Abstract
Standardized benchmarking approaches are required to assess the accuracy of variants called from sequence data. Although variant-calling tools and the metrics used to assess their performance continue to improve, important challenges remain. Here, as part of the Global Alliance for Genomics and Health (GA4GH), we present a benchmarking framework for variant calling. We provide guidance on how to match variant calls with different representations, define standard performance metrics, and stratify performance by variant type and genome context. We describe limitations of high-confidence calls and regions that can be used as truth sets (for example, single-nucleotide variant concordance of two methods is 99.7% inside versus 76.5% outside high-confidence regions). Our web-based app enables comparison of variant calls against truth sets to obtain a standardized performance report. Our approach has been piloted in the PrecisionFDA variant-calling challenges to identify the best-in-class variant-calling methods within high-confidence regions. Finally, we recommend a set of best practices for using our tools and evaluating the results.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Raw sequence data used in the PrecisionFDA Truth Challenge were previously deposited in the NCBI SRA with the accession codes SRX847862 to SRX848317. Benchmark calls from GIAB used in the PrecisionFDA challenges and in the examples in Tables 3 and 4 are available at ftp://ftp-trace.ncbi.nlm.nih.gov/giab/ftp/release/. VCFs submitted to the PrecisionFDA challenge and benchmarking results are available at https://precision.fda.gov/, where browse access is granted immediately upon requesting account.
Code availability
All code for benchmarking developed for this manuscript are linked to from the GA4GH Benchmarking Team GitHub repository at https://github.com/ga4gh/benchmarking-tools. The hap.py benchmarking toolkit is available at https://github.com/Illumina/hap.py.
Change history
21 March 2019
In the version of this article initially published online, two pairs of headings were switched with each other in Table 4: “Recall (PCR free)” was switched with “Recall (with PCR),” and “Precision (PCR free)” was switched with “Precision (with PCR).” The error has been corrected in the print, PDF and HTML versions of this article.
References
Yang, Y. et al. Molecular findings among patients referred for clinical whole-exome sequencing. J. Am. Med. Assoc. 312, 1870–1879 (2014).
Xue, Y., Ankala, A., Wilcox, W. R. & Hegde, M. R. Solving the molecular diagnostic testing conundrum for Mendelian disorders in the era of next-generation sequencing: single-gene, gene panel, or exome/genome sequencing. Genet. Med. 17, 444–451 (2015).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Zook, J. M. et al. Integrating human sequence data sets provides a resource of benchmark SNP and indel genotype calls. Nat. Biotechnol. 32, 246–251 (2014).
Zook, J. M. et al. Extensive sequencing of seven human genomes to characterize benchmark reference materials. Sci. Data 3, 160025 (2016).
Eberle, M. A. et al. A reference data set of 5.4 million phased human variants validated by genetic inheritance from sequencing a three-generation 17-member pedigree. Genome Res. 27, 157–164 (2017).
Zook, J. et al. An open resource for accurately benchmarking small variant and reference calls. Nat. Biotechnol. https://doi.org/10.1038/s41587-019-0074-6 (2019).
Li, H. et al. New synthetic-diploid benchmark for accurate variant calling evaluation. Preprint at bioRxiv https://doi.org/10.1101/223297 (2017).
Highnam, G. et al. An analytical framework for optimizing variant discovery from personal genomes. Nat. Commun. 6, 6275 (2015).
Cleary, J. G. et al. Comparing variant call files for performance benchmarking of next-generation sequencing variant calling pipelines. Preprint at bioRxiv https://doi.org/10.1101/023754 (2015).
Sun, C. & Medvedev, P. VarMatch: robust matching of small variant datasets using flexible scoring schemes. Bioinformatics 33, 1301–1308 (2017).
Talwalkar, A. et al. SMaSH: a benchmarking toolkit for human genome variant calling. Bioinformatics 30, 2787–2795 (2014).
The Variant Call Format Specification https://samtools.github.io/hts-specs/VCFv4.3.pdf (2017).
Chen, B. et al. Good Laboratory Practices for Molecular Genetic Testing for Heritable Diseases and Conditions (Centers for Disease Control and Prevention, 2009).
Mattocks, C. J. et al. A standardized framework for the validation and verification of clinical molecular genetic tests. Eur. J. Hum. Genet. 18, 1276–1288 (2010).
Gargis, A. S. et al. Assuring the quality of next-generation sequencing in clinical laboratory practice. Nat. Biotechnol. 30, 1033–1036 (2012).
Rehm, H. L. et al. ACMG clinical laboratory standards for next-generation sequencing. Genet. Med. 15, 733–747 (2013).
Aziz, N. et al. College of American Pathologists’ laboratory standards for next-generation sequencing clinical tests. Arch. Pathol. Lab. Med. 139, 481–493 (2015).
Roy, S. et al. Standards and guidelines for validating next-generation sequencing bioinformatics pipelines: a joint recommendation of the association for molecular pathology and the college of american pathologists. J. Mol. Diagn. 20, 4–27 (2018).
Krusche, P. Haplotype comparison tools / hap.py. http://github.com/illumina/hap.py (2018).
