Abstract
Pervasive transcriptional activity is observed across diverse species. The genomes of extant organisms have undergone billions of years of evolution, making it unclear whether these genomic activities represent effects of selection or ‘noise’1,2,3,4. Characterizing default genome states could help understand whether pervasive transcriptional activity has biological meaning. Here we addressed this question by introducing a synthetic 101-kb locus into the genomes of Saccharomyces cerevisiae and Mus musculus and characterizing genomic activity. The locus was designed by reversing but not complementing human HPRT1, including its flanking regions, thus retaining basic features of the natural sequence but ablating evolved coding or regulatory information. We observed widespread activity of both reversed and native HPRT1 loci in yeast, despite the lack of evolved yeast promoters. By contrast, the reversed locus displayed no activity at all in mouse embryonic stem cells, and instead exhibited repressive chromatin signatures. The repressive signature was alleviated in a locus variant lacking CpG dinucleotides; nevertheless, this variant was also transcriptionally inactive. These results show that synthetic genomic sequences that lack coding information are active in yeast, but inactive in mouse embryonic stem cells, consistent with a major difference in ‘default genomic states’ between these two divergent eukaryotic cell types, with implications for understanding pervasive transcription, horizontal transfer of genetic information and the birth of new genes.
Similar content being viewed by others
Main
The majority of the human genome may be transcribed1,2,3,4, even though only a small fraction is annotated as discrete mature RNA species5,6. Debate remains over whether the approximately 75% of the genome that is covered by detectable transcripts4, and the approximately 80% of such transcripts for which there is predicted biochemical activity2, represent truly functional activity or random and pervasive ‘noise’7,8,9. In another eukaryotic species, the yeast S. cerevisiae, a similar fraction of the genome is transcribed10, although the genome is relatively gene-dense with an average intergenic distance11 of around 400 bp compared with the approximately 100,000 bp in the human genome12. This raises the question of whether all eukaryotic genomes are transcribed at the same level, regardless of their structure. Understanding the ‘default state’ of a genome—that is, the way a sequence lacking evolved features is acted on by the host—would be useful in interpreting the meaning of such transcriptional activity.
A genome that is active by default would present ample opportunity for transcriptional machinery to bind non-specifically, leading to spurious activity, whereas a genome that is inactive by default would generally preclude such low-specificity activity. The true default state of a genome, if such a thing exists, is difficult to determine, owing to billions of years of evolutionary pressure that has acted on existing sequences. It is thus unclear to what extent observed genomic states are passively present by default, or actively produced by chromatin-interacting proteins that recognize specific sequences selected for over time. A true default genomic state can be queried by observing activity of a newly introduced, evolutionarily naive locus. Indeed, a hypothetical ‘random genome’ experiment has been proposed as the ideal negative control for interpreting reports of large-scale genomic activity13, in which megabase-sized fragments of random DNA can be introduced into a cell and its activity compared with that of the endogenous genome. However, owing to technical limitations, such experiments have not yet been performed.
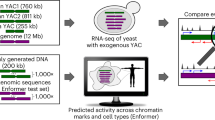
To date there has not been any well-controlled characterization of novel DNA loci in mammalian genomes, or a comparison of genomic activity for the same locus in different organismal contexts. Current techniques in synthetic genomics enable the design, assembly and delivery of very large pieces of DNA14,15. Locus-scale DNA constructs, up to hundreds of kilobases long, can be assembled de novo in yeast assembly vectors (YAVs), which exist as episomal DNA circles separate from native yeast and bacterial genomes. The ability to synthesize large DNA loci de novo enables complete design freedom over the sequence of synthetic DNA, although this realization has been limited in practice. In recent years, we have developed a workflow for synthetic regulatory genomics involving the de novo assembly of large DNA loci, including an intermediate step involving S. cerevisiae, for delivery and characterization in a desired eukaryotic context, typically mouse embryonic stem (ES) cells16,17,18,19,20. This enables straightforward design and assembly of novel DNA loci that do not exist in nature, and characterization of such loci in the distinct genomic contexts of S. cerevisiae and M. musculus. By introducing novel DNA loci to both yeast and mouse cells we can compare the default genomic states of two distinct eukaryotic hosts. Here we decided on a simple yet informative approach: to write an entire locus backwards as an initial foray into exploration of the behaviour of truly random sequences in distinct types of living cells, and an initial approximation of a ‘random genome’ experiment.
Engineering of synthetic loci
To design a novel large piece of DNA, we reversed the sequence of the human hypoxanthine phosphoribosyltransferase 1 (HPRT1) locus (Fig. 1a). By using the reverse sequence (not the reverse complement), which we refer to as HPRT1R, we ensured that the new locus lacks coding information but retains sequence features such as GC content, homopolymer runs and repeat frequency and position, that might otherwise confound analysis. This approach also provides a forward, coding ‘control’ locus, the natural HPRT1 sequence, which we previously synthesized and delivered to mouse ES cells, where it was expressed18. Statistics describing sequence composition of both synthetic loci (Table 1) indicate that although reversing the HPRT1 sequence ablates evolved regulatory elements, many potentially functional sequences, which have low information content, are still present and might be expected to occur by chance in DNA sequences of sufficient length.
a, Schematic illustrating the strategy of reversing the HPRT1 sequence to produce the HPRT1R sequence. b, The human HPRT1 locus was cloned into a assemblon vector and flanked by lox recombination sites for Big-IN integration. HPRT1R was assembled de novo from 28 synthetic segments, shown below the locus. Vector components include centromere (CEN)–autonomously replicating sequence (ARS) and LEU2 for propagation and selection in S. cerevisiae, bacterial artificial chromosome (BAC) oriS and oriV (low copy and inducible high copy origins, respectively) and the kanamycin resistance gene (Kanr) for propagation and selection in Escherichia coli, and eGFP–T2A–BSD for transient selection in mammalian cells. Chr., chromosome. c, Genomic contexts for interrogating synthetic locus activity. Episomal (Epi) and genomically integrated (chromosome XI) in S. cerevisiae, and genomically integrated (chromosome X and chromosome 3) in M. musculus. The chromosome 3 integration is monoallelic on the BL6 locus, leaving Sox2 intact on the CAST locus. d,e, DNA sequencing coverage plots from next-generation sequencing verification of assembled and integrated synthetic loci. Yeast samples were whole-genome sequenced and mouse ES cell samples were characterized by Capture-seq. Sc Epi, episomal in S. cerevisiae; Sc chr. XI, integrated on S. cerevisiae chromosome XI; Mm chr. X, integrated on M. musculus chromosome X; Mm chr. 3, integrated on M. musculus chromosome 3. (1) and (2) indicate two independent mouse ES cell clones. GC content shown as a line plot and colour-scaled. For HPRT1 Mm chr. X (1), dotted lines on the right show 2x coverage depth for most of the synthetic locus and 1x coverage depth at the edges. The relative position of the reversed HPRT1 coding sequence is indicated above the BAC in b and below the coverage plots in e.
