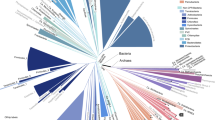
Abstract
The Genome Taxonomy Database is a phylogenetically consistent, genome-based taxonomy that provides rank-normalized classifications for ~150,000 bacterial and archaeal genomes from domain to genus. However, almost 40% of the genomes in the Genome Taxonomy Database lack a species name. We address this limitation by using commonly accepted average nucleotide identity criteria to set bounds on species and propose species clusters that encompass all publicly available bacterial and archaeal genomes. Unlike previous average nucleotide identity studies, we chose a single representative genome to serve as the effective nomenclatural ‘type’ defining each species. Of the 24,706 proposed species clusters, 8,792 are based on published names. We assigned placeholder names to the remaining 15,914 species clusters to provide names to the growing number of genomes from uncultivated species. This resource provides a complete domain-to-species taxonomic framework for bacterial and archaeal genomes, which will facilitate research on uncultivated species and improve communication of scientific results.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Genome metadata used to establish the proposed species clusters are available on the GTDB website in the files ar122_metadata.tsv and bac120_metadata.tsv. Metadata for the 24,706 GTDB species representatives are in the file sp_clusters.tsv. Genomes in the GTDB satisfying the high-quality MIMAG criteria47 are indicated in the file hq_mimag_genomes.tsv. Genome sequences are available from the NCBI Assembly database, including the 153 archaeal MAGs in BioProject PRJNA593905.
Code availability
The methodology used to establish the GTDB species clusters is implemented in version GTDB-R89 of the GTDB Species Cluster Toolkit (https://github.com/Ecogenomics/gtdb-species-clusters), a Python program available under the GNU General Public License v.3.0.
Change history
04 May 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41587-020-0539-7
References
Kyrpides, N. C. et al. Genomic Encyclopedia of Bacteria and Archaea: sequencing a myriad of type strains. PLoS Biol. 12, e1001920 (2014).
Mukherjee, S. et al. 1,003 reference genomes of bacterial and archaeal isolates expand coverage of the tree of life. Nat. Biotechnol. 35, 676–683 (2017).
Parks, D. H. et al. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat. Microbiol. 2, 1533–1542 (2017).
Chen, I. A. et al. IMG/M v.5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 47, D666–D677 (2019).
Pasolli, E. et al. Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle. Cell 176, 649–662 (2019).
Konstantinidis, K. T. & Tiedje, J. M. Towards a genome-based taxonomy for prokaryotes. J. Bacteriol. 187, 6258–6264 (2005).
Thompson, C. C. et al. Microbial taxonomy in the post-genomic era: rebuilding from scratch? Arch. Microbiol. 197, 359–370 (2015).
Garrity, G. M. A new genomics-driven taxonomy of Bacteria and Archaea: are we there yet? J. Clin. Microbiol. 54, 1956–1963 (2016).
Hugenholtz, P., Sharshewski, A. & Parks, D. H. in Microbial Evolution (ed. Ochman, H.) 55–65 (Cold Spring Harbor Laboratory Press, 2016).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Cohan, F. M. What are bacterial species? Annu. Rev. Microbiol. 56, 457–487 (2002).
Konstantinidis, K. T., Ramette, A. & Tiedje, J. M. The bacterial species definition in the genomic era. Phil. Trans. R. Soc. Lond. B Biol. Sci. 361, 1929–1940 (2006).
Fraser, C., Alm, E. J., Polz, M. F., Spratt, B. G. & Hanage, W. P. The bacterial species challenge: making sense of genetic and ecological diversity. Science 323, 741–746 (2009).
Bobay, L. M. & Ochman, H. Biological species are universal across life’s domains. Genome Biol. Evol. 9, 491–501 (2017).
Ciufo, S. et al. Using average nucleotide identity to improve taxonomic assignments in prokaryotic genomes at the NCBI. Int. J. Syst. Evol. Microbiol. 68, 2386–2392 (2018).
Chun, J. et al. Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int. J. Syst. Evol. Microbiol. 68, 461–466 (2018).
Whitman, W. B. Genome sequences as the type material for taxonomic descriptions of prokaryotes. Syst. Appl. Microbiol. 38, 217–222 (2015).
Konstantinidis, K. T. & Tiedje, J. M. Genomic insights that advance the species definition for prokaryotes. Proc. Natl Acad. Sci. USA 102, 2567–2572 (2005).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Olm, M. R. et al. Consistent metagenome-derived metrics verify and define bacterial species boundaries. mSystems 5, e00731-19 (2020).
Richter, M. & Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl Acad. Sci. USA 45, 19126–19131 (2009).
Varghese, N. J. et al. Microbial species delineation using whole genome sequences. Nucleic Acids Res. 43, 6761–6771 (2015).