Hasan, M. S., Wu, X., Watson, L. T., Li, Z. & Zhang, L. UPS-indel: a universal positioning system for indels. Preprint at bioRxiv https://doi.org/10.1101/133553 (2017).
Tan, A., Abecasis, G. R. & Kang, H. M. Unified representation of genetic variants. Bioinformatics 31, 2202–2204 (2015).
Kaplanis, J. et al. Exome-wide assessment of the functional impact and pathogenicity of multi-nucleotide mutations. Preprint at bioRxiv https://doi.org/10.1101/258723 (2018).
Ball, M. P. et al. A public resource facilitating clinical use of genomes. Proc. Natl Acad. Sci. USA 109, 11920–11927 (2012).
Lincoln, S. E. et al. An interlaboratory study of complex variant detection. Preprint at bioRxiv https://doi.org/10.1101/218529 (2017).
Ewing, A. D. et al. Combining tumor genome simulation with crowdsourcing to benchmark somatic single-nucleotide-variant detection. Nat. Methods 12, 623–630 (2015).
Novak, A. M. et al. Genome graphs. Preprint at bioRxiv https://doi.org/10.1101/101378 (2017).
Paten, B., Novak, A. M., Eizenga, J. M. & Garrison, E. Genome graphs and the evolution of genome inference. Genome Res. 27, 665–676 (2017).
Garrison, E. et al. Variation graph toolkit improves read mapping by representing genetic variation in the reference. Nat. Biotechnol. 36, 875–879 (2018).
Schneider, V. A. et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res. 27, 849–864 (2017).
Acknowledgements
We thank GA4GH, especially S. Keenan, D. Lloyd, and R. Nag, for their support in hosting and organizing the Benchmarking Team. We thank the many contributors to Benchmarking Team and GIAB discussions over the past few years, especially D. Church, S. Lincoln, H. Li, A. Talwalker, K. Jacobs, and B. O’Fallon. Certain commercial equipment, instruments, or materials are identified to specify adequate experimental conditions or reported results. Such identification does not imply recommendation or endorsement by the NIST or the Food and Drug Administration, nor does it imply that the equipment, instruments, or materials identified are necessarily the best available for the purpose.
Author information
Authors and Affiliations
Consortia
Contributions
P.K., L.T., P.C.B., C.E.M., F.M.d.l.V., M.A.E., R.T., B.F., M.F., M.S., and J.M.Z. wrote the manuscript. P.K., L.T., F.M.d.l.V., B.L.M., and M.G.-P. designed and implemented the benchmarking tools. Z.T., S.L., G.A., and J.M.Z. designed and/or analyzed results from the PrecisionFDA Challenges. P.K., L.T., G.A., B.A.C., M.S., and J.M.Z. designed the project. All authors contributed to GA4GH Benchmarking Team discussions about this work.
Corresponding author
Ethics declarations
Competing interests
P.K., B.L.M., M.G., and M.A.E. are employees of, and/or hold stock in, Illumina. R.T. is an employee of, and holds stock in, Invitae. G.A. is an employee of DNAnexus. B.F. is an employee of Veritas Genetics and holds leadership positions in AMP, CLSI, CAP, and ClinGen. L.T. is an employee of Real Time Genomics. C.E.M. is a founder of Onegevity Health and Biotia, Inc.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
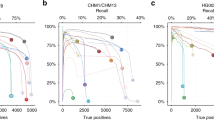
Supplementary Figure 1 Example standardized HTML report output from hap.py.
(a) Tier 1 high-level metrics output in the default view. (b) Precision-recall curve using QUAL field, where the black point is all indels, the blue point is only PASS indels, the dotted blue line is the precision-recall curve for all indels, and the solid blue line is the precision-recall curve for PASS indels. (c) Tier 2 more detailed metrics and stratifications by variant type and genome context.
Supplementary Figure 2 Hybrid Genome in a Bottle and Platinum Genomes truthset.
The hybrid truth set combines variants from Genome in a Bottle and Platinum Genomes into a single, more comprehensive gold standard. Intersection counts are shown for Genome in a Bottle (GiaB) v3.3.2 GRCh37 compared with Platinum Genomes (PG) v2016.1 as reported by hap.py v0.3.7. The union of both callsets was then re-validated using k-mer testing of inherited haplotypes in the CEPH 1463 pedigree, with all passing calls added to the hybrid truth set (Supplementary Note 4).
Supplementary Figure 3 Two examples in NA12878 where local phasing of variants can affect the interpretation.
(a) In this case, if the SNVs are interpreted independently then they are two missense mutations, and if they are interpreted together then a stop codon has been gained. (b) In this case, if the SNVs are interpreted independently then there is one missense mutation and one gained stop codon, and if they are interpreted together then it is just a missense mutation. If these events were heterozygous without phasing information, then the interpretation would be ambiguous from the VCF.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Tables 1–2 and Supplementary Notes 1–5
Rights and permissions
About this article
Cite this article
Krusche, P., Trigg, L., Boutros, P.C. et al. Best practices for benchmarking germline small-variant calls in human genomes. Nat Biotechnol 37, 555–560 (2019). https://doi.org/10.1038/s41587-019-0054-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0054-x