The synthetic HPRT1 and HPRT1R loci, hereafter referred to as assemblons, were assembled in yeast assembly vectors (Fig. 1b) (see Supplementary Table 1 for details of the YAVs). YAVs facilitate Cre-mediated delivery into landing pads pre-installed in the yeast or mouse genomes (Extended Data Fig. 1a–d) (see Supplementary Table 2 for details of the landing pad), enabling readout in four contexts: in yeast, as episomes and genomic integrants, and in mouse ES cells, as genomic integrants at two distinct genomic locations (Fig. 1c). The HPRT1 locus was shuttled from a previous assemblon18 into a YAV allowing Big-IN delivery17, and the synthetic HPRT1R locus was assembled de novo from synthetic DNA pieces (Extended Data Fig. 1e,f) (see Supplementary Table 3 for synthetic DNA sequences, and Supplementary Table 4 for oligonucleotide sequences). All assemblons were verified by next-generation sequencing (Fig. 1d,e). The synthetic loci were integrated into the yeast YKL162C-A gene, a previously identified safe harbour site21, and into the mouse genome by overwriting the Hprt1 locus on the X chromosome, and at Sox2, overwriting one endogenous allele on chromosome 3 (Extended Data Fig. 1a,b) (specific genomic coordinates in Supplementary Table 5). Successful integrants were isolated and ultimately verified by whole-genome sequencing in yeast and targeted resequencing17 (Capture-seq) in mouse ES cells (Fig. 1d,e). The Capture-seq protocol involves targeted sequencing of genomic regions flanking the integration site, enabling copy number estimation of integrated loci based on comparison to the flanking regions, which have a single copy of the Hprt1 site on the X chromosome (the BL6xCAST mouse ES cells used are male with an XY karyotype) and two copies of the Sox2 site on chromosome 3. Analysis of Capture-seq data showed that one mouse ES cell clone had synthetic HPRT1 integrated as two copies (Fig. 1d), whereas all other synthetic loci were integrated as single copies. mouse ES cells with successful integration of synthetic HPRT1 were also selected for their ability to grow in hypoxanthine-aminopterin-thymidine (HAT)-supplemented medium22, demonstrating that HPRT1, which was previously shown to be expressed in mouse cells18, is able to functionally complement the loss of mouse Hprt1.
Synthetic loci are active in yeast
We first assessed activity of the novel synthetic loci in yeast, both as episomes and as chromosomal integrations (yeast strain details in Supplementary Table 6). For sequencing-based assays, replicates agreed well, as assessed by Pearson correlation of genome-wide signal depth (Extended Data Figs. 2 and 3) and by comparison with publicly available sequencing data for the same or similar assays (Extended Data Fig. 4a). Assaying chromatin accessibility by assay for transposase-accessible chromatin using sequencing (ATAC-seq), we observed multiple peaks of highly accessible chromatin across the entire synthetic locus for both HPRT1 and HPRT1R (Fig. 2a–d and Extended Data Fig. 2a,b). Using an adjacent region of the yeast genome as a reference, ATAC-seq peaks coincided with promoter regions of transcribed genes (Extended Data Fig. 4a,b). For both HPRT1 and HPRT1R synthetic loci, the highly accessible regions were conserved across replicates and between episomal and integrated loci (Extended Data Fig. 2a,b). Average ATAC-seq coverage depth was greater across the synthetic HPRT1 and HPRT1R loci compared with the genome average calculated over 100-kb sliding windows (Extended Data Fig. 4d), which was also evident when comparing the integrated synthetic loci to their surrounding native genomic regions (Fig. 2b,d and Extended Data Fig. 2b). ATAC-seq coverage depth was also greater for episomal loci compared with integrated loci, even when normalizing for estimated copy number based on DNA sequencing coverage (Extended Data Figs. 1g and 4d). The synthetic loci contained more ATAC-seq peaks over 100 kb compared with the genome average (Extended Data Fig. 4e), and although we observed peaks coinciding with the HPRT1 transcription start site (TSS), and the relative TSS position in HPRT1R, peaks generally did not correspond with known HPRT1 functional elements.
a,b, Sequencing tracks for ATAC-seq, H3K4me3 CUT&RUN, RNA-seq and CAGE-seq reads aligned to the synthetic HPRT1 locus as an episome (Epi) (a) or integrated on chromosome XI (b). c,d, Sequencing tracks of ATAC-seq, H3K4me3 CUT&RUN, RNA-seq and CAGE-seq reads aligned to the synthetic HPRT1R locus as an episome (c) or integrated on chromosome XI (d). The synthetic locus regions (HPRT1 and HPRT1R) are shaded in b,d. The HPRT1 coding sequence is indicated in a,b and the relative position corresponding to the reversed coding sequence is indicated in c,d. For loci integrated into chromosome XI (b,d), approximately 50 kb of flanking yeast genome is shown upstream and downstream of the integrated synthetic loci with annotated yeast genes indicated. RNA-seq and CAGE-seq tracks are stranded, displayed with reverse strand reads inverted and below forward strand reads. Sequencing tracks are shown for one replicate for each genomic context. e,f, Metaplots of ATAC-seq (e) and H3K4me3 CUT&RUN (f) signal at the TSS (defined by experimental CAGE-seq peaks) ±0.5 kb for HPRT1 and HPRT1R episomal assemblons as well as the yeast (Sc) genome. Shaded region shows standard error. g, Example strategy for insertion of the Sphis5 coding sequence at two experimentally identified TSSs. RNA-seq and CAGE-seq tracks are shown, as well as CAGE-seq peaks. The Sphis5 coding sequence (green arrow) is inserted with the 5′ untranslated region at the 5′ boundary of the CAGE-seq peak. Fwd, forward; rev, reverse. h, Spot assays for yeast with Sphis5 integration on the HPRT1 or HPRT1R episome, and their parental strains, on SC–Leu–His medium. For Sphis5 insertion strains, the number indicates the position of the Sphis5 insertion along the synthetic locus.
We next checked for H3K4me3 at the synthetic loci, a marker of active transcription of nearby genes. Using cleavage under targets & release using nuclease23 (CUT&RUN), we found broad coverage of H3K4me3 over both synthetic loci (Fig. 2a–d and Extended Data Fig. 2c,d). H3K4me3 coverage of HPRT1 and HPRT1R appears broader than coverage over the yeast genome, where tight peaks coincide with known promoter regions (Extended Data Fig. 4a,b). H3K4me3 coverage was also greater across episomal HPRT1 and HPRT1R loci compared to the yeast genome average and the synthetic loci contained more H3K4me3 peaks per 100 kb than the genome average (Extended Data Fig. 4f,g).