Yoon, S. H., Ha, S. M., Lim, J., Kwon, S. & Chun, J. A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110, 1281–1286 (2017).
Ondov, B. D. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 17, 132 (2016).
Goris, J. et al. DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int. J. Syst. Evol. Microbiol. 57, 81–91 (2007).
Rodriguez-R, L. M. et al. The Microbial Genomes Atlas (MiGA) webserver: taxonomic and gene diversity analysis of Archaea and Bacteria at the whole genome level. Nucleic Acids Res. 46, W282–W288 (2018).
Parker, C. T., Tindall, B. J. & Garrity, G. M. International Code of Nomenclature of Prokaryotes. Int. J. Syst. Evol. Microbiol. 69, S1–S111 (2019).
Federhen, S. et al. Meeting report: GenBank microbial genomic taxonomy workshop (12–13 May, 2015). Stand. Genomic Sci. 11, 15 (2016).
Barco, R. A. et al. A genus definition for Bacteria and Archaea based on a standard genome relatedness index. mBio 11, e02475–19 (2020).
Whitman, W. B. Modest proposals to expand the type material for naming of prokaryotes. Int. J. Syst. Evol. Microbiol. 66, 2108–2112 (2016).
Konstantinidis, K. T., Rosselló-Móra, R. & Amann, R. Uncultivated microbes in need of their own taxonomy. ISME J. 11, 2399–2406 (2017).
Chuvochina, M. et al. The importance of designating type material for uncultured taxa. Syst. Appl. Microbiol. 42, 15–21 (2019).
Kitts, P. A. et al. Assembly: a resource for assembled genomes at NCBI. Nucleic Acids Res. 44, D73–D80 (2016).
Parte, A. C. LPSN—List of Prokaryotic names with Standing in Nomenclature (bacterio.net), 20 years on. Int. J. Syst. Evol. Microbiol. 68, 1825–1829 (2018).
Reimer, L. C. BacDive in 2019: bacterial phenotypic data for high-throughput biodiversity analysis. Nucleic Acids Res. 47, D631–D636 (2019).
Verslyppe, B., De Smet, W., De Baets, B., De Vos, P. & Dawyndt, P. StrainInfo introduces electronic passports for microorganisms. Syst. Appl. Microbiol. 37, 42–50 (2014).
Federhen, S. Type material in the NCBI Taxonomy Database. Nucleic Acids Res. 43, D1086–D1098 (2015).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Ficht, T. Brucella taxonomy and evolution. Future Microbiol. 5, 859–866 (2010).
Verger, J. M., Grimont, F., Grimont, P. A. D. & Grayon, M. Brucella, a monospecific genus as shown by deoxyribonucleic acid hybridization. Int. J. Syst. Evol. Microbiol. 35, 292–295 (1985).
Riojas, M. A., McGough, K. J., Rider-Riojas, C. J., Rastogi, N. & Hazbón, M. H. Phylogenomic analysis of the species of the Mycobacterium tuberculosis complex demonstrates that Mycobacterium africanum, Mycobacterium bovis, Mycobacterium caprae, Mycobacterium microti and Mycobacterium pinnipedii are later heterotypic synonyms of Mycobacterium tuberculosis. Int. J. Syst. Evol. Microbiol. 68, 324–332 (2018).
Liu, G. H.et al. Genome-based reclassification of Bacillus plakortidis Borchert et al. 2007 and Bacillus lehensis Ghosh et al. 2007 as a later heterotypic synonym of Bacillus oshimensis Yumoto et al. 2005; Bacillus rhizosphaerae Madhaiyan et al. 2011 as a later heterotypic synonym of Bacillus clausii Nielsen et al. 1995. Antonie Van Leeuwenhoek, doi:112, 1725–1730 (2019).
Oren, A. Reclassification of Halomonas caseinilytica Wu et al. 2008 as a later synonym of Halomonas sinaiensis—comments on the proposal by Hwang et al., Antonie Van Leeuwenhoek 109:1345–1352, 2016. Antonie Van Leeuwenhoek 110, 171 (2017).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of Bacteria and Archaea. Nat. Biotechnol. 35, 725–731 (2017).
Peix, A., Ramírez-Bahena, M. H. & Velázquez, E. Historical evolution and current status of the taxonomy of genus Pseudomonas. Infect. Genet. Evol. 9, 1132–1147 (2009).
Bhandari, V., Ahmod, N. Z., Shah, H. N. & Gupta, R. S. Molecular signatures for Bacillus species: demarcation of the Bacillus subtilis and Bacillus cereus clades in molecular terms and proposal to limit the placement of new species into the genus Bacillus. Int. J. Syst. Evol. Microbiol. 63, 2712–2726 (2013).
