Abstract
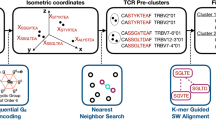
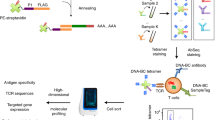
CD4+ T cells are critical to fighting pathogens, but a comprehensive analysis of human T-cell specificities is hindered by the diversity of HLA alleles (>20,000) and the complexity of many pathogen genomes. We previously described GLIPH, an algorithm to cluster T-cell receptors (TCRs) that recognize the same epitope and to predict their HLA restriction, but this method loses efficiency and accuracy when >10,000 TCRs are analyzed. Here we describe an improved algorithm, GLIPH2, that can process millions of TCR sequences. We used GLIPH2 to analyze 19,044 unique TCRβ sequences from 58 individuals latently infected with Mycobacterium tuberculosis (Mtb) and to group them according to their specificity. To identify the epitopes targeted by clusters of Mtb-specific T cells, we carried out a screen of 3,724 distinct proteins covering 95% of Mtb protein-coding genes using artificial antigen-presenting cells (aAPCs) and reporter T cells. We found that at least five PPE (Pro-Pro-Glu) proteins are targets for T-cell recognition in Mtb.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and in its Supplementary Information files.
Code availability
Two compiled standalone versions of GLIPH2 (Executable for MacOS ≥ 10.14.14 and Linux server Centos 7) are provided as Supplementary Code. Also, a web tool for GLIPH2 analysis is available at http://50.255.35.37:8080/.
References
Zhu, J. & Paul, W. E. CD4 T cells: fates, functions, and faults. Blood 112, 1557–1569 (2008).
Corbett, E. L. et al. The growing burden of tuberculosis: global trends and interactions with the HIV epidemic. Arch. Intern. Med. 163, 1009–1021 (2003).
Lindestam Arlehamn, C. S. & Sette, A. Definition of CD4 immunosignatures associated with MTB. Front. Immunol. 5, 124 (2014).
Lindestam Arlehamn, C. S. et al. A quantitative analysis of complexity of human pathogen-specific CD4 T cell responses in healthy M. tuberculosis infected South Africans. PLoS Pathog. 12, e1005760 (2016).
Altman, J. D. et al. Phenotypic analysis of antigen-specific T lymphocytes. Science 274, 94–96 (1996).
Bentzen, A. K. et al. Large-scale detection of antigen-specific T cells using peptide-MHC-I multimers labeled with DNA barcodes. Nat. Biotechnol. 34, 1037–1045 (2016).
Hadrup, S. R. et al. Parallel detection of antigen-specific T-cell responses by multidimensional encoding of MHC multimers. Nat. Methods 6, 520–526 (2009).
Newell, E. W., Klein, L. O., Yu, W. & Davis, M. M. Simultaneous detection of many T-cell specificities using combinatorial tetramer staining. Nat. Methods 6, 497–499 (2009).
Newell, E. W. et al. Combinatorial tetramer staining and mass cytometry analysis facilitate T-cell epitope mapping and characterization. Nat. Biotechnol. 31, 623–629 (2013).
Zhang, S.-Q. et al. High-throughput determination of the antigen specificities of T cell receptors in single cells. Nat. Biotechnol. 36, 1156–1159 (2018).
Simoni, Y. et al. Bystander CD8+ T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557, 575–579 (2018).
Mishto, M. & Liepe, J. Post-translational peptide splicing and T cell responses. Trends Immunol. 38, 904–915 (2017).
Laumont, C. M. et al. Noncoding regions are the main source of targetable tumor-specific antigens. Sci. Transl Med. 10, eaau5516 (2018).
Kahles, A. et al. Comprehensive analysis of alternative splicing across tumors from 8,705 patients. Cancer Cell 34, 211–224.e6 (2018).
Engelhard, V. H., Altrich-Vanlith, M., Ostankovitch, M. & Zarling, A. L. Post-translational modifications of naturally processed MHC-binding epitopes. Curr. Opin Immunol. 18, 92–97 (2006).
Cobbold, M. et al. MHC class I-associated phosphopeptides are the targets of memory-like immunity in leukemia. Sci. Transl. Med 5, 203ra125 (2013).
Bacher, P. et al. Human anti-fungal Th17 immunity and pathology rely on cross-reactivity against Candida albicans. Cell 176, 1340–1355.e15 (2019).
Bacher, P. et al. Regulatory T cell specificity directs tolerance versus allergy against aeroantigens in humans. Cell 167, 1067–1078.e16 (2016).
Glanville, J. et al. Identifying specificity groups in the T cell receptor repertoire. Nature 547, 94–98 (2017).
Huang, H. et al. Select sequencing of clonally expanded CD8+ T cells reveals limits to clonal expansion. Proc. Natl Acad. Sci. USA 116, 8995–9001 (2019).
de Sola Pool, I. & Kochen, M. Contacts and influence. Soc. Networks 1, 5–51 (1978).
Dash, P. et al. Quantifiable predictive features define epitope-specific T cell receptor repertoires. Nature 547, 89–93 (2017).
Pogorelyy, M. V. et al. Detecting T cell receptors involved in immune responses from single repertoire snapshots. PLoS Biol. 17, e3000314 (2019).
Zhang, H. et al. Investigation of antigen-specific T-cell receptor clusters in human cancers. Clin. Cancer Res. 26, 1359–1371 (2020).
Han, A., Glanville, J., Hansmann, L. & Davis, M. M. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat. Biotechnol. 32, 684–692 (2014).
Rubelt, F. et al. Individual heritable differences result in unique cell lymphocyte receptor repertoires of naive and antigen-experienced cells. Nat. Commun. 7, 11112 (2016).
