Abstract
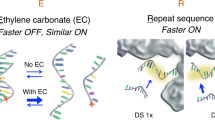
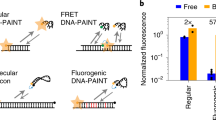
DNA point accumulation in nanoscale topography (DNA-PAINT) increases the resolution and multiplexing capabilities of super-resolution imaging, but cellular DNA interferes with DNA–DNA hybridization between target and probe in the nucleus. Here, we introduce left-handed DNA (L-DNA) oligomers that do not hybridize to natural right-handed DNA (R-DNA) and demonstrate that L-DNA-PAINT has the same specificity and multiplexing capability as R-DNA-PAINT, but improves the imaging of nuclear targets by substantially reducing background signal.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The microscopy data that support the findings of this study will be available from the corresponding author upon reasonable request, due to storage limitations. Source data are provided with this paper.
References
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–796 (2006).
Heilemann, M., Margeat, E., Kasper, R., Sauer, M. & Tinnefeld, P. Carbocyanine dyes as efficient reversible single-molecule optical switch. J. Am. Chem. Soc. 127, 3801–3806 (2005).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Schnitzbauer, J., Strauss, M. T., Schlichthaerle, T., Schueder, F. & Jungmann, R. Super-resolution microscopy with DNA-PAINT. Nat. Protoc. 12, 1198–1228 (2017).
Schueder, F. et al. Multiplexed 3D super-resolution imaging of whole cells using spinning disk confocal microscopy and DNA-PAINT. Nat. Commun. 8, 2090 (2017).
Erdelyi, M. et al. Correcting chromatic offset in multicolor super-resolution localization microscopy. Opt. Express 21, 10978–10988 (2013).
Jungmann, R. et al. Quantitative super-resolution imaging with qPAINT. Nat. Methods 13, 439–442 (2016).
Jungmann, R. et al. Single-molecule kinetics and super-resolution microscopy by fluorescence imaging of transient binding on DNA origami. Nano Lett. 10, 4756–4761 (2010).
Ricci, M. A., Manzo, C., García-Parajo, M. F., Lakadamyali, M. & Cosma, M. P. Chromatin fibers are formed by heterogeneous groups of nucleosomes in vivo. Cell 160, 1145–1158 (2015).
Otterstrom, J. et al. Super-resolution microscopy reveals how histone tail acetylation affects DNA compaction within nucleosomes in vivo. Nucleic Acids Res. 47, 8470–8484 (2019).
Lakadamyali, M. & Cosma, M. P. Visualizing the genome in high resolution challenges our textbook understanding. Nat. Methods 17, 371–379 (2020).
Hauser, N. C. et al. Utilising the left-helical conformation of L-DNA for analysing different marker types on a single universal microarray platform. Nucleic Acids Res. 34, 5101–5111 (2006).
Archetti, A. et al. Waveguide-PAINT offers an open platform for large field-of-view super-resolution imaging. Nat. Commun. 10, 1267 (2019).
Nieuwenhuizen, R. P. J. et al. Measuring image resolution in optical nanoscopy. Nat. Methods 10, 557–562 (2013).
Chagin, V. O. et al. 4D Visualization of replication foci in mammalian cells corresponding to individual replicons. Nat. Commun. 7, 11231 (2016).
Osterrieder, N., Wallaschek, N. & Kaufer, B. B. Herpesvirus genome integration into telomeric repeats of host cell chromosomes. Annu. Rev. Virol. 1, 215–235 (2014).
Kaufer, B. B. Detection of integrated herpesvirus genomes by fluorescence in situ hybridization (FISH). Methods Mol. Biol. 1064, 141–152 (2013).
Beliveau, B. J. et al. Single-molecule super-resolution imaging of chromosomes and in situ haplotype visualization using Oligopaint FISH probes. Nat. Commun. 6, 7147 (2015).
Beliveau, B. J. et al. OligoMiner provides a rapid, flexible environment for the design of genome-scale oligonucleotide in situ hybridization probes. Proc. Natl Acad. Sci. USA 115, E2183–E2192 (2018).
Frottin, F. et al. The nucleolus functions as a phase-separated protein quality control compartment. Science 365, 342–347 (2019).
Gasser, S. M. Nuclear architecture: past and future tense. Trends Cell Biol. 26, 473–475 (2016).
Kempfer, R. & Pombo, A. Methods for mapping 3D chromosome architecture. Nat. Rev. Genet. 21, 207–226 (2019).
Boettiger, A. N. et al. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 529, 418–422 (2016).
Fabricius, V., Lefèbre, J., Geertsema, H., Marino, S. F. & Ewers, H. Rapid and efficient C-terminal labeling of nanobodies for DNA-PAINT. J. Phys. D: Appl. Phys. https://doi.org/10.1101/389445 (2018).
Icha, J., Kunath, C., Rocha-Martins, M. & Norden, C. Independent modes of ganglion cell translocation ensure correct lamination of the zebrafish retina. J. Cell Biol. 215, 259–275 (2016).