To determine whether observed chromatin accessibility and H3K4me3 patterns relate to transcription, we performed RNA sequencing (RNA-seq) for strains with episomal and chromosomally integrated synthetic loci (Fig. 2a–d and Extended Data Fig. 2e,f). RNA-seq reads mapped across both HPRT1 and HPRT1R synthetic loci, and RNA-seq peaks were consistent between replicates and between episomal and integrated loci. The synthetic loci showed RNA-seq coverage depth similar to the genome average, which is gene-dense (Extended Data Fig. 4h). We used cap analysis gene expression and sequencing24 (CAGE-seq) to map TSSs (Fig. 2a–d and Extended Data Fig. 2g,h), and found that both synthetic loci have around 3 times more CAGE-seq peaks per 100 kb than the yeast genome average (Extended Data Fig. 4i). CAGE-seq peaks map to the 5′ end of annotated and expressed yeast genes (Extended Data Fig. 4a,b), adding confidence that peaks observed in the synthetic loci are true TSSs. Using the 5′ boundary of CAGE-seq peaks as reference points, we produced metaplots of ATAC-seq and H3K4me3 signals for TSSs in the synthetic HPRT1 and HPRT1R loci and throughout the yeast genome (Fig. 2e,f and Extended Data Fig. 4c). The metaplot profiles for the synthetic loci generally match that for the yeast genome and, although they lack the precise nucleosome repeat definition seen in genome profiles obtained by averaging tens of thousands of TSSs, rough periodicity can be observed. RNA-seq and CAGE-seq signals do not appear to correspond to known gene features in the HPRT1 locus, and in fact appear to be generally depleted around the HPRT1 transcription unit and its corresponding region in the HPRT1R locus. We performed motif analysis using MEME25 on the predicted promoter regions—defined as 200 bp upstream and 100 bp downstream of identified TSSs—on ATAC-seq peaks and on ATAC-seq peaks that overlap predicted promoters (Extended Data Fig. 5). We identified a number of significantly enriched sequence motifs, including stretches of A and T reported to precede TSSs26, as well as the dyad symmetric CTCNGNCTC/GAGNCNGAG motif, and the palindromic GGTC(G/C)GACC/CCAG(C/G)CTGG motif. Although the identified motifs do not exactly match known transcription factor binding sites (TFBSs), predicting TFBSs using Tomtom27 identified a number of potential sites for stress-responsive transcription factor including Crz1, Rpn4 and Gsm1. Performing the same analysis on overlapping CAGE-predicted promoters and ATAC-seq peaks genome-wide, the only significant motifs identified are poly-A and poly-T regions.
To assess whether observed sites of transcription initiation can produce functional mRNAs, we cloned the his5 transcription unit from Schizosaccharomyces pombe (Sphis5) downstream of the predicted promoters for eight identified transcription start sites (Fig. 2g and Extended Data Fig. 6a–c). As the parental BY4741 strain is His–, only yeast expressing the Sphis5 transgene can survive on histidine dropout medium. We observed His+ colonies following transformation-mediated integration of the Sphis5 gene into four sites each of the HPRT1 and HPRT1R episomes and confirmed the His+ phenotype compared with the parental yeast strains (Fig. 2h and Extended Data Fig. 6d), demonstrating that novel TSSs are able to drive transcription of functional mRNAs and proteins. All eight tested sites appear to produce sufficiently high levels of transcription, as there were no observable fitness differences between the Sphis5 strains, even when grown in the presence of 3-amino-1,2,4-triazole (3-AT), a competitive inhibitor of the Sphis5 gene product used to titrate relatively small differences in gene expression. To determine whether any transcription factors predicted to bind to the putative promoter region motifs are responsible for transcription from the integrated transgenes, we deleted three candidate factor genes, CRZ1, RPN4 and GSM1, from yeast strains with Sphis5 inserted at HPRT1 −13493. This putative promoter region has the CTCNGNCTC/GAGNCNGAG motif that is predicted to bind these non-essential transcription factors. After identifying successful knockouts, we observed specific reduction in growth in the absence of histidine for the CRZ1 and RPN4 knockouts (Extended Data Fig. 8e).
Synthetic loci are inactive in mouse ES cells
We next assessed the activity of synthetic loci in mouse ES cells, performing ATAC-seq, RNA-seq and CUT&RUN for RNA polymerase II (RNAP2), H3K4me3, H3K27ac and H3K27me3. Sequencing results showed high correlation between replicates (Extended Data Figs. 7 and 8), as well as similar enrichment patterns as seen in mouse ES cells previously (Extended Data Fig. 9a). For HPRT1 integrated at Hprt1, we observed peaks for ATAC-seq, H3K4me3, RNAP2 and H3K27ac at the HPRT1 TSS, and RNA-seq reads mapping specifically to the HPRT1 exons (Fig. 3a and Extended Data Fig. 7a,b). These observations agree with public data from mouse ES cells28 and human ES cell lines29, and with data from an intact Hprt1 locus from this study (with HPRT1R integrated at Sox2) (Extended Data Fig. 9b–d). By contrast, the HPRT1R locus showed no activity when integrated at this same location on the X chromosome, with no peaks for ATAC-seq, H3K4me3, H3K27ac or RNAP2 anywhere across the locus (Fig. 3b and Extended Data Fig. 7c,d). There was, however, an enrichment of H3K27me3, particularly in the flanking regions. An identical chromatin signature is observed when HPRT1R is integrated at Sox2 on chromosome 3, with no peaks for ATAC-seq, H3K4me3, H3K27ac or RNAP2 and an enrichment of H3K27me3 in the flanking regions (Fig. 3c and Extended Data Fig. 7e,f). Although there is no observable transcriptional activity at HPRT1R in either genomic context, there is nonetheless a very small number of RNA-seq reads (0–4 across all replicates) mapping within the locus, which is less than the median of RNA-seq reads mapping to 100-kb windows of geneless regions genome-wide (10–30 mapped reads per 100-kb window) (Fig. 3d).
a–c, Sequencing tracks for ATAC-seq, H3K4me3, RNAP2, H3K27ac and H3K27me3 CUT&RUN, and RNA-seq over the synthetic HPRT1 and HPRT1R loci integrated into the mouse genome. One clonal replicate is shown for each integration: HPRT1 chromosome X (2) (a), HPRT1R chromosome X (1) (b) and HPRT1R chromosome 3 (1) (c). Synthetic locus regions (HPRT1 and HPRT1R) are shaded. The HPRT1 coding sequence is indicated in a. Approximately 50 kb of flanking mouse genome is included, with annotated mouse genes indicated. RNA-seq tracks are stranded, displayed with reverse strand reads inverted and below forward strand reads. d, RNA-seq read counts for the synthetic loci (circles) and for 100-kb geneless regions of the mouse genome (squares, median of 70,107 100-kb sliding windows).
HPRT1R noCpG is transcriptionally silent
The synthetic HPRT1R locus is highly enriched relative to mammalian DNA for CpG dinucleotides (Table 1), which have been implicated in Polycomb recruitment30,31,32,33,34,35. This locus has increased GC content and an enrichment of CpG islands in the flanking regions compared with the middle, gene body region (Fig. 4a), corresponding to increased H3K27me3 signal in mouse ES cells and RNA-seq signal in yeast (Extended Data Fig. 10a,b). To determine whether the artificial enrichment of CpGs at this locus underlies the increased levels of H3K27me3 and low transcriptional activity, we designed a variant of HPRT1R in which every CpG was eliminated by randomly deleting either the C or G. This resulted in a new synthetic locus, HPRT1RnoCpG, 5,600 bp shorter than HPRT1R and completely lacking CpGs. This locus was assembled de novo into the same YAV as HPRT1 and HPRT1R and delivered to mouse ES cells at the Hprt1 and Sox2 integration sites, and the locus integrity was verified by sequencing at each step (Extended Data Fig. 10c–e). By performing the same sequencing assays as for HPRT1 and HPRT1R, we found that HPRT1RnoCpG had lost H3K27me3 enrichment found in the flanking regions of HPRT1R but, notably, remained transcriptionally quiescent (Fig. 4b,c and Extended Data Fig. 10f–i).
