Abstract
Inducible expression of neoantigens in mice would enable the study of endogenous antigen-specific naïve T cell responses in disease and infection, but has been difficult to generate because leaky antigen expression in the thymus results in central T cell tolerance. Here we develop inversion-induced joined neoantigen (NINJA), using RNA splicing, DNA recombination and three levels of regulation to prevent leakiness and allow tight control over neoantigen expression. We apply NINJA to create tumor cell lines with inducible neoantigen expression, which could be used to study antitumor immunity. We also show that the genetic regulation in NINJA mice bypasses central and peripheral tolerance mechanisms and allows for robust endogenous CD8 and CD4 T cell responses on neoantigen induction in peripheral tissues. NINJA will enable studies of how T cells respond to defined neoantigens in the context of peripheral tolerance, transplantation, autoimmune diseases and cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper, its Extended Data Figures and Supplementary material. Source data are provided with this paper.
References
Curtsinger, J. M., Lins, D. C. & Mescher, M. F. Signal 3 determines tolerance versus full activation of naive CD8 T cells: dissociating proliferation and development of effector function. J. Exp. Med. 197, 1141–1151 (2003).
Harding, F. A., McArthur, J. G., Gross, J. A., Raulet, D. H. & Allison, J. P. CD28-mediated signalling co-stimulates murine T cells and prevents induction of anergy in T-cell clones. Nature 356, 607–609 (1992).
Hugues, S. et al. Distinct T cell dynamics in lymph nodes during the induction of tolerance and immunity. Nat. Immunol. 5, 1235–1242 (2004).
Iwasaki, A. & Medzhitov, R. Toll-like receptor control of the adaptive immune responses. Nat. Immunol. 5, 987–995 (2004).
Joshi, N. S. et al. Inflammation directs memory precursor and short-lived effector CD8(+) T cell fates via the graded expression of T-bet transcription factor. Immunity 27, 281–295 (2007).
Cebula, M. et al. An inducible transgenic mouse model for immune mediated hepatitis showing clearance of antigen expressing hepatocytes by CD8+ T cells. PLoS ONE 8, e68720 (2013).
Cheung, A. F., Dupage, M. J., Dong, H. K., Chen, J. & Jacks, T. Regulated expression of a tumor-associated antigen reveals multiple levels of T-cell tolerance in a mouse model of lung cancer. Cancer Res. 68, 9459–9468 (2008).
Strandt, H. et al. Neoantigen expression in steady-state langerhans cells induces CTL tolerance. J. Immunol. 199, 1626–1634 (2017).
Brockschnieder, D., Pechmann, Y., Sonnenberg-Riethmacher, E. & Riethmacher, D. An improved mouse line for Cre-induced cell ablation due to diphtheria toxin A, expressed from the Rosa26 locus. Genesis 44, 322–327 (2006).
Lee, P. et al. Conditional lineage ablation to model human diseases. Proc. Natl Acad. Sci. USA 95, 11371–11376 (1998).
Malhotra, D. et al. Tolerance is established in polyclonal CD4(+) T cells by distinct mechanisms, according to self-peptide expression patterns. Nat. Immunol. 17, 187–195 (2016).
Probst, H. C., Lagnel, J., Kollias, G. & van den Broek, M. Inducible transgenic mice reveal resting dendritic cells as potent inducers of CD8+ T cell tolerance. Immunity 18, 713–720 (2003).
Esterhazy, D. et al. Compartmentalized gut lymph node drainage dictates adaptive immune responses. Nature 569, 126–130 (2019).
Luo, X. et al. ECDI-fixed allogeneic splenocytes induce donor-specific tolerance for long-term survival of islet transplants via two distinct mechanisms. Proc. Natl Acad. Sci. USA 105, 14527–14532 (2008).
Kontos, S., Kourtis, I. C., Dane, K. Y. & Hubbell, J. A. Engineering antigens for in situ erythrocyte binding induces T-cell deletion. Proc. Natl Acad. Sci. USA 110, E60–E68 (2013).
Hlavaty, K. A. et al. Tolerance induction using nanoparticles bearing HY peptides in bone marrow transplantation. Biomaterials 76, 1–10 (2016).
Lutterotti, A. et al. Antigen-specific tolerance by autologous myelin peptide-coupled cells: a phase 1 trial in multiple sclerosis. Sci. Transl. Med. 5, 188ra175 (2013).
Wilson, D. S. et al. Synthetically glycosylated antigens induce antigen-specific tolerance and prevent the onset of diabetes. Nat. Biomed. Eng. 3, 817–829 (2019).
