2018).
Michlits, G. et al. CRISPR-UMI: single-cell lineage tracing of pooled CRISPR–Cas9 screens. Nat. Methods 14, 1191–1197 (2017).
Schmierer, B. et al. CRISPR/Cas9 screening using unique molecular identifiers. Mol. Syst. Biol. 13, 945 (2017).
Arbab, M. et al. Determinants of base editing outcomes from target library analysis and machine learning. Cell 182, 463–480 (2020).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Ghandi, M. et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569, 503–508 (2019).
Carter, H. et al. Cancer-specific high-throughput annotation of somatic mutations: computational prediction of driver missense mutations. Cancer Res. 69, 6660–6667 (2009).
Ng, P. C. & Henikoff, S. Predicting deleterious amino acid substitutions. Genome Res. 11, 863–874 (2001).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Miosge, L. A. et al. Comparison of predicted and actual consequences of missense mutations. Proc. Natl Acad. Sci. USA 112, E5189–E5198 (2015).
Sun, S. et al. An extended set of yeast-based functional assays accurately identifies human disease mutations. Genome Res. 26, 670–680 (2016).
Chen, H. et al. Comprehensive assessment of computational algorithms in predicting cancer driver mutations. Genome Biol. 21, 43 (2020).
Markusic, D., Oude-Elferink, R., Das, A. T., Berkhout, B. & Seppen, J. Comparison of single regulated lentiviral vectors with rtTA expression driven by an autoregulatory loop or a constitutive promoter. Nucleic Acids Res. 33, e63 (2005).
Yi, S. A. et al. HPV-mediated nuclear export of HP1γ drives cervical tumorigenesis by downregulation of p53. Cell Death Differ. 27, 2537–2551 (2020).
Eekels, J. J. M. et al. A competitive cell growth assay for the detection of subtle effects of gene transduction on cell proliferation. Gene Ther. 19, 1058–1064 (2012).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Sequist, L. V. et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci. Transl. Med. 3, 75ra26 (2011).
Ganesan, P. et al. Epidermal growth factor receptor P753S mutation in cutaneous squamous cell carcinoma responsive to cetuximab-based therapy. J. Clin. Oncol. 34, e34–e37 (2016).
Stabile, L. P. et al. Combined targeting of the estrogen receptor and the epidermal growth factor receptor in non-small cell lung cancer shows enhanced antiproliferative effects. Cancer Res. 65, 1459–1470 (2005).
Landrum, M. J. et al. ClinVar: improvements to accessing data. Nucleic Acids Res. 48, D835–D844 (2020).
Chen, Y. et al. PHLDA1, another PHLDA family protein that inhibits Akt. Cancer Sci. 109, 3532–3542 (2018).
Nagai, M. A. Pleckstrin homology-like domain, family A, member 1 (PHLDA1) and cancer. Biomed. Rep. 4, 275–281 (2016).
Botti, E. et al. Developmental factor IRF6 exhibits tumor suppressor activity in squamous cell carcinomas. Proc. Natl Acad. Sci. USA 108, 13710–13715 (2011).
Jobling, R. et al. Monozygotic twins with variable expression of Van der Woude syndrome. Am. J. Med. Genet. A 155A, 2008–2010 (2011).
Stupack, D. G. Caspase-8 as a therapeutic target in cancer. Cancer Lett. 332, 133–140 (2013).
Jia, D. et al. Crebbp loss drives small cell lung cancer and increases sensitivity to HDAC inhibition. Cancer Discov. 8, 1422–1437 (2018).
Pasqualucci, L. et al. Inactivating mutations of acetyltransferase genes in B-cell lymphoma. Nature 471, 189–195 (2011).
Cuella-Martin, R. et al. Functional interrogation of DNA damage response variants with base editing screens. Cell 184, 1081–1097 (2021).
Sánchez-Rivera, F. J. et al. Base editing sensor libraries for high-throughput engineering and functional analysis of cancer-associated single nucleotide variants. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01172-3 (2022).
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Li, X. et al. Base editing with a Cpf1–cytidine deaminase fusion. Nat. Biotechnol. 36, 324–327 (2018).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015).
Nishimasu, H. et al. Engineered CRISPR–Cas9 nuclease with expanded targeting space. Science 361, 1259–1262 (2018).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
Anders, C., Bargsten, K. & Jinek, M. Structural plasticity of PAM recognition by engineered variants of the RNA-guided endonuclease Cas9. Mol. Cell 61, 895–902 (2016).