a, GC content overlaid with HPRT1R H3K27me3 CUT&RUN in mouse ES cells. GC content was calculated over 5-kb windows, and the range from 35–49% is indicated as a black line overlaying the sequencing track. CpG islands, as predicted using EMBOSS CpGplot68, are indicated. b,c, Sequencing tracks for ATAC-seq, H3K4me3, RNAP2, H3K27ac and H3K27me3 CUT&RUN, and RNA-seq over the HPRT1RnoCpG locus integrated into the mouse genome on chromosome X, clone HPRT1RnoCpG chromosome X (1) (b) and on chromosome 3, clone HPRT1RnoCpG chromosome 3 (1) (c). The synthetic locus region (HPRT1RnoCpG) is shaded. Approximately 50 kb of flanking mouse genome is included with annotated mouse genes indicated. RNA-seq tracks are stranded, displayed with reverse strand reads inverted and below forward strand reads. The H3K27me3 CUT&RUN track for HPRT1R (copied from Fig. 3) is included below the respective tracks for HPRT1RnoCpG at each genomic position.
Discussion
By introducing synthetic reversed loci into both yeast and mouse ESCs, genomic activity across large swaths of evolutionarily naive DNA sequence can be assessed, shedding light on apparent fundamental differences in default genomic states between the two divergent cell types. In yeast, we observed widespread activity of both synthetic loci, despite the lack of promoters evolved for yeast gene expression, which did not correlate with known functional elements in the HPRT1 locus. Previous studies in yeast have reported pervasive transcription10,36,37,38,39, which is generally predicted to arise from functionally specific or productive transcription. While this work was in preparation, other groups have similarly demonstrated widespread activity from an 18-kb random sequence40, a 244-kb sequence designed for data storage41, and exogenous human42 and bacterial43 chromosomes in yeast. These results, along with those from our two synthetic reversed loci, provide independent examples of active transcription in medium-to-large DNA sequences that lack evolved yeast regulatory sequences. These studies all support a hypothesis that the default state in yeast is open and active, and newly introduced loci provide ample substrates for spurious transcription initiation. Naturally, exogenous DNA may be introduced through horizontal gene transfer in the form of conventional viruses and other infectious molecules, even across relatively vast phylogenetic distances44,45,46,47. Yeast is largely insulated by a thick cell wall during the majority of its life cycle, and lacks conventionally transmitted viruses. Thus, yeast might be an outlier, able to afford an open, active default state to a greater extent than other eukaryotic cells.
Pervasive transcription may represent an adaptive strategy in fast-growing single-celled organisms such as yeast, in which widespread transcription of even non-coding sequences provides a chance for new, potentially favourable ‘neogenes’ to arise48. We observed that the RNA-seq signal was enriched in GC-rich flanking regions of both synthetic loci, possibly reflecting the increased stability of GC-rich transcripts49,50,51, and indeed new genes in yeast preferentially emerge in GC-rich intergenic regions52. We show here that GC-rich regions also underlie increased levels of transcription across completely non-functional loci, enabling a preview of what happens before genes are established, and generally supporting an ‘expression-first’ hypothesis53 for neogene formation. Following these strains harbouring synthetic loci over multiple generations may reveal whether the widespread activity seen here is tuned, or even whether novel genes can arise from such newly introduced sequences.
In contrast to yeast, activity of the novel HPRT1R synthetic locus was largely shut down in mouse ES cells. The HPRT1 locus behaved as expected, faithfully recapitulating the activity of endogenous HPRT1 in human ES cells and of Hprt1 in mouse ES cells, indicating that the mouse ES cells have had ample opportunity to properly chromatinize the synthetic loci at the time of our analysis. HPRT1R showed no evidence of transcriptional activity in mouse ES cells, but did show that enrichment of H3K27me3 correlated with increased GC content and CpG islands, with an almost identical chromatin profile when integrated at two completely distinct genomic locations. It has been demonstrated previously that GC-rich sequences are important for Polycomb recruitment, particularly when the region is devoid of activating sequence motifs30,31,32,33,34,35. Although specific transcription factors and non-coding RNAs have also been hypothesized to have a role in Polycomb recruitment, our results are in line with observations by other groups showing that shorter sequences from Escherichia coli31 or artificial sequences designed in silico34, similarly devoid of mammalian TFBSs, were capable of recruiting PRC2 when integrated into the genome of mouse ES cells, suggesting that specific sequences are not required for PRC2 targeting.
Mammalian genomes are generally depleted of CpG dinucleotides54,55. In the HPRT1R locus, all GpC dinucleotides from the forward HPRT1 are converted to CpGs, resulting in a significant 3.75-fold enrichment of this dinucleotide compared to HPRT1, with an observed/expected CpG ratio of 1.05 compared to 0.28 in the forward sequence. CpG dinucleotides have specifically been implicated in Polycomb recruitment32,33,34,35, and could explain why H3K27me3 enrichment is observed in GC-rich regions. H3K27me3 enrichment was absent in the CpG-less version of HPRT1R, but transcriptional activity was not restored. Thus, CpG enrichment indeed has a major role in Polycomb recruitment, but this active silencing is not responsible for the observed lack of activity. It will be informative to further modify the HPRT1R locus, introducing activating sequence elements such as enhancers, promoters or entire genes, and identify what elements are sufficient for introducing transcriptional activity.
The presence of extremely few RNA-seq reads mapping to the synthetic loci suggests that spurious transcription initiation is not common, and that much low-level transcription observed genome-wide may be a signal artefact or experimental noise, and does not reflect widespread functional transcription. Conversely, transcription observed at a significantly higher level may well be functional. It should be noted that these results are obtained from embryonic stem cells, and may not translate to all mammalian cells. Indeed, it has recently been reported that short random sequences inserted into Drosophila embryos can function as developmental enhancers, but are largely non-functional in early embryos56, perhaps suggesting that early embryonic genomes are generally less permissive to activity from novel sequences. As with the synthetic loci in yeast, it will be informative to measure activity arising from the HPRT1R locus over time, or following differentiation into different cell types that might express transcription factors that recognize chance binding sites occurring in the synthetic locus.
In comparing the same synthetic loci between the two different eukaryotic contexts, we can see substantial differences in each cell type’s requirement for transcriptional activity. Both forward and reverse loci contain thousands of TFBSs for both species, and also probably contain minor evolved sequence features, such as weak–weak base pairing and mutation periodicity around positioned nucleosomes57, as well as short palindromes, that would not be ablated by sequence reversal. Despite this, the loci are only broadly transcriptionally active in yeast, and we do indeed see enrichment of AT-rich stretches and short palindromes in the putative promoters, indicating that these relatively minor features may nonetheless suffice for initiating transcription. Conversely, the loci are broadly inactive in mouse ES cells, suggesting that the basic requirements for transcription in this genomic context are much more limited. Our reversed locus, although long for a sequence insertion into the mouse genome, still only represents a small fraction of an entire genome. Animal genomes are generally transcription-sparse, with even non-conserved long non-coding RNAs identified on average every 50–100 kb in the human genome58,59. Our reversed sequence might not be long enough for chance occurrence of a sufficiently dense cluster of proper TFBSs to initiate transcription in mouse ES cells, although the current sequence length is clearly sufficient for abundant spurious transcription in yeast cells.