Hogquist, K. A., Baldwin, T. A. & Jameson, S. C. Central tolerance: learning self-control in the thymus. Nat. Rev. Immunol. 5, 772–782 (2005).
Anderson, M. S. et al. Projection of an immunological self shadow within the thymus by the aire protein. Science 298, 1395–1401 (2002).
Klein, L., Roettinger, B. & Kyewski, B. Sampling of complementing self-antigen pools by thymic stromal cells maximizes the scope of central T cell tolerance. Eur. J. Immunol. 31, 2476–2486 (2001).
McGargill, M. A., Derbinski, J. M. & Hogquist, K. A. Receptor editing in developing T cells. Nat. Immunol. 1, 336–341 (2000).
Ohashi, P. S. et al. Ablation of ‘tolerance’ and induction of diabetes by virus infection in viral antigen transgenic mice. Cell 65, 305–317 (1991).
Zehn, D. & Bevan, M. J. T cells with low avidity for a tissue-restricted antigen routinely evade central and peripheral tolerance and cause autoimmunity. Immunity 25, 261–270 (2006).
Kisielow, P., Bluthmann, H., Staerz, U. D., Steinmetz, M. & von Boehmer, H. Tolerance in T-cell-receptor transgenic mice involves deletion of nonmature CD4+8+ thymocytes. Nature 333, 742–746 (1988).
Sha, W. C. et al. Positive and negative selection of an antigen receptor on T cells in transgenic mice. Nature 336, 73–76 (1988).
Miller, J. F. & Flavell, R. A. T-cell tolerance and autoimmunity in transgenic models of central and peripheral tolerance. Curr. Opin. Immunol. 6, 892–899 (1994).
Agudo, J. et al. GFP-specific CD8 T cells enable targeted cell depletion and visualization of T-cell interactions. Nat. Biotechnol. 33, 1287–1292 (2015).
Schlake, T. & Bode, J. Use of mutated FLP recognition target (FRT) sites for the exchange of expression cassettes at defined chromosomal loci. Biochemistry 33, 12746–12751 (1994).
Raymond, C. S. & Soriano, P. ROSA26Flpo deleter mice promote efficient inversion of conditional gene traps in vivo. Genesis 48, 603–606 (2010).
Schnutgen, F. et al. Enhanced gene trapping in mouse embryonic stem cells. Nucleic Acids Res. 36, e133 (2008).
Homann, D. et al. Mapping and restriction of a dominant viral CD4+ T cell core epitope by both MHC class I and MHC class II. Virology 363, 113–123 (2007).
Mannering, S. I. et al. The insulin A-chain epitope recognized by human T cells is posttranslationally modified. J. Exp. Med. 202, 1191–1197 (2005).
Skowera, A. et al. CTLs are targeted to kill beta cells in patients with type 1 diabetes through recognition of a glucose-regulated preproinsulin epitope. J. Clin. Invest. 118, 3390–3402 (2008).
Dunne, J. L., Overbergh, L., Purcell, A. W. & Mathieu, C. Posttranslational modifications of proteins in type 1 diabetes: the next step in finding the cure? Diabetes 61, 1907–1914 (2012).
Carrillo-Vico, A., Leech, M. D. & Anderton, S. M. Contribution of myelin autoantigen citrullination to T cell autoaggression in the central nervous system. J. Immunol. 184, 2839–2846 (2010).
Dorum, S. et al. The preferred substrates for transglutaminase 2 in a complex wheat gluten digest are peptide fragments harboring celiac disease T-cell epitopes. PLoS ONE 5, e14056 (2010).
Grunewald, J. et al. Mechanistic studies of the immunochemical termination of self-tolerance with unnatural amino acids. Proc. Natl Acad. Sci. USA 106, 4337–4342 (2009).
Myers, L. K. et al. Relevance of posttranslational modifications for the arthritogenicity of type II collagen. J. Immunol. 172, 2970–2975 (2004).
Joshi, N. S. et al. Regulatory T cells in tumor-associated tertiary lymphoid structures suppress anti-tumor T cell responses. Immunity 43, 579–590 (2015).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
DuPage, M., Dooley, A. L. & Jacks, T. Conditional mouse lung cancer models using adenoviral or lentiviral delivery of Cre recombinase. Nat. Protoc. 4, 1064–1072 (2009).