Kleinstiver, B. P. et al. Engineered CRISPR–Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Walton, R. T., Christie, K. A., Whittaker, M. N. & Kleinstiver, B. P. Unconstrained genome targeting with near-PAMless engineered CRISPR–Cas9 variants. Science 368, 290–296 (2020).
Zhou, C. et al. Off-target RNA mutation induced by DNA base editing and its elimination by mutagenesis. Nature 571, 275–278 (2019).
Thuronyi, B. W. et al. Continuous evolution of base editors with expanded target compatibility and improved activity. Nat. Biotechnol. 37, 1070–1079 (2019).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Gaudelli, N. M. et al. Directed evolution of adenine base editors with increased activity and therapeutic application. Nat. Biotechnol. 38, 892–900 (2020).
Kurt, I. C. et al. CRISPR C-to-G base editors for inducing targeted DNA transversions in human cells. Nat. Biotechnol. 39, 41–46 (2021).
Zhao, D. et al. Glycosylase base editors enable C-to-A and C-to-G base changes. Nat. Biotechnol. 39, 35–40 (2021).
Hanson, G. & Coller, J. Codon optimality, bias and usage in translation and mRNA decay. Nat. Rev. Mol. Cell Biol. 19, 20–30 (2018).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Meier, J. A., Zhang, F. & Sanjana, N. E. GUIDES: sgRNA design for loss-of-function screens. Nat. Methods 14, 831–832 (2017).
Ramirez, R. D. et al. Immortalization of human bronchial epithelial cells in the absence of viral oncoproteins. Cancer Res. 64, 9027–9034 (2004).
Ellis, B. L., Potts, P. R. & Porteus, M. H. Creating higher titer lentivirus with caffeine. Hum. Gene Ther. 22, 93–100 (2011).
Dang, Y. et al. Optimizing sgRNA structure to improve CRISPR–Cas9 knockout efficiency. Genome Biol. 16, 280 (2015).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Billon, P. et al. CRISPR-mediated base editing enables efficient disruption of eukaryotic genes through induction of STOP codons. Mol. Cell 67, 1068–1079 (2017).
Behan, F. M. et al. Prioritization of cancer therapeutic targets using CRISPR–Cas9 screens. Nature 568, 511–516 (2019).
Hart, T., Brown, K. R., Sircoulomb, F., Rottapel, R. & Moffat, J. Measuring error rates in genomic perturbation screens: gold standards for human functional genomics. Mol. Syst. Biol. 10, 733 (2014).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041 (2017).
Kandasamy, K. et al. NetPath: a public resource of curated signal transduction pathways. Genome Biol. 11, R3 (2010).
Wang, G. & Fersht, A. R. Mechanism of initiation of aggregation of p53 revealed by Φ-value analysis. Proc. Natl Acad. Sci. USA 112, 2437-2442 (2015).
Zhao, D. et al. Combinatorial CRISPR–Cas9 metabolic screens reveal critical redox control points dependent on the KEAP1–NRF2 regulatory axis. Mol. Cell 69, 699–708 (2018).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in unique molecular identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Zhu, S. et al. Guide RNAs with embedded barcodes boost CRISPR-pooled screens. Genome Biol. 20, 20 (2019).
Xu, P. et al. Genome-wide interrogation of gene functions through base editor screens empowered by barcoded sgRNAs. Nat. Biotechnol. 39, 1403–1413 (2021).
Acknowledgements
We thank J. W. Choi for assisting with computational analysis. This work was supported, in part, by the National Research Foundation of Korea (grants 2017R1A2B3004198 (H.H.K.), 2017M3A9B4062403 (H.H.K.) and 2018R1A5A2025079 (H.H.K)); the Brain Korea 21 Plus Project (Yonsei University College of Medicine); the Yonsei Signature Research Cluster Program of 2021-22-0014 (H.H.K.); a grant of the MD-PhD/Medical Scientist Training Program (S.L.) through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea; Lung Cancer SPORE P50 (CA070907; J.D.M.); and the Korean Health Technology R&D Project, Ministry of Health and Welfare, Republic of Korea (grant HI21C1314 (H.H.K.)).