The question remains of whether there are default states for gene expression. It is certainly true that the vast majority of the yeast genome is heavily transcribed and translated into proteins, whereas the opposite is true of mammalian protein-coding genes. The vast majority of animal DNA is packaged during replication into nucleosomes containing histones H3.1 and H3.2 (ref. 60), whereas subsequently activated regions are replaced by more yeast-like histone H3.3 nucleosomes, which are thought to be fundamentally more compatible with transcription61. Different genomes may respond differently to random DNA—for example, replacing yeast core nucleosomes with their human counterparts62 leads to a generally less permissive chromatin state, owing to intrinsically higher affinity of human nucleosomes for DNA63,64. However, we acknowledge that no sequence is truly random, and a future test of this hypothesis might involve evaluating thousands of sequences created by a random number generator. Indeed, as genome-engineering technologies advance, Eddy’s hypothetical multi-mega-base random genome becomes ever more plausible13.
In conclusion, we have used our ability to build unnatural synthetic loci as a tool to probe the default genomic states in two different eukaryotic cell types—yeast and mouse ES cells. We show that even without evolved regulatory elements, minor sequence features appear to elicit genomic activity, underlying pervasive transcription in yeast, and CpG-dependent H3K27me3 enrichment in mouse ES cells. Our results suggest that the default state in yeast is open and active, and this widespread transcription may facilitate exploitation of rare instances of horizontal transfer and provide raw fodder for neogene formation. By contrast, the default state in mouse ES cells is inactive, suggesting much more complex and limiting requirements for transcription. Here we have characterized large, evolutionarily naive sequences in different cell types, which exhibit distinct default genomic states.
Methods
Design of synthetic loci
The synthetic HPRT1 locus has been described previously18. The synthetic HPRT1R locus was designed by reversing (but not reverse-complementing) the sequence of the human HPRT1 locus corresponding to hg38 chromosome X:134429208-134529874. HPRT1RnoCpG was designed starting with the HPRT1R sequence, using a Python script to scan the sequence for occurrences of CG and randomly delete either the C or the G. As this sequence transformation can result in the formation of new CG instances, the script was reiterated until no CG sequences remained. We used software developed in house to split the synthetic loci into smaller DNA segments for commercial DNA synthesis. HPRT1R was split into 28 segments, 27 of ~4 kb and one of ~2 kb, and HPRT1RnoCpG was split into 36 segments, 35 of ~3 kb and one of 1,300 bp. Each synthetic segment had overlaps of ~300 bp, in both termini, with the neighbouring segments. MenDEL69 was used to design primers for junction PCR screening of yeast clones harbouring the correct assembly. Synthetic DNA segments were ordered from Qinglan Biotech, and junction PCR primers were ordered from IDT.
Synthetic loci sequence features
Dinucleotides were counted across each synthetic locus. Expected CpG number was calculated as (no. of C × no. of G)/sequence length and CpG ratio was calculated as observed CpG/expected CpG. Yeast TFBSs were predicted by scanning the DNA sequences with the YEASTRACT+ database65. Mouse TFBSs were predicted using FIMO66 in the MEME suite using the JASPAR vertebrate motif database67.
Yeast assembly and BAC recovery
All yeast work was performed starting with the parental strain BY4741 using standard yeast media. HPRT1R was assembled from 28 synthetic DNA segments, first as two half-assemblies that were then combined using eSwAP-In18. HPRT1RnoCpG was assembled from 36 synthetic segments in one step. For both HPRT1R and HPRT1RnoCpG assemblies, ~50 ng each of linearized and gel-purified yeast assembly vector (YAV) (pLM1110 (ref. 17), Addgene #168460) backbone DNA and purified assembly fragments were transformed into yeast using the high-efficiency lithium acetate method70. Transformants were plated on synthetic complete media lacking uracil or leucine (SC–Ura, SC–Leu) depending on the selectable marker (URA3 for HPRT1R segments 1–15 half-assembly, and LEU2 for HPRT1R segments 15–28 half-assembly and for HPRT1RnoCpG full assembly). Successful assemblies were screened by junction quantitative PCR (qPCR) on crude yeast genomic DNA (gDNA) prepared from 48 colonies from each assembly transformation. Crude yeast gDNA was prepared by performing three cycles of boiling in 20 mM NaOH at 98 °C for 3 min, followed by cooling at 4 °C for 1 min. Junction qPCRs were set up using an Echo 650 liquid handler (Labcyte) by dispensing 20 nl crude gDNA and 10 nl premixed junction primer pairs (50 µM) into a LightCycler 1536 Multiwell Plate (Roche 05358639001) containing 1 µl 1× LightCycler 1536 DNA Green mix (Roche 05573092001). qPCR reactions were performed using a LightCycler 1536 Instrument (Roche 05334276001) and successful assemblies were identified based on positive results for all junctions, defined as a having a Ct value lower than 30 (with exceptions for primer pairs determined to be consistently poor). Candidate assemblies were verified by next-generation sequencing. Libraries were prepared from 100 ng of DNA using the NEBNext Ultra II FS DNA Library Prep Kit for Illumina (NEB E7805L) with NEBNext Multiplex Oligos for Illumina (E7600S), according to the manufacturer’s protocol for FS DNA Library Prep Kit with Inputs ≤100 ng. Sequencing reactions were run on a NextSeq 500 system (Illumina SY-415-1001). Sequence-verified assemblons were recovered from yeast using the Zymoprep Yeast Miniprep I kit (Zymo Research D2001) and electroporated into TransforMax EPI300 Electrocompetent E. coli (Lucigen EC300150), recovered in LB + 5 mM MgCl2 at 30 °C for 1 h and then selected on LB + kanamycin agar plates. Bacteria colonies were screened by colony PCR for one or two assembly junctions to confirm that they contained the assemblon, then assemblon DNA was isolated from overnight cultures using ZR BAC DNA Miniprep kit (Zymo Research D4048) and verified by next-generation sequencing. eSwAP-In18 was used to combine the two HPRT1R half-assemblies. The sequence-verified assembly of segments 15–28 was purified from E. coli and digested with I-SceI and NotI to release the HPRT1R portion along with the LEU2 marker. This digested segment was transformed into yeast harbouring the assemblon with segments 1–15, along with a Cas9–guide RNA (gRNA) expression vector, pYTK-Cas9 (ref. 71), with a URA3-targeting gRNA. The Cas9-induced break in the URA3 marker was repaired with the HPRT1R-15–28-LEU2 segment using homology provided by the common segment 15 and common sequence downstream of the selection markers. eSwAP-In transformants were selected on SC–Leu and colonies were picked to screen by junction PCR using a subset of primers spanning the entire locus. Candidate clones were verified by next-generation sequencing and recovered into E. coli as previously described.