Saviola, A. J. et al. Chromatin profiles of chromosomally integrated human herpesvirus. Front. Microbiol. https://doi.org/10.3389/fmicb.2019.01408 (2019).
Wallaschek, N. et al. The telomeric repeats of human herpesvirus 6A (HHV-6A) are required for efficient virus integration. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1005666 (2016).
Thévenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998).
Acknowledgements
We thank all members of the Ewers laboratory for helpful discussions. This study was supported by the ERC (starting grant no. Stg 677673 awarded to B.B.K.) and Deutsche Forschungsgemeinschaft (project no. 278001972 - TRR 186 and SFB 958 to H.E.).
Author information
Authors and Affiliations
Contributions
H.J.G. and H.E. developed the initial workflow, conceived the project and established the L-DNA-PAINT approach. G.A. conducted the FISH experiments. V.F. synthesized and labeled the left-handed nanobody and secondary antibody. H.J.G. acquired and analyzed the data. H.J.G. and H.E. wrote the manuscript. G.A. and B.B.K. edited the manuscript. B.B.K. offered guidance and resources for the project. H.E. supervised the project. All authors have reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Kinetics of L-DNA and R-DNA PAINT.
a, Titration experiment of L-DNA imagers (LP3) on L-DNA binders (LB3) and R-DNA imagers (P3) on R-DNA binders (LB3). Single molecule images are 5 µm x 5 µm. b, Plot of total number of localizations per frame in the titration experiment was quantified for L- and R-DNA and found to be very similar. No single molecules were detected for 100 nM of imagers and as such excluded from these graphs. Boxplots (centre is the median; box are the IQR; whiskers are 1.5 × IQR; mean values are represented by open squares; X represent the 1% and 99% percentiles). c, Fluorescence on-times as a result of imager-binder hybridization were extracted. The histogram represents the distribution of on-times for R-DNA and L-DNA and was fitted with a single exponential, yielding a half time of 230 ± 20 ms and 220 ± 10 ms, respectively. Data were gathered from 3 independent measurements. d, The time between subsequent imager-binder hybridization event is defined as the off-time. For 1 nM of imager, the off-times were found to be 61 ± 4 s and 59 ± 6 s for R- and L-DNA, respectively. Data were gathered from 3 independent measurements.
Extended Data Fig. 2 Nuclear localization density is enhanced by R-DNA imagers.
P1 and LP1 as well as P12 and LP12, that are coupled to an Atto655 fluorophore, show a significant difference in the ratio of localization density in the nucleus over the cytoplasm. No significant difference was detected between Atto655 or Cy3B as fluorescent dyes. P-values are obtained from two-sided paired t-Tests 16 cells stained with Cy3B-P1 and 33 cells for P1 and LP1 in 3 independent experiments and 20 cells in 2 independent measurements for P13 and LP13. Boxplots (center is the median; box are the IQR; whiskers are 1.5 × IQR; mean values are represented by open squares).
Extended Data Fig. 3 Plot of relative localization density of R-DNA over L-DNA for multiplexing experiments initiated with either R-DNA or L-DNA.
Cells were imaged with sequence-identical R-DNA and L-DNA imagers, sequentially. No significant difference in the relative localization densities of R-DNA over L-DNA was found when the image sequence was initiated with R-DNA (pink) or L-DNA (blue). P-value was obtained by two-tailed t-test of 61 and 52 cells for R-DNA and L-DNA initiation, respectively, in 17 independent experiments. Boxplots (centre is the median; box are the IQR; whiskers are 1.5 × IQR; mean values are represented by open squares).
Extended Data Fig. 4 Detection of integrated HHV-6A genomes by FISH.
FISH was performed on HEK293T cells harboring two copies of an integrated HHV-6A genome as described previously18. The two viral integration sites were detected in the cell nuclei using HHV-6A specific probes by FISH, matching the detection using the DNA-PAINT approach, and could be successfully reproduced in 30 independent FISH experiments. Scale bar is 2 µm.
Extended Data Fig. 5 Localization of R-DNA and L-DNA imagers in the nucleus of FISH samples.
R-DNA imagers strongly localize to the nucleus of FISH samples, both without DNA-FISH probes and with DNA-FISH probes coupled to complementary R-DNA binders. L-DNA imagers are not detected in the FISH samples omitting the FISH probes, but show specific loci where the samples are labeled with DNA-FISH probes. Scale bars are 5 µm. All images are reconstructed from 5000 frames and 2 nM imager concentrations.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data for Fig. 1e.
Source Data Fig. 2
Intensity profile for Fig. 2b.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Rights and permissions
About this article
Cite this article
Geertsema, H.J., Aimola, G., Fabricius, V. et al. Left-handed DNA-PAINT for improved super-resolution imaging in the nucleus. Nat Biotechnol 39, 551–554 (2021). https://doi.org/10.1038/s41587-020-00753-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-00753-y
This article is cited by
-
Picasso-server: a community-based, open-source processing framework for super-resolution data
Communications Biology (2022)
-
DNA-PAINT takes a left turn
Nature Methods (2021)