Author information
Authors and Affiliations
Contributions
Y.K., S.L. and H.H.K. conceived and designed the study. Y.K. and S.L. performed most of the experiments. J.P. critically contributed to computational analysis. S.C. critically assisted in the wet experiments. Y.K. and S.L. analyzed the data based on comments of H.H.K. J.D.M. generated and provided HBEC30KT-shTP53 cells (P cells). D.C. and T.P. contributed to the mathematical analysis (Supplementary Note 2). Y.K. and H.H.K. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
Yonsei University has filed a patent application based on this work, in which Y.K., S.L. and H.H.K. are listed as inventors. J.D.M. receives licensing fees from the National Institutes of Health and the University of Texas Southwestern Medical Center for distributing human cell lines. All the other authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Exon transcript profiles of P cells.
a, Expression of TP53 mRNA in P cells and HBEC30KT cells. FPKM, fragments per kilobase of transcript per million. Boxplots are represented for n = 3 biologically independent samples as follows: center line of box indicating the median, box limits indicating the upper and lower quartile; whiskers show the 1.5 times interquartile range. b, Gene set enrichment analysis (GSEA) of exon transcript profiles of HBEC30KT, P cells, and HCC4017, a lung cancer cell line. The single sample GSEA score (ssGSEA score) represents the degree to which the genes in a particular gene set are up- or down-regulated within the sample. RNA expression data were retrieved from Kim et al34.
Extended Data Fig. 2 Generation of libraries C and A.
a, The process of selecting sgRNA-target pairs for the generation of libraries C and A. SNVs, single nucleotide variants; sgRNA, single guide RNA. b, Generation of lentiviral libraries of sgRNA-encoding and target sequence pairs with unique molecular identifiers (UMIs). Oligonucleotides containing a 20-nt guide sequence, and the corresponding target sequence were synthesized and cloned into the pLenti-gRNA-puro vector to create plasmid library 1. The plasmids were then digested with BsmBI restriction enzyme and ligated with fragments containing the sgRNA scaffold sequences and UMIs to create plasmid library 2. Lentiviral libraries generated from plasmid library 2 were then transduced into cells expressing cytosine base editor (CBE) or adenine base editor (ABE) in a doxycycline-inducible manner.
Extended Data Fig. 3 Base editing efficiencies and indel frequencies at integrated target sequences.
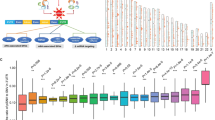
Base editing efficiencies measured at each position in the indicated region for target nucleotide Cs (a) or As (b) in integrated surrogate target sequences. Position 1 is the 5’ end of the target sequence and position 20 is immediately upstream of the NGG PAM. The numbers of analyzed target sequences (n) are as follows: n = 5,865 (position −4), 5,393 (position −3), 5,782 (position -2), 5,815 (position -1), 5,292 (position 1), 5,614 (position 2), 5,697, 6,394, 10,586, 9,382, 8,837, 5,421, 6,130, 5,339, 5,541, 5,796, 5,058, 5,723, 5,955, 5,348, 5,779, 5,437, 4,884, 5,502 (position 20) for ABE (a); n = 19,475 (position -4), 20,753 (position -3), 20,110 (position -2), 19,425 (position -1), 19,984 (position 1), 20,004 (position 2), 17,873, 24,870, 35,421, 33,186, 32,807, 19,895, 19,195, 20,227, 19,549, 18,986, 20,367, 18,793, 18,361, 20,478, 19,605, 20,975, 21,542, 22,952 (position 20) for CBE (b). Boxplots are represented as follows: center line of box indicating the median, box limits indicating the upper and lower quartile; whiskers show the 10th and 90th percentiles. Outliers are shown using dots. c, Indel frequencies measured 10 days after the transduction of sgRNA target pairs. The number of analyzed target sequence is indicated at the top of each dataset. (n = 62,000 (Library C, Replicate 1), 77,201 (Library C, Replicate 2), 21,617 (Library A, Replicate 1) and 20,913 (Library A, Replicate 2). Boxplots are represented as follows: center white dot of box indicating the median, box limits indicating the upper and lower quartile; the distributions of indel frequencies are represented with kernel densities. d, Nonsynonymous base editing efficiencies at the integrated target sequences of synonymous control sgRNAs and other sgRNAs in the given datasets. The number of synonymous and other sgRNAs are as follows; 431 and 21,055 (Library A, Replicate 1), 413 and 20,372 (Library A, Replicate 2), 2,272 and 59,390 (Library C, Replicate 1), and 2,795 and 73,691 (Library C, replicate 2), respectively. Boxplots are represented as follows: center line of box indicating the median, box limits indicating the upper and lower quartile; whiskers show the 1.5 times interquartile range.