The HPRT1 locus was transplanted from its original assembly vector18 by restriction digestion of purified assemblon DNA with NotI and NruI to release the HPRT1 locus, followed by co-transformation of the digested locus (~1.5 μg) along with the new, linearized, pLM1110 assembly vector (~100 ng) and linker DNAs that included loxP and loxM sites flanked by 200 bp of homology to the assembly vector and HPRT1 locus (~50 ng each). Forty-eight colonies were picked following transformation and selection and crude yeast gDNA was screened by PCR using primers spanning the vector-HPRT1 junctions. Candidate clones were verified by next-generation sequencing and recovered into E. coli as described above.
Assemblons were recovered from TransforMax EPI300 E. coli for delivery to mouse ES cells. Cultures of 250 ml cultures were grown at 30 °C with shaking overnight in LB + kanamycin + 0.04% arabinose to induce copy number amplification of the assemblon BAC. DNA was purified using the NucleoBond XtraBAC kit (Takara Bio 740436.25) and stored at 4 °C for less than one week before delivery to mouse ES cells.
Integrating loci into the yeast genome
A landing pad containing a URA3 cassette flanked by loxM and loxP sites was installed at YKL162C-A21 in yeast strains harbouring either HPRT1 or HPRT1R assemblons. The landing pad was co-transformed, along with linker DNAs with terminal homologies to the yeast genomic locus and to the landing pad cassette (~200 ng each), into yeast as described above. Colonies were selected on SC–Ura plates, and 4 colonies were picked from each transformation and screened by PCR using primers spanning the genome–landing pad junctions. Landing pad integration was verified by Sanger sequencing of PCR products spanning the genome–landing pad junctions. The synthetic HPRT1 and HPRT1R loci were integrated by Cre-mediated recombination. A HIS3 plasmid expressing Cre-recombinase from a galactose-inducible promoter (pSH62 (ref. 72), Euroscarf P30120) was introduced by yeast transformation, single colonies were picked and grown to saturation in SC–His–Leu with raffinose, subcultured 1:100 in SC–His media with galactose, and plated on SC + 5-Fluoroorotic acid (5FOA) plates after 2 days of growth. 5FOA-resistant colonies were picked, screened by PCR using primers spanning the yeast genome–HPRT1 or HPRT1R junctions, and verified by next-generation whole-genome sequencing as described above. Engineered yeast strains are available upon request.
Sphis5 insertion and transcription factor knockouts
The His5 gene, including 5′ and 3′ untranslated regions, was cloned by PCR using Q5 high-fidelity DNA polymerase (New England Biolabs M0494L) from S. pombe genomic DNA. PCR primers were designed to add 40 bp of homology on each side for the desired target location in the synthetic HPRT1 or HRPT1R sequence, or in the yeast genome. Sphis5 PCR products were purified using the DNA Clean and Concentrator 5 kit (Zymo Research D4004) and transformed into HPRT1 or HPRT1R episome-harbouring yeast strains, as described above. Transformations were selected on SC–His–Leu plates and correct insertions were determined by PCR using a forward primer annealing in the in the predicted promoter regions within the HPRT1 or HPRT1R locus or yeast genome, outside of the homology arm, and a reverse primer annealing inside of the Sphis5 sequence.
Select transcription factor genes were knocked out of His+ yeast strains by cloning the URA3 expression cassette from pAV116 (Addgene #63183) using primers designed to add 40-bp homology arms targeting the genomic region upstream and downstream of the transcription factor coding sequence. URA3 PCR products were purified using the DNA Clean and Concentrator 5 kit (Zymo Research D4004) and transformed into His+ yeast strains as above. Transformations were selected on SC–Leu–Ura and correct knockouts were verified by PCR using two sets of primers spanning the URA3–genome junctions.
Yeast spot assays
Fitness of yeast strains following Sphis5 insertions and transcription factor knockouts was assessed by spot assay. Yeast strains were grown to saturation in selective media and diluted to OD600 of 1 in sterile water. Five tenfold serial dilutions were made of each strain, and 5 μl of each dilution was spotted on agar plates using a multichannel pipette. Plates were incubated at 37 °C for 2 days before imaging. 3-AT, a competitive inhibitor of the Sphis5 gene product, was used to better identify small magnitude changes in expression.
Mouse ES cell culture
C57BL6/6J × CAST/EiJ (BL6xCAST) ΔPiga mouse ES cells, which enable PIGA-based Big-IN genome rewriting, have been described previously17. Mouse ES cells were cultured in 80/20 medium, which consists of 80% 2i medium (1:1 mixture of Advanced DMEM/F12 (ThermoFisher 12634010) and Neurobasal-A (ThermoFisher 10888022) supplemented with 1% N2 Supplement (ThermoFisher 17502048), 2% B27 Supplement (ThermoFisher 17504044), 1% GlutaMAX (ThermoFisher 35050061), 1% penicillin-streptomycin (ThermoFisher 15140122), 0.1 mM 2-mercaptoethanol (Sigma M3148), 1,250 U ml−1 LIF (ESGRO ESG1107l), 3 μM CHIR99021 (R&D Systems 4423), and 1 μM PD0325901 (Sigma PZ0162)), and 20% mouse ES cell medium (KnockOut DMEM (ThermoFisher 10829018) supplemented with 15% FBS (BenchMark 100106), 0.1 mM 2-mercaptoethanol, 1% GlutaMAX, 1% MEM non-essential amino acids (ThermoFisher 11140050), 1% nucleosides (EMD Millipore ES-008-D), 1% penicillin-streptomycin, and 1,250 U ml−1 LIF). Mouse ES cells were maintained on plates coated with 0.1% gelatin (EMD Millipore ES-006-B) at 37 °C in a humidified incubator with 5% CO2. C57BL6/6J × CAST/EiJ (BL6xCAST) mouse ES cells were originally provided by D. Spector, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY. The BL6xCAST cell line was authenticated in next-generation capture-sequencing experiments, confirming cells as C57BL6/6J × CAST/EiJ hybrids on the basis of species-specific single-nucleotide polymorphisms. Cell lines were verified to be mycoplasma free prior to the study. There was no indication of contamination of any kind.
Integrating synthetic loci into mouse ES cells
Integration of synthetic loci was performed using the Big-IN method17. First, a landing pad, LP-PIGA2, containing a polycistronic cassette, pEF1 α-PuroR-P2A-PIGA-P2A-mScarlet-EF1αpA, for selection and counterselection and flanked by loxM and loxP sites, was modified with homology arms for targeting the landing pad to the mouse Hprt1 locus. Specifically, ~130-bp homology arms (amplified from a mouse Hprt1 BAC) flanked by gRNA sites for the Hprt1-targeting gRNAs (see below) and protospacer adjacent motifs were cloned flanking the lox sites using BsaI Golden Gate Assembly. LP-PIGA2 was delivered to BL6xCAST ΔPiga mouse ES cells, along with Cas9–gRNA-expression plasmids (pSpCas9(BB)-2A-GFP, Addgene #48138) expressing gRNAs that target sites flanking the Hprt1 locus, by nucleofection using the Neon Transfection System (ThermoFisher) as described17. One million cells were used per transfection with 5 μg of the landing pad plasmid and 2.5 μg each of Cas9–gRNA-expression plasmids. Cells were selected with 1 μg ml−1 puromycin starting day 1 post-transfection, with 6-thioguanine (Sigma-Aldrich A4660) starting day 7 post-transfection to select for the loss of Hprt1, and with 1 µM ganciclovir (Sigma PHR1593) to select against the landing pad plasmid backbone that contained a HSV1-ΔTK expression cassette. Candidate clones were picked on day 10, screened by qPCR using primers spanning the mouse genome–landing pad junctions and with primers for validating the loss of the endogenous Hprt1 gene and the absence of landing pad backbone or pSpCas9 plasmid integration. Mouse ES cell clones were further verified by next-generation baited Capture-seq17 that the Hprt1 locus was deleted and the landing pad was present on target. Genomic integration of a landing pad at Sox2 has been described20, replacing only the BL6 allele in the hybrid BL6xCAST cell line, leaving the CAST Sox2 allele intact. Engineered mouse ES cell lines are available upon request.
