Abstract
A principal challenge in the analysis of tissue imaging data is cell segmentation—the task of identifying the precise boundary of every cell in an image. To address this problem we constructed TissueNet, a dataset for training segmentation models that contains more than 1 million manually labeled cells, an order of magnitude more than all previously published segmentation training datasets. We used TissueNet to train Mesmer, a deep-learning-enabled segmentation algorithm. We demonstrated that Mesmer is more accurate than previous methods, generalizes to the full diversity of tissue types and imaging platforms in TissueNet, and achieves human-level performance. Mesmer enabled the automated extraction of key cellular features, such as subcellular localization of protein signal, which was challenging with previous approaches. We then adapted Mesmer to harness cell lineage information in highly multiplexed datasets and used this enhanced version to quantify cell morphology changes during human gestation. All code, data and models are released as a community resource.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The TissueNet dataset is available at https://datasets.deepcell.org/ for noncommercial use.
Code availability
All software for dataset construction, model training, deployment and analysis is available on our github page https://github.com/vanvalenlab/intro-to-deepcell. All code to generate the figures in this paper is available at https://github.com/vanvalenlab/publication-figures/tree/master/2021-Greenwald_Miller_et_al-Mesmer.
References
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014).
Keren, L. et al. MIBI-TOF: a multiplexed imaging platform relates cellular phenotypes and tissue structure. Sci. Adv. 5, eaax5851 (2019).
Huang, W., Hennrick, K. & Drew, S. A colorful future of quantitative pathology: validation of Vectra technology using chromogenic multiplexed immunohistochemistry and prostate tissue microarrays. Hum. Pathol. 44, 29–38 (2013).
Lin, J.-R. et al. Highly multiplexed immunofluorescence imaging of human tissues and tumors using t-CyCIF and conventional optical microscopes. eLife 7, e31657 (2018).
Gerdes, M. J. et al. Highly multiplexed single-cell analysis of formalin-fixed, paraffin-embedded cancer tissue. Proc. Natl Acad. Sci. 110, 11982–11987 (2013).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with CODEX multiplexed imaging. Cell 174, 968–981.e15 (2018).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Lee, J. H. et al. Highly multiplexed subcellular RNA sequencing in situ. Science 343, 1360–1363 (2014).
Moffitt, J. R. et al. Molecular, spatial and functional single-cell profiling of the hypothalamic preoptic region. Science 362, eaau5324 (2018).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Lubeck, E., Coskun, A. F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat Methods 11, 360–361 (2014).
Eng, C.-H. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH+. Nature 568, 235–239 (2019).
Rozenblatt-Rosen, O. et al. The human tumor atlas network: charting tumor transitions across space and time at single-cell resolution. Cell 181, 236–249 (2020).
Snyder, M. P. et al. The human body at cellular resolution: the NIH Human Biomolecular Atlas Program. Nature 574, 187–192 (2019).
Regev, A. et al. The human cell atlas white paper. Preprint at https://arxiv.org/abs/1810.05192v1 (2018).
Keren, L. et al. A structured tumor-immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387.e19 (2018).
Milo, R. & Phillips, R. Cell Biology by the Numbers 1st edn (Garland Science, 2015).
Mescher, A. Junqueira’s Basic Histology: Text and Atlas 13th edn (McGraw Hill, 2013).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
de Chaumont, F. Icy: an open bioimage informatics platform for extended reproducible research. Nat. Methods 9, 690–696 (2012).
Belevich, I., Joensuu, M., Kumar, D., Vihinen, H. & Jokitalo, E. Microscopy image browser: a platform for segmentation and analysis of multidimensional datasets. PLoS Biol. 14, e1002340 (2016).
Ronneberger, O., Fischer, P. & Brox, T. in Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015 (eds Navab, N. et al.) 234–241 (Lecture Notes in Computer Science 9351, Springer, 2015).
Valen, D. A. V. et al. Deep learning automates the quantitative analysis of individual cells in live-cell imaging experiments. PLoS Comput. Biol. 12, e1005177 (2016).
Caicedo, J. C. et al. Nucleus segmentation across imaging experiments: the 2018 Data Science Bowl. Nat. Methods 16, 1247–1253 (2019).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Hollandi, R. et al. nucleAIzer: a parameter-free deep learning framework for nucleus segmentation using image style transfer. Cell Syst. 10, 453–458.e6 (2020).