Beiko, R. G. Microbial malaise: how can we classify the microbiome? Trends Microbiol. 23, 671–679 (2015).
Osterman, B. & Moriyon, I. International committee on systematics of prokaryotes; subcommittee on the taxonomy of Brucella: minutes of the meeting, 17 September 2003, Pamplona, Spain. Int. J. Syst. Evol. Microbiol. 56, 1173–1175 (2006).
Fenwick, A. J. & Carroll, K. C. Practical problems when incorporating rapidly changing microbial taxonomy into clinical practice. Clin. Chem. Lab. Med. 57, e238–e240 (2019).
Lan, R. & Reeves, P. R. Escherichia coli in disguise: molecular origins of Shigella. Microbes Infect. 4, 1125–1132 (2002).
Pettengill, E. A., Pettengill, J. B. & Binet, R. Phylogenetic analyses of Shigella and enteroinvasive Escherichia coli for the identification of molecular epidemiological markers: whole-genome comparative analysis does not support distinct genera designation. Front. Microbiol. 6, 1573 (2016).
Hanage, W. P. Fuzzy species revisited. BMC Biol. 11, 41 (2013).
Evans, J. T. & Denef, V. J. To dereplicate or not to dereplicate? Preprint at bioRxiv https://doi.org/10.1101/848176 (2019).
Zhu, Q. et al. Phylogenomics of 10,575 genomes reveals evolutionary proximity between domains Bacteria and Archaea. Nat. Commun. 10, 5477 (2019).
Hug, L. A. et al. A new view of the tree of life. Nat. Microbiol. 1, 16048 (2016).
Chaumeil, P. A., Mussig, A., Hugenholtz, P. & Parks, D. H. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2020).
Leinonen, R., Sugawara, H. & Shumway, M. The Sequence Read Archive. Nucleic Acids Res. 39, D19–D21 (2011).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comp. Biol. 7, e1002195 (2011).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 26, 1641–1650 (2009).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of Bacteria and Archaea. ISME J. 6, 610–618 (2012).
Acknowledgements
We thank the members of an NSF-sponsored Microbial Taxonomy Workshop (NSF no. 1841658) for helpful discussions relating to establishing species clusters. This project was supported by an Australian Research Council Laureate Fellowship (grant no. FL150100038) awarded to P.H. and an Australian Research Council Future Fellowship (grant no. FT170100213) awarded to C.R.
Author information
Authors and Affiliations
Contributions
D.H.P. and P.H. wrote the paper with constructive suggestions from all other authors. D.H.P. designed the initial study. M.C., C.R. and P.H. provided nomenclatural advice and manual curation of species representatives where necessary. D.H.P., P.-A.C. and A.J.M. performed the bioinformatic analyses.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Tables 1–3, 5, 7, 8, 11, 14, 15, 17 and 19.
Supplementary Table 4
Genomes satisfying the assignment criteria of multiple species with validly or effectively published names
Supplementary Table 6
Species reclassified as synonyms because they have an ANI > 97% to a species with naming priority
Supplementary Table 9
Species where the difference in the mean ANI from the medoid and mean ANI from the GTDB-selected representative is >2
Supplementary Table 10
Genomes satisfying the assignment criteria for multiple GTDB species clusters
Supplementary Table 12
Species clusters that were incongruent with the proposed species clusters across all five trials evaluating the impact of forming species clusters from randomly selected representative genomes
Supplementary Table 13
Evaluation of the monophyly of the proposed species clusters
Supplementary Table 16
GTDB and NCBI species assignments for the 24,080 proposed representative genomes with a species classification in the NCBI taxonomy
Supplementary Table 18
List of 87 genomes retained in the genome dataset despite failing the QC criteria as they represent genomes of high nomenclatural or taxonomic importance
Rights and permissions
About this article
Cite this article
Parks, D.H., Chuvochina, M., Chaumeil, PA. et al. A complete domain-to-species taxonomy for Bacteria and Archaea. Nat Biotechnol 38, 1079–1086 (2020). https://doi.org/10.1038/s41587-020-0501-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0501-8
This article is cited by
-
Exploring virus-host-environment interactions in a chemotrophic-based underground estuary
Environmental Microbiome (2024)
-
Biofilm colonization and succession in a full-scale partial nitritation-anammox moving bed biofilm reactor
Microbiome (2024)
-
Particle-attached bacteria act as gatekeepers in the decomposition of complex phytoplankton polysaccharides
Microbiome (2024)
-
Macroalgal microbiomes unveil a valuable genetic resource for halogen metabolism
Microbiome (2024)
-
Evaluating and improving the representation of bacterial contents in long-read metagenome assemblies
Genome Biology (2024)