Delivery of the synthetic locus payloads was performed as described17 using the Amaxa 2b nucleofector (program A-23). In brief, 5 million cells were nucleofected with 5 μg pCAG-iCre (Addgene #89573) and 5 μg of assemblon DNA. Nucleofected mouse ES cells were treated with 10 µg ml−1 blasticidin for 2 days starting 1 day post-transfection to transiently select for the presence of the synthetic assemblons, and then with 2 nM proaerolysin for 2 days starting day 7 post-transfection to select for loss of PIGA in the landing pad cassette. Cells delivered with HPRT1 were also selected with HAT medium (ThermoFisher Scientific 21060017) starting day 7 post-transfection. Clones were picked on day 9 post-transfection, expanded, and screened first by qPCR aided by an Echo 550 liquid handler (Labcyte) as described20 using primers spanning the junctions between the mouse genome and HPRT1 or HPRT1R synthetic loci, and verified by Capture-seq17. For each locus integration we established two clonal cell lines from independent integration events.
Whole-genome sequencing and Capture-seq
Whole-genome sequencing and Capture-seq were performed as previously described17. Biotinylated bait DNA was generated by nick translation from purified BACs and plasmids of interest: the mouse Hprt1- and Sox2-containing BACs (RP23-412J16, RP23-274P9 respectively, BACPAC Resources Center), the synthetic HPRT1, HPRT1R, and HPRT1RnoCpG BACs, LP-PIGA2, pCAG-iCre and pSpCas9(BB)-2A-GFP.
Sequencing and initial data processing were performed according to as previously described17 with modifications. Illumina libraries were sequenced in paired-end mode on an Illumina NextSeq 500 operated at the Institute for Systems Genetics. All data were initially processed using a uniform mapping pipeline. Sequencing adapters were trimmed with Trimmomatic v0.39 (ref. 73). Whole-genome and Capture-seq reads were aligned using BWA v0.7.17 (ref. 74) to a reference genome (SacCer_April2011/sacCer3 or GRCm38/mm10), including unscaffolded contigs and alternate references, as well as independently to HPRT1 and HPRT1R custom references for relevant samples. PCR duplicates were marked using samblaster v0.1.24 (ref. 75). Generation of per base coverage depth tracks and quantification was performed using BEDOPS v2.4.35 (ref. 76). Data were visualized using the University of California, Santa Cruz Genome Browser. On-target, single-copy integrations are validated using DELLY77 call copy number variations, and bamintersect17 to identify unexpectedly mapping read pairs. Using these quality control steps, DELLY will identify duplications or deletions, and bamintersect will identify duplications based on read pairs mapping either between the end and the start of the synthetic locus (if duplicated in tandem) or between the synthetic locus and an unexpected genomic location (if duplicated by off-target integration). The sequencing processing pipeline is available at https://github.com/mauranolab/mapping.
ATAC-seq
For yeast, two independent clones for each strain were inoculated into 5 ml of SC–Leu (for assemblon strains) or YPD (for integration strains) for overnight culture at 30 °C. Saturated overnight cultures were diluted to an OD600 of 0.1 and cultured for 6 h at 30 °C, until OD600 reached ~0.6. Around 5 × 106 cells were taken from each culture, pelleted at 3,000g for 5 min, washed twice with 500 μl spheroplasting buffer (1.4 M sorbitol, 40 mM HEPES-KOH pH 7.5, 0.5 mM MgCl2), resuspended in 100 μl spheroplasting buffer with 0.2 U μl−1 zymolyase (Zymo Research E1004), then incubated for 30 min at 30 °C on a rotator. Spheroplasts were washed twice with 500 μl spheroplasting buffer then resuspended in 50 μl 1× TD buffer with TDE (Illumina 20034197). Tagmentation was performed for 30 min at 37 °C on a rotator and DNA was purified using the DNA Clean and Concentrator 5 kit (Zymo Research D4004). PCR was performed as previously described78 using 11 total cycles. The libraries were sequenced with 36-bp paired-end reads on a NextSeq 500 for ~1 million reads per sample.
For mouse ES cells, two independent cultures of each cell line were grown to medium confluency in 6-well plates. Cells were harvested by washing once with PBS, dissociated into single-cell suspension with TrypLE Express (ThermoFisher 12604013) and then neutralizing with equal volume mouse ES cell medium. Cells were counted and 50,000 were taken for tagmentation. Cells were pelleted at 500g for 5 min at 4 °C, washed with 50 μl cold PBS, resuspended in 50 μl cold ATAC lysis buffer (10 mM Tris-HCl, pH 7.4, 10 mM NaCl, 3 mM MgCl2, 0.1% IGEPAL CA-630), spun down at 500g for 10 mins at 4 °C, resuspended in 50 μl TDE mix, and incubated at 37 °C on rotator for 30 mins. DNA was purified using the DNA Clean and Concentrator 5 kit (Zymo Research D4004). PCR was performed as previously described78 using 10 total cycles. The libraries were sequenced with 36-bp paired-end reads for ~50 million reads per sample.
Illumina libraries were sequenced on an Illumina NextSeq 500 operated at the Institute for Systems Genetics. Sequencing adapters were trimmed with Trimmomatic v0.39 (ref. 73). Reads were aligned using bowtie2 v2.2.9 (ref. 79) to custom references in which the synthetic locus sequences were present on separate chromosomes or inserted at their specific integration sites in the SacCer_April2011/sacCer3 or GRCm38/mm10 genomes (produced using the reform tool; https://gencore.bio.nyu.edu/reform/). Coverage tracks were produced in bigWig format using bamCoverage (deepTools v3.5.0)80 with bin size 10 and smooth length 100, normalized using RPGC to an effective genome size of 12,000,000 for sacCer3 and 2652783500 for mm10, and visualized using IGV v2.12.3 (ref. 81). Peaks were called using macs2 v2.1.0 (ref. 82) with the parameters: --nomodel -f BAMPE --keep-dup all -g 1.2e7 (sacCer3)/1.87e9 (mm10). Relative coverage analysis was performed as described below.