Koyuncu, C. F., Gunesli, G. N., Cetin-Atalay, R. & Gunduz-Demir, C. DeepDistance: a multi-task deep regression model for cell detection in inverted microscopy images. Med. Image Anal. 63, 101720 (2020).
Yang, L. et al. NuSeT: A deep learning tool for reliably separating and analyzing crowded cells. PLoS Comput. Biol. 16, e1008193 (2020).
Yu, W. et al. CCDB:6843, mus musculus, Neuroblastoma. CIL. Dataset. https://doi.org/10.7295/W9CCDB6843
Koyuncu, C. F., Cetin‐Atalay, R. & Gunduz‐Demir, C. Object‐oriented segmentation of cell nuclei in fluorescence microscopy images. Cytometry A 93, 1019–1028 (2018).
Ljosa, V., Sokolnicki, K. L. & Carpenter, A. E. Annotated high-throughput microscopy image sets for validation. Nat. Methods 9, 637–637 (2012).
Kumar, N. et al. A multi-organ nucleus segmentation challenge. IEEE Trans. Med. Imaging 39, 1380–1391 (2020).
Verma, R. et al. MoNuSAC2020: A Multi-organ Nuclei Segmentation and Classification Challenge. IEEE Trans. Med. Imaging 10.1109/TMI.2021.3085712 (2021).
Moen, E. et al. Accurate cell tracking and lineage construction in live-cell imaging experiments with deep learning. Preprint at bioRxiv https://doi.org/10.1101/803205 (2019).
Gamper, J. et al. PanNuke dataset extension, insights and baselines. Preprint at https://arxiv.org/abs/2003.10778v7 (2020).
Bannon, D. et al. DeepCell Kiosk: scaling deep learning–enabled cellular image analysis with Kubernetes. Nat. Methods 18, 43–45 (2021).
Haberl, M. G. et al. CDeep3M—plug-and-play cloud-based deep learning for image segmentation. Nat. Methods 15, 677–680 (2018).
Ouyang, W., Mueller, F., Hjelmare, M., Lundberg, E. & Zimmer, C. ImJoy: an open-source computational platform for the deep learning era. Nat. Methods 16, 1199–1200 (2019).
von Chamier, L. et al. Democratising deep learning for microscopy with ZeroCostDL4Mic. Nat. Commun. 12, 2276 (2021).
Hughes, A. J. et al. Quanti.us: a tool for rapid, flexible, crowd-based annotation of images. Nat. Methods 15, 587–590 (2018).
Ouyang, W., Le, T., Xu, H. & Lundberg, E. Interactive biomedical segmentation tool powered by deep learning and ImJoy. F1000Research 10, 142 (2021).
Wolny, A. et al. Accurate and versatile 3D segmentation of plant tissues at cellular resolution. eLife 9, e57613 (2020).
DeepCell Label: https://github.com/vanvalenlab/deepcell-label
Lin, T.-Y. et al. Feature pyramid networks for object detection. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 2117–2125 (IEEE, 2017).
Tan, M., Pang, R. & Le, Q. V. EfficientDet: scalable and efficient object detection. In 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 10778–10787 (IEEE, 2020).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (IEEE, 2016).
Zuiderveld, K. in Graphics Gems (ed Heckbert, P. S.) Ch. VIII.5 (Academic Press, 1994).
Chevalier, G. Make smooth predictions by blending image patches, such as for image segmentation. https://github.com/Vooban/Smoothly-Blend-Image-Patches
Meyer, F. & Beucher, S. Morphological segmentation. J. Vis. Commun. Image R 1, 21–46 (1990).
Weigert, M., Schmidt, U., Haase, R., Sugawara, K. & Myers, G. Star-convex polyhedra for 3D object detection and segmentation in microscopy. In IEEE Winter Conference on Applications of Computer Vision (WACV) 3655–3662 (IEEE, 2020).
Fu, C.-Y., Shvets, M. & Berg, A. C. RetinaMask: learning to predict masks improves state-of-the-art single-shot detection for free. Preprint at https://arxiv.org/abs/1901.03353v1 (2019).