Extended Data Fig. 4 Performance of high-throughput evaluations.
a, Distribution of median normalized log fold changes (LFCs) of 338 sgRNAs targeting essential genes depending on the nonsynonymous base editing efficiencies determined at the integrated target sequences in library C2. NT, nontargeting sgRNAs. The number of sgRNAs n = 359 (NT), 5 (<20%), 10 (20%~40%), 55 (40%~60%), 268 (>60%). Boxplots are represented as follows: center line of box indicating the median, box limits indicating the upper and lower quartile; whiskers show the 1.5 times interquartile range. (in comparison with NT, student’s t-test; NS, not significant, *P = 1.5 × 10−4, **P = 2.2 × 10−32). b,c, Receiver operating characteristic-area under the curve (ROC-AUC) analysis of LFCs for sgRNAs predicted to induce stop codons in common essential genes versus nontargeting sgRNAs in library C2 (b) and library C (c) at increasing thresholds of nonsynonymous base editing efficiencies. AUC values are indicated in parentheses. d, ROC-AUC analysis of LFCs for sgRNAs predicted to induce stop codons in common essential genes versus nontargeting controls at increasing thresholds of the number of UMIs in each sgRNA in library C. An area under curve for each UMI cutoff is shown in the parenthesis. e, Correlations between median LFCs of UMIs for sgRNAs and LFCs of UMI CPM (counts per million) for the same sgRNAs in library C2. Red dots indicate sgRNAs predicted to induce nonsense mutations in selected common essential genes. The number of sgRNAs n = 3,229 (merged), 2,913 (other sgRNAs, blue dots), 217 (sgRNAs targeting essential genes, red dots), 99 (nontargeting sgRNAs, black dots). Pearson correlation coefficients (r) are shown.
Extended Data Fig. 5 Design of small libraries and reproducibility of base editing efficiencies using these libraries.
a-b, Design of small libraries C1, C2, and A1 (a) and C3 and A2 (b). UMIs, unique molecular identifiers. c, Correlations between nonsynonymous base editing efficiencies at the integrated target sequences of biological replicates. The color of each dot was determined by the number of neighboring dots (that is, dots within a distance that is three times the radius of the dot). The base editing efficiencies were determined ten days after the initial transduction of each library into P-C or P-A cells. Only sgRNAs with more than 100 raw read counts in each replicate were included. Pearson correlation coefficients (r) are shown. The number of sgRNAs n = 3,181 (library C1), 3,063 (library C2), and 1,520 (library A1).
Extended Data Fig. 6 The number of protein variants generated by an sgRNA.
a, The proportion of sgRNAs that induce a primary protein variant. The numbers of sgRNAs are indicated in parentheses. b, The number of significant (frequency > 10%) protein variants generated by sgRNAs that induce multiple protein variants without a primary protein variant. The numbers of protein variants are indicated in parentheses.
Extended Data Fig. 7 Association between computationally predicted functions of variants and measured functions of variants.
a, The scores from driver detection algorithms (CTAT-cancer and CHASM) for 4,143 protein variants. The number of variants n = 15 (depleting), 39 (likely depleting), 864 (possibly depleting), 2,141 (neutral), 1,056 (possibly outgrowing), 25 (likely outgrowing), and 3 (outgrowing). b, The scores from algorithms that predict the functional effects of variants (SIFT and PolyPhen-2) for 3,899 protein variants. The number of variants n = 12 (depleting), 38 (likely depleting), 807 (possibly depleting), 2,009 (neutral), 1,008 (possibly outgrowing), 22 (likely outgrowing), and 3 (outgrowing). c,d, Distribution of SIFT scores (c) and PolyPhen-2 scores (d) for missense variants in common essential genes according to the LFC in library C. The number of variants n = 10 (<−0.4), 65 (−0.4~0), 82 (0~0.4). Boxplots are represented as follows: center line of box indicating the median, box limits indicating the upper and lower quartile; whiskers show the 1.5 times interquartile range.
Extended Data Fig. 8 Allele frequency tracking after transduction of sgRNA-encoding sequences into P-C or P-A cells.
sgRNA-encoding lentivirus was transduced into P-C and P-A cells at day 0 and doxycycline was added to induce expression of CBE and ABE, respectively, and maintained until day 10, after which doxycycline was removed. The functional classification results obtained from the high-throughput experiments and those from these individual experiments are shown in red and green, respectively, on the top of each graph. The mean values of two independent samples are indicated.