Luche, H., Weber, O., Nageswara Rao, T., Blum, C. & Fehling, H. J. Faithful activation of an extra-bright red fluorescent protein in ‘knock-in’ Cre-reporter mice ideally suited for lineage tracing studies. Eur. J. Immunol. 37, 43–53 (2007).
Nielsen, M. et al. Reliable prediction of T-cell epitopes using neural networks with novel sequence representations. Protein Sci. 12, 1007–1017 (2003).
Andreatta, M. & Nielsen, M. Gapped sequence alignment using artificial neural networks: application to the MHC class I system. Bioinformatics 32, 511–517 (2016).
Acknowledgements
We thank Joshi laboratory members for reviewing the manuscript, and S. Jameson, J. Obhrai and E. Sun for helpful discussions. We also thank the Yale Cancer Center (P30 CA016359 40), Yale Flow Cytometry Core, Yale School of Medicine Histology Facility, P. Cresswell for confocal microscopy, the Swanson Biotechnology Center Preclinical Modeling facility of MIT for ES cell targeting, the University of Iowa Viral Vector Core for recombinant adenoviral vectors, P. Soriano for Ad-FLPo vector and the NIH Tetramer Core Facility. This work was supported by grants from the Howard Hughes Medical Institute (T.J.), the K22 transition to Independence grant no. NCI-K22CA200912 (N.S.J.), the Damon Runyon Cancer Foundation (N.S.J.), The G. Harold & Leila Y. Mathers Foundation (N.S.J.) and the National Institute of Diabetes and Digestive and Kidney Diseases of the National Institutes of Health under award no. P30KD034989 (N.S.J.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. T.J. is a Howard Hughes Investigator and a Daniel K. Ludwig Scholar.
Author information
Authors and Affiliations
Contributions
M.D., B.F., N.S.J. and T.J. designed the research. M.D., B.F., Y.L., M.N., I.W., J.F.C., K.A.C., G.G.F., E.A.-G., D.Y.L., G.P.C., V.G., L.M.S., A.B., J.H.W., C.C., I.M., P.G. and P.C. performed the research. M.D., B.F. and N.D.J. analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Introduction of splice sites into NM.
a, Possible splice donor and acceptor sequence candidates in the NM to create Exon 2. b, Design and inversion of Exon 2, with transcription in OFF state resulting in splicing directly from Exon 1 to Exon 3. Insertion of non-compatible Frt sites shown in light and dark blue arrows. c, FLPo recombinase activity results in permanent inversion and transcription of all exons of the NM.
Extended Data Fig. 2 Development of advanced versions of NM.
a, Schematic shows 7 versions of the NM, with modifications made at each step and the fluorescence status at either ON or OFF state. b, Flow cytometry histograms of GFP or YFP fluorescence in each version, with or without FLPo activation. Representative of 3 independent experiments. c, Western blotting for GRP94 (control, top panel) or N-terminal GFP (bottom panel) on lysates from 293T cells transiently transfected with each version of NM, with (top blot) or without (bottom blot) FLPo. Positive control (+) is cell lysate from KP-C4A3D6 after FLPo. Representative of 3 independent experiments.
Extended Data Fig. 3 Introduction of frameshift into NM.
Schematic of NM.3, which shows the two possible in silico insertions of the spliced neoantigen construct from NM.2 in YFP, where in one skipping exon 2 results in a frameshift and premature stop codon. b-c,The transcription product and fluorescence status in the ON or OFF state is shown for the frameshift version (b) or no frameshift version (c). d, Flow cytometry histograms of YFP fluorescence of the constructs from c. The frameshift version is only YFP positive after FLPo exposure. Representative of 3 independent experiments. e, 293T cells transiently transfected with plasmids expressing the indicated version of the NINJA NM were imaged by confocal microscopy after staining with an antibody specific for either the N-term portion of GFP (middle panels), for a conformational epitope of GFP (bottom panels) or with no antibody (top panels). BLUE = DAPI, RED = folded GFP, GREEN = fluorescent GFP. Representative images are shown (n = 3). f, Hydrophobicity score (top graph) in relation to amino acid position along the NM (red/grey/blue rectangle). In version NM.5 (black line) positions GP43-GP59 are predicted to be a transmembrane domain (bottom graph), and this elevated hydrophobicity is abrogated when replaced by a FLAG domain in NM.7 (red line).