Schürch, C. M. et al. Coordinated cellular neighborhoods orchestrate antitumoral immunity at the colorectal cancer invasive front. Cell 182, 1341–1359.e19 (2020).
Ali, H. R. et al. Imaging mass cytometry and multiplatform genomics define the phenogenomic landscape of breast cancer. Nat. Cancer 1, 163–175 (2020).
Gaglia, G. et al. HSF1 phase transition mediates stress adaptation and cell fate decisions. Nat. Cell Biol. 22, 151–158 (2020).
Nelson, D. E. et al. Oscillations in NF-κB signaling control the dynamics of gene expression. Science 306, 704–708 (2004).
Kumar, K. P., McBride, K. M., Weaver, B. K., Dingwall, C. & Reich, N. C. Regulated nuclear-cytoplasmic localization of interferon regulatory factor 3, a subunit of double-stranded RNA-activated factor 1. Mol. Cell Biol. 20, 4159–4168 (2000).
Wolff, A. C. et al. Recommendations for human epidermal growth factor receptor 2 testing in Breast Cancer: American Society of Clinical Oncology/College of American pathologists clinical practice guideline update. J. Clin. Oncol. 31, 3997–4013 (2013).
Risom, T. et al. Transition to invasive breast cancer is associated with progressive changes in the structure and composition of tumor stroma. Preprint at bioRxiv https://doi.org/10.1101/2021.01.05.425362 (2021)
Ark Analysis. https://github.com/angelolab/ark-analysis
Koss, L. G. Diagnostic Cytology and Its Histopathologic Bases. (J.B. Lippincott Company, 1979).
Erlebacher, A. Immunology of the maternal-fetal interface. Annu. Rev. Immunol. 31, 387–411 (2013).
Greenbaum, S. et al. Spatio-temporal coordination at the maternal-fetal interface promotes trophoblast invasion and vascular remodeling in the first half of human pregnancy. Preprint at bioRxiv https://doi.org/10.1101/2021.09.08.459490 (2021).
Garrido-Gomez, T. et al. Defective decidualization during and after severe preeclampsia reveals a possible maternal contribution to the etiology. Proc. Natl Acad. Sci. USA 114, E8468–E8477 (2017).
Deep Cell Core Library. Deep learning for single-cell analysis. https://github.com/vanvalenlab/deepcell-tf
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Graham, S. et al. Hover-Net: simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med. Image Anal. 58, 101563 (2019).
Tsai, H.-F., Gajda, J., Sloan, T. F. W., Rares, A. & Shen, A. Q. Usiigaci: instance-aware cell tracking in stain-free phase contrast microscopy enabled by machine learning. SoftwareX 9, 230–237 (2019).
Kiemen, A. et al. In situ characterization of the 3D microanatomy of the pancreas and pancreatic cancer at single cell resolution. bioRxiv 2020.12.08.416909 (2020) https://doi.org/10.1101/2020.12.08.416909
Cao, J. et al. Establishment of a morphological atlas of the Caenorhabditis elegans embryo using deep-learning-based 4D segmentation. Nat. Commun. 11, 6254 (2020).
Schulz, D. et al. Simultaneous multiplexed imaging of mRNA and proteins with subcellular resolution in breast cancer tissue samples by mass cytometry. Cell Syst. 6, 531 (2018).
McKinley, E. T. et al. Optimized multiplex immunofluorescence single-cell analysis reveals tuft cell heterogeneity. JCI Insight 2, e93487 (2017).
Patel, S. S. et al. The microenvironmental niche in classic Hodgkin lymphoma is enriched for CTLA-4- positive T-cells that are PD-1-negative. Blood 134, 2059–2069 (2019).
Jackson, H. W. et al. The single-cell pathology landscape of breast cancer. Nature 578, 615–620 (2020).
Rashid, R. et al. Highly multiplexed immunofluorescence images and single-cell data of immune markers in tonsil and lung cancer. Sci. Data 6, 323 (2019).
McCaffrey, E. F. et al. Multiplexed imaging of human tuberculosis granulomas uncovers immunoregulatory features conserved across tissue and blood. Preprint at bioRxiv https://doi.org/10.1101/2020.06.08.140426 (2020).
Walt, Svander et al. scikit-image: image processing in Python. PeerJ 2, e453 (2014).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980v9 (2014).