Extended Data Fig. 9 The results of competitive proliferation assays.
a, An example for flow cytometry gating strategy used in the competitive proliferation assays. b, Mean relative enrichment values ± standard deviation of three replicates. Student’s t test was performed under the null hypothesis that the proportions of sgRNA-transduced and nontargeting sgRNA-transduced cells would be the same. Two nontargeting sgRNAs were used as the control and the mean values of relative enrichment were used as the control.
Extended Data Fig. 10 Notable gene groups associated with outgrowing/likely outgrowing and depleting/likely depleting sgRNAs and variants.
a, (Left panel) The fraction of functionally classified sgRNAs (top) targeting cancer gene census (CGC)5 genes and primary protein variants (bottom) encoded by CGC genes in the outgrowing and likely outgrowing groups. Results from all libraries except library eC were combined. P-values from two-sided Fisher’s exact test are shown. The number of sgRNAs or variants either targeting or encoded by CGC genes among all sgRNAs or variants in each group are shown on the x-axes. (Right panel) Detailed distribution of sgRNAs predicted to introduce mutations in CGC genes (top) and variants generated in CGC genes (bottom). The number of sgRNAs or variants corresponding to each gene is specified in parentheses. b, The fraction of functionally classified sgRNAs (left) targeting Depmap common essential genes (CEGs) and protein variants (right) encoded by CEGs in the depleting and likely depleting groups. Results from all libraries except library eC were combined. P-values from two-sided Fisher’s exact test are shown. The numbers of sgRNAs or variants either targeting or encoded by CEG genes among all sgRNAs or variants in each group are shown on the x-axes.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2 and Supplementary Figs. 1–7
Supplementary Table
Supplementary Table 1. Composition of sgRNA-encoding libraries C, A, C1, C2, C3, A1, A2, dA and eC. Barcode sequences used for sorting, sgRNA sequences and target sequences, including neighboring sequences (5′-neighboring sequence (4 bp) + target sequence (20 bp + 3-bp PAM = 23 bp) + 3′-neighboring sequence (3 bp) = 30 bp of genomic DNA sequence). Information about intended mutations and DeepCBE or DeepABE efficiency scores are also included (provided as a separate Excel file). Supplementary Table 2. The results of MAGeCK analyses. RPM of four replicateUMI, LFCs, median LFCs (mLFCs), positive or negative MAGeCK RRA P values and LFCs of UMI CPM are shown for each sgRNA (provided as a separate Excel file). Supplementary Table 3. Functional classifications of sgRNAs and protein variants. a, Functional classification of sgRNAs based on the proliferation and survival (sheet 1). b, Functional classification of sgRNAs in library eC (sheet 2). c, Base editing outcomes and allele frequencies at the integrated target sequences (dependency on EGF signaling) (sheet 3). d, Potential classification of sgRNAs with low base editing efficiencies (sheet 4) (provided as a separate Excel file). Supplementary Table 4. Results of allele frequency tracking after delivery of an individual sgRNA for 20 selected sgRNAs. After lentiviral transduction of the specified individual sgRNA, protein variant frequencies were calculated from DNA sequence analysis. Endogenous DNA sequence variants encoding the same amino acid change were combined into one protein variant (provided as a separate Excel file). Supplementary Table 5. Oligonucleotides used in this study (provided in a separate Excel file).
Rights and permissions
About this article
Cite this article
Kim, Y., Lee, S., Cho, S. et al. High-throughput functional evaluation of human cancer-associated mutations using base editors. Nat Biotechnol 40, 874–884 (2022). https://doi.org/10.1038/s41587-022-01276-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-022-01276-4
This article is cited by
-
High-throughput evaluation of genetic variants with prime editing sensor libraries
Nature Biotechnology (2024)
-
Recent advances in CRISPR-based functional genomics for the study of disease-associated genetic variants
Experimental & Molecular Medicine (2024)
-
Precise genome-editing in human diseases: mechanisms, strategies and applications
Signal Transduction and Targeted Therapy (2024)
-
Generation of precision preclinical cancer models using regulated in vivo base editing
Nature Biotechnology (2024)
-
Direct measurement of engineered cancer mutations and their transcriptional phenotypes in single cells
Nature Biotechnology (2023)