Bagaev, D. V. et al. VDJdb in 2019: database extension, new analysis infrastructure and a T-cell receptor motif compendium. Nucleic Acids Res. 48, D1057–D1062 (2020).
Butler, M. O. et al. A panel of human cell-based artificial APC enables the expansion of long-lived antigen-specific CD4+ T cells restricted by prevalent HLA-DR alleles. Int. Immunol. 22, 863–873 (2010).
Rosskopf, S. et al. Creation of an engineered APC system to explore and optimize the presentation of immunodominant peptides of major allergens. Sci. Rep. 6, 31580 (2016).
Thorstenson, Y. R. et al. Allelic resolution NGS HLA typing of Class I and Class II loci and haplotypes in Cape Town, South Africa. Hum. Immunol. 79, 839–847 (2018).
Fortune, S. M. et al. Mutually dependent secretion of proteins required for mycobacterial virulence. Proc. Natl Acad. Sci. USA 102, 10676–10681 (2005).
Garces, A. et al. EspA acts as a critical mediator of ESX1-dependent virulence in Mycobacterium tuberculosis by affecting bacterial cell wall integrity. PLoS Pathog. 6, e1000957 (2010).
Karosiene, E. et al. NetMHCIIpan-3.0, a common pan-specific MHC class II prediction method including all three human MHC class II isotypes, HLA-DR, HLA-DP and HLA-DQ. Immunogenetics 65, 711–724 (2013).
Odermatt, N. T., Sala, C., Benjak, A. & Cole, S. T. Essential nucleoid associated protein mIHF (Rv1388) controls virulence and housekeeping genes in Mycobacterium tuberculosis. Sci. Rep. 8, 14214 (2018).
Brennan, M. J. et al. The enigmatic PE/PPE multigene family of mycobacteria and tuberculosis vaccination. Infect. Immun. 85, e00969 (2017).
Wagih, O. ggseqlogo: a versatile R package for drawing sequence logos. Bioinformatics 33, 3645–3647 (2017).
Blum, J. S., Wearsch, P. A. & Cresswell, P. Pathways of antigen processing. Annu. Rev. Immunol. 31, 443–473 (2013).
Boucau, J. & Le Gall, S. Antigen processing and presentation in HIV infection. Mol. Immunol. 113, 67–74 (2019).
Song, I. et al. Broad TCR repertoire and diverse structural solutions for recognition of an immunodominant CD8+ T cell epitope. Nat. Struct. Mol. Biol. 24, 395–406 (2017).
Robins, H. S. et al. Comprehensive assessment of T-cell receptor β-chain diversity in αβ T cells. Blood 114, 4099–4107 (2009).
Davis, M. M. & Boyd, S. D. Recent progress in the analysis of αβT cell and B cell receptor repertoires. Curr. Opin Immunol. 59, 109–114 (2019).
Mahomed, H. et al. Predictive factors for latent tuberculosis infection among adolescents in a high-burden area in South Africa. Int. J. Tuberc. Lung Dis.15, 331–336 (2011).
O’Neill, D. W. & Bhardwaj, N. Differentiation of peripheral blood monocytes into dendritic cells. Curr. Protoc. Immunol. 22, 24 (2005).
Acknowledgements
We would like to thank the Stanford Human Immune Monitoring Center for their high-throughput sequencing support for this project, M. Mindrinos and co-workers at Sirona Genomics for the HLA typing, S. Xue (Department of Immunology, University College London) for providing the Jurkat 76 T-cell line, J. Li for providing HLA-typed PBMCs, L. Chen and S. Chiou for valuable discussions regarding GLIPH2 optimization, H. Mahomed, W. Hanekom and members of the Adolescent Cohort Study (ACS) group for enrolment and follow-up of the Mtb-infected adolescents, R. DiFazio for help making the schematic overview and Y. Chien for constructive criticism of the manuscript, and J. Pavlovitch-Bedzyk for proofreading. This work was supported by the Bill and Melinda Gates Foundation (grant OPP1113682) and the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
H.H., C.W. and M.M.D. conceptualized the study. H.H. performed the experiments with assistance from F.R. C.W. authored the codebase, upgraded the algorithm and performed its benchmark. T.J.S. provided PBMCs from Mtb-infected adolescents. F.R. provided bulk sequencing and bulk TCR analysis. H.H., C.W. and F.R. performed the analysis. H.H. and M.M.D. wrote the manuscript with input from all authors. M.M.D. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Information
Supplementary Figures 1–7
Supplementary Table 1
Mtb-specific TCR sequences and summary
Supplementary Table 2
TCR sequences from VDJdb
Supplementary Table 3
TCR specificity groups from GLIPH2 analysis
Supplementary Table 4
Gene list of the whole Mtb ORF clone set
Supplementary Code
Two compiled standalone versions of GLIPH2
Rights and permissions
About this article
Cite this article
Huang, H., Wang, C., Rubelt, F. et al. Analyzing the Mycobacterium tuberculosis immune response by T-cell receptor clustering with GLIPH2 and genome-wide antigen screening. Nat Biotechnol 38, 1194–1202 (2020). https://doi.org/10.1038/s41587-020-0505-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0505-4
This article is cited by
-
Anchor Clustering for million-scale immune repertoire sequencing data
BMC Bioinformatics (2024)
-
Single-cell immune repertoire analysis
Nature Methods (2024)
-
Autoreactive T cells target peripheral nerves in Guillain–Barré syndrome
Nature (2024)
-
Antigen identification strategies and preclinical evaluation models for advancing tuberculosis vaccine development
npj Vaccines (2024)
-
Transcriptional signature of durable effector T cells elicited by a replication defective HCMV vaccine
npj Vaccines (2024)