Kluyver, T. et al. in Positioning and Power in Academic Publishing: Players, Agents and Agendas (eds Schmidt, B. & Loizides, F.) (IOS Press, 2016).
Chollet, F. et al. Keras. https://keras.io (2015).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).
Reback, J. et al. pandas-dev/pandas: Pandas 1.1.3. https://doi.org/10.5281/zenodo.3509134 (2020).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. Preprint at https://arxiv.org/abs/1201.0490v4 (2012).
Waskom, M. et al. mwaskom/seaborn. https://doi.org/10.5281/zenodo.592845 (2020).
Abadi, M. et al. TensorFlow: large-scale machine learning on heterogeneous distributed systems. Preprint at https://arxiv.org/abs/1603.04467v2 (2016).
Hoyer, S. & Hamman, J. xarray: N-D labeled arrays and datasets in Python. J. Open Res. Softw. 5, 10 (2017).
Acknowledgements
We thank K. Borner, L. Cai, M. Covert, A. Karpathy, S. Quake and M. Thomson for interesting discussions; D. Glass and E. McCaffrey for feedback on the manuscript; T. Vora for copy editing; R. Angoshtari, G. Barlow, B. Bodenmiller, C. Carey, R. Coffey, A. Delmastro, C. Egelston, M. Hoppe, H. Jackson, A. Jeyasekharan, S. Jiang, Y. Kim, E. McCaffrey, E. McKinley, M. Nelson, S.-B. Ng, G. Nolan, S. Patel, Y. Peng, D. Philips, R. Rashid, S. Rodig, S. Santagata, C. Schuerch, D. Schulz, Di. Simons, P. Sorger, J. Weirather and Y. Yuan for providing imaging data for TissueNet; the crowd annotators who powered our human-in-the-loop pipeline; and all patients who donated samples for this study. This work was supported by grants from the Shurl and Kay Curci Foundation, the Rita Allen Foundation, the Susan E. Riley Foundation, the Pew Heritage Trust, the Alexander and Margaret Stewart Trust, the Heritage Medical Research Institute, the Paul Allen Family Foundation through the Allen Discovery Centers at Stanford and Caltech, the Rosen Center for Bioengineering at Caltech and the Center for Environmental and Microbial Interactions at Caltech (D.V.V.). This work was also supported by 5U54CA20997105, 5DP5OD01982205, 1R01CA24063801A1, 5R01AG06827902, 5UH3CA24663303, 5R01CA22952904, 1U24CA22430901, 5R01AG05791504 and 5R01AG05628705 from NIH, W81XWH2110143 from DOD, and other funding from the Bill and Melinda Gates Foundation, Cancer Research Institute, the Parker Center for Cancer Immunotherapy and the Breast Cancer Research Foundation (M.A.). N.F.G. was supported by NCI CA246880-01 and the Stanford Graduate Fellowship. B.J.M. was supported by the Stanford Graduate Fellowship and Stanford Interdisciplinary Graduate Fellowship. T.D. was supported by the Schmidt Academy for Software Engineering at Caltech.
Author information
Authors and Affiliations
Contributions
N.F.G., L.K., M.A. and D.V.V. conceived the project. E.M. and D.V.V. conceived the human-in-the-loop approach. L.K. and M.A. conceived the whole-cell segmentation approach. G.M., T.D., E.M., W.G. and D.V.V. developed DeepCell Label. G.M., N.F.G., E.M., I.C., W.G. and D.V.V. developed the human-in-the-loop pipeline. M.S.S., C.P., W.G. and D.V.V. developed Mesmer’s deep learning architecture. W.G., N.F.G. and D.V.V. developed model training software. C.P. and W.G. developed cloud deployment. M.S.S., S.C., W.G. and D.V.V. developed metrics software. W.G. developed plugins. N.F.G., A. Kong, A. Kagel, J.S. and O.B.-T. developed the multiplex image analysis pipeline. A. Kagel and G.M. developed the pathologist evaluation software. N.F.G., G.M. and T.H. supervised training data creation. N.F.G., C.C.F., B.J.M., K.X.L., M.F., G.C., Z.A., J.M. and S.W. performed quality control on the training data. E.S., S.G. and T.R. generated MIBI-TOF data for morphological analyses. S.C.B. helped with experimental design. N.F.G., W.G. and D.V.V. trained the models. N.F.G., W.G., G.M. and D.V.V. performed data analysis. N.F.G., G.M., M.A. and D.V.V. wrote the manuscript. M.A. and D.V.V. supervised the project. All authors provided feedback on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.A. is an inventor on patent US20150287578A1. M.A. is a board member and shareholder in IonPath Inc. T.R. has previously consulted for IonPath Inc. D.V.V and E.M. have filed a provisional patent for this work. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 DeepCell Label annotation workflow.