Extended Data Fig. 4 Peptides from splice junction in NM are not presented as antigens.
a, The antisense DNA sequence of exon 2 encodes GP34-41 with a preceding amino acid encoded by nCA, which could encode an Ala, Ser, Thr, or Pro residue b, the predicted binding of each peptide (SIINFEKL control, GP34-41, or GP33-41 with K33A, K33S, K33T, or K33P mutations). K33P did not bind to H2-Db or stimulate T cell activation. c, Line graphs show the median fluorescence intensity (MFI) for surface expression of the indicated MHC molecule after incubation with different concentrations of the indicated peptides.
Extended Data Fig. 5 Design and development of RM.
a, Flow cytometry plots of GFP expression in transiently transfected 293s with either NM.7 alone or in combination with FLPoER or FLPoER251. FLPoER251, while leakier, is more responsive to 4-OHT treatment than FLPoER and its activity was not increased by estrogen (E2) treatment. Data reflects n=3 technical replicates per group; 3 independent experiments. (Unpaired two-tailed t tests, *, P = 0.0170, ns, P = 0.896. Measure of centre for no treatment = 15.6(+/-2.8)%, for E2 = 16.1(+/-1.5)%, FOR 4-oht = 27.9(+/-1/3)%. Error bars = mean with SEM.) b, An early regulatory module design, with pTRE:Lox-STOP-Lox (LSL):FLPoER251 2x CGG insulator, and the NM. We discovered in c, that the NM in this construct was recombined by FLPo activity in E coli, which led to the later inverted design.
Extended Data Fig. 6 Schematic diagram of completed NINJA construct.
Final version of the NINJA construct, including the final RM and NM.
Extended Data Fig. 7 Targeting of ES cell clones for NINJA.
a, Schematic showing the Rosa 26 targeting construct used for generation of the NINJA mouse. b, Southern blotting confirmation of successful target insertion of NINJA. Single experiment. c, Confirmation of germline transmission in two pups via PCR in the Rosa locus. Representative of >1000 experiments.
Extended Data Fig. 8 Time course of neoantigen-specific CD8 T cell response in NINJA.
Images show local accumulation and expansion of fLuc+ P14 T cells adoptively transferred into NINJA mice that were subsequently infected S.C. in the footpad with Ad-FLPo (107 PFU/mouse) and imaged by IVIS at the indicated day after infection. Representative mice are shown (n = 3). Intensity of signal (blue to red) indicates accumulation of T cells.
Extended Data Fig. 9 Ad-Cre infection does not lead to leaky neoantigen expression.
Quantification of endogenous H2Db/GP33-43-specific Thy1.2+CD8+ cells from the spleen (gray dots) and draining LNs (white dots) of NINJA mice 8 days after S.C. infection in the footpad with the indicated mixes of Ad-FLPo + Ad-Cre or Ad-FLPo + Ad-GFP (total dose 107 PFU/mouse) was performed by flow cytometry. Representative experiment is shown (n = 5). n.s. = difference not significant by two-tailed unpaired t test for comparisons of (Ad-FLPo + Ad-Cre) vs. (Ad-FLPo + Ad-GFP) responses in the spleen (P = 0.9) and in the draining LNs (P = 0.3). Average values ± SD are shown.
Extended Data Fig. 10 Analysis of thymic development in NINJA strains.
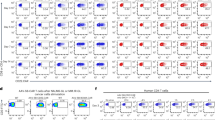
Dot plots from Fig. 3b show the average frequency ± SD of each thymocyte population as determined by FC analysis for the indicated mouse strains (each dot represents one mouse, n ≥ 3, 2 experimental repeats). n.s., not significant by two-tailed unpaired t test (0.1 ≤ P ≤ 0.9).
Supplementary information
Supplementary Information
Supplementary Notes and Tables 1–3.
Source data
Source Data Fig. 1
Unprocessed western blot.
Source Data Extended Data Fig. 2
Unprocessed western blot.
Rights and permissions
About this article
Cite this article
Damo, M., Fitzgerald, B., Lu, Y. et al. Inducible de novo expression of neoantigens in tumor cells and mice. Nat Biotechnol 39, 64–73 (2021). https://doi.org/10.1038/s41587-020-0613-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0613-1
This article is cited by
-
The screening, identification, design and clinical application of tumor-specific neoantigens for TCR-T cells
Molecular Cancer (2023)
-
Dissecting metastasis using preclinical models and methods
Nature Reviews Cancer (2023)
-
Functional T cell tolerance by peripheral tissue-based checkpoint control
Nature Immunology (2023)
-
PD-1 maintains CD8 T cell tolerance towards cutaneous neoantigens
Nature (2023)