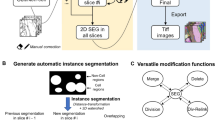
a, How multichannel images are represented and edited in DeepCell Label. b, Scalable backend for DeepCell Label that dynamically adjusts required resources based on usage, allowing concurrent annotators to work in parallel. c, Human-in-the-loop workflow diagram. Images are uploaded to the server, run through Mesmer to make predictions, and cropped to facilitate error correction. These crops are sent to the crowd to be corrected, stitched back together, run through quality control to ensure accuracy, and used to train an updated model.
Extended Data Fig. 2 Mesmer benchmarking.
a, PanopticNet architecture. Images are fed into a ResNet50 backbone coupled to a feature pyramid network. Two semantic heads produce pixel-level predictions. The first head predicts whether each pixel belongs to the interior, border, or background of a cell, while the second head predicts the center of each cell. b, Relative proportion of preprocessing, inference, and postprocessing time in PanopticNet architecture. c, Evaluation of precision, recall, and Jaccard index for Mesmer and previously published models (right) and models trained on TissueNet (left). d, Summary of TissueNet accuracy for Mesmer and selected models to facilitate future benchmarking efforts e,f Breakdown of most prevalent error types (e) and less prevalent error types (f) for Mesmer and previously published models illustrates Mesmer’s advantages over previous approaches. g, Comparison of the size distribution of prediction errors for Mesmer (left) with nuclear segmentation followed by expansion (right) shows that Mesmer’s predictions are unbiased.
Extended Data Fig. 3 TissueNet accuracy comparisons.
a, Accuracy of specialist models trained on each platform type (rows) and evaluated on data from other platform types (columns) indicates good agreement within immunofluorescence and mass spectrometry-based methods, but not across distinct methods. b, Accuracy of specialist models trained on each tissue type (rows) and evaluated on data from other tissue types (columns) demonstrates that models trained on only a single tissue type do not generalize as well to other tissue types. c, Quantification of F1 score as a function of the size of the dataset used for training. d-h, Quantification of individual error types as a function of the size of the dataset used for training. i, Representative images where Mesmer accuracy was poor, as determined by the image specific F1 score. j, Impact of image blurring on model accuracy. k, Impact of image downsampling and then upsampling on model accuracy. l, Impact of adding random noise to image on model accuracy. All scale bars are 50 μM.
Extended Data Fig. 4 3D segmentation.
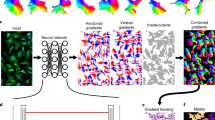
Proof of principle for using Mesmer’s segmentation predictions to generate 3D segmentations. A z-stack of 3D data is fed to Mesmer, which generates separate 2D predictions for each slice. We computationally link the segmentations predictions from each slice to form 3D objects. This approach can form the basis for human-in-the-loop construction of training data for 3D models.
Supplementary information
Rights and permissions
About this article
Cite this article
Greenwald, N.F., Miller, G., Moen, E. et al. Whole-cell segmentation of tissue images with human-level performance using large-scale data annotation and deep learning. Nat Biotechnol 40, 555–565 (2022). https://doi.org/10.1038/s41587-021-01094-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-01094-0
This article is cited by
-
Spatial multi-omics: novel tools to study the complexity of cardiovascular diseases
Genome Medicine (2024)
-
Task design for crowdsourced glioma cell annotation in microscopy images
Scientific Reports (2024)
-
Multiplex protein imaging in tumour biology
Nature Reviews Cancer (2024)
-
MAPS: pathologist-level cell type annotation from tissue images through machine learning
Nature Communications (2024)
-
Innate and adaptive immune cell interaction drives inflammasome activation and hepatocyte apoptosis in murine liver injury from immune checkpoint inhibitors
Cell Death & Disease (2024)