RNA-seq
For yeast, the remaining culture that was not used for ATAC-seq was centrifuged at 3,000g for 5 min to pellet cells, washed once with water, pelleted again at 3,000g for 5 min, and cell pellets were frozen at −80 °C. Frozen pellets were resuspended in 200 μl lysis buffer (50 mM Tris-HCl pH 8, 100 mM NaCl) and lysed by disruption with an equal volume of acid washed glass beads, vortexing 10× 15 s. 300 μl lysis buffer was added and samples were mixed by inversion followed by a short centrifugation to collect all liquid in the tube. Supernatant (450 μl) was mixed with an equal volume of phenol:chloroform:isoamyl alcohol, vortexed for 1 min, and centrifuged at maximum speed for 5 min. 350 μl of the aqueous layer was then mixed with an equal volume of phenol:chloroform:isoamyl alcohol, vortexed for 1 min, and centrifuged at maximum speed for 5 min. RNA was precipitated from 300 μl of the aqueous phase by adding 30 μl of 3 M sodium acetate and 800 μl of cold 99.5% ethanol, briefly vortexing, and centrifuging at maximum speed for 10 min. The pellet was rinsed with 70% ethanol and dried at room temperature before dissolving in 100 μl of RNase-free DNase set (Qiagen 79254) and incubating at room temperature for 10 min to remove DNA. RNA was purified using the RNeasy Plus Mini kit (Qiagen 74136) and eluted in 30 μl RNase-free water. RNA-seq libraries were prepared from 1 μg total RNA using the QIAseq FastSelect -rRNA Yeast kit (Qiagen 334217) and QIAseq Stranded RNA Library kit (Qiagen 180743) according to the manufacturer’s protocol. The libraries were sequenced on a NextSeq 500 with 75 bp paired-end reads for ~45 million reads per sample.
For mouse ES cells, the remaining cells that were not used for ATAC-seq were pelleted at 500g for 5 min and RNA was isolated using Qiagen RNeasy Plus Mini kit, resuspending in 350 μl buffer RLT Plus + β-mercaptoethanol, with homogenization using QIAshredder columns (Qiagen 79654). RNA-seq libraries were prepared from 1 μg total RNA using QIAseq FastSelect -rRNA HMR (Qiagen 334386) and QIAseq Stranded RNA kits (Qiagen 180743) according to the manufacturer’s protocol. The libraries were sequenced with 75-bp paired-end reads for ~50 million reads per sample.
Illumina libraries were sequenced on an Illumina NextSeq 500 operated at the Institute for Systems Genetics. Sequencing adapters were trimmed with Trimmomatic v0.39 (ref. 73). STAR (v2.5.2a)83 was used to align reads, without providing a gene annotation file, to custom references in which the synthetic HPRT1 and HPRT1R sequences were present on separate chromosomes or inserted at their specific integration sites in the SacCer_April2011/sacCer3 or GRCm38/mm10 genomes (produced using the reform tool; https://gencore.bio.nyu.edu/reform/). Coverage tracks were produced in bigWig format using bamCoverage (deepTools v3.5.0)80 with bin size 10 and smooth length 100, filtering by strand, normalizing using TMM84, and visualized using IGV v2.12.3 (ref. 81). Relative coverage analysis was performed as described below.
CUT&RUN
For yeast, two independent colonies for each strain were inoculated into 5 ml of SC–Leu (for assemblon strains) or YPD (for integration strains) for overnight culture at 30 °C. Saturated overnight cultures were diluted to OD600 of 0.1 and cultured for ~6 h at 30 °C, until OD600 reached ~0.6. Cells were pelleted at 3,000g for 5 min, washed twice with water, and resuspended in spheroplasting buffer (1.4 M sorbitol, 40 mM HEPES-KOH pH 7.5, 0.5 mM MgCl2, 0.5 mM 2-mercaptoethanol). Spheroplasting was performed by adding 0.125 U μl−1 Zymolyase (Zymo Research E1004) and incubating at 37 °C for 45 min on a rotator. Nuclei were prepared as previously described85. Resuspended nuclei were split into aliquots of ~108 nuclei each and snap frozen in liquid nitrogen.
For mouse ES cells, two independent cultures for each engineered cell line cells were harvested from tissue culture dishes using TrypLE Express (ThermoFisher 12604013), dissociated into single-cell suspension, and quenched with mouse ES cell medium. Crosslinking was performed by adding formaldehyde to a final concentration of 0.1% (v/v) and incubating at room temperature for 5 min with occasional mixing by inversion. Crosslinking was stopped by quenching with 125 mM glycine and incubating at room temperature for 5 min with occasional mixing by inversion. DMSO was added to a final concentration of 10% (v/v) and cells were frozen in aliquots of ~106 cells.
Isolated yeast nuclei (~108 per sample) or crosslinked mouse ES cells (~106 per sample) were thawed and processed for CUT&RUN using the CUTANA ChIC/CUT&RUN kit (EpiCypher 14-1048) according to the manufacturer’s protocol. Antibodies were all used at 0.5 μg: rabbit IgG negative control (EpiCypher 13-0042), H3K4me3 (EpiCypher 13-0041), H3K27ac (EpiCypher 13-0045), H3K27me3 (Active Motif 39055, RRID: AB_2561020), RNAP2 (Santa Cruz Biotechnology sc-56767). Sequencing libraries were prepared using the NEBNext Ultra II DNA Library Prep Kit for Illumina (New England Biolabs E7645L) and sequenced with 75 bp paired-end reads for ~15 M reads for H3K4me3 and Pol II samples, and ~20 M reads for H3K27ac and H3K27me3 samples.
Illumina libraries were sequenced on an Illumina NextSeq 500 operated at the Institute for Systems Genetics. Sequencing adapters were trimmed with Trimmomatic v0.39 (ref. 73). Reads were aligned using bowtie2 v2.2.9 (ref. 79) to custom references in which the synthetic HPRT1 and HPRT1R sequences were present on separate chromosomes or inserted at their specific integration sites in the SacCer_April2011/sacCer3 or GRCm38/mm10 genomes (produced using the reform tool; https://gencore.bio.nyu.edu/reform/). Coverage tracks were produced in bigWig format using bamCoverage (deepTools v3.5.0)80 with bin size 10 and smooth length 100, normalized using RPGC to an effective genome size of 12,000,000 for sacCer3 and 2,652,783,500 for mm10, and visualized using IGV v2.12.3 (ref. 81). Peaks were called using macs2 v2.1.0 (ref. 82) with the parameters: --nomodel -f BAMPE --keep-dup all -g 1.2e7 (sacCer3)/1.87e9 (mm10). Relative coverage analysis was performed as described below.
CAGE-seq
RNA was isolated as described above for RNA-seq, using two replicate colonies for each yeast strain. CAGE libraries were prepared as previously described24,starting with 5 μg RNA, with the following modifications. SuperScript IV Reverse Transcriptase (Invitrogen 18090010) was used for the reverse transcription step. AMPure XP beads (Beckman Coulter A63881) were used for all bead cleanup steps. We also used custom-made linker and primer oligonucleotides so that linkers are universal to all samples and primers contain sample-specific barcodes. Libraries were amplified using universal forward and reverse primers with 20 cycles of PCR. Libraries were sequenced on with 75 bp paired-end reads for ~22 million reads per sample.
Illumina libraries were sequenced on an Illumina NextSeq 500 operated at the Institute for Systems Genetics. Sequencing adapters were trimmed with Trimmomatic v0.39 (ref. 73). The 5′ reads only were aligned using bowtie2 v2.2.9 (ref. 79) to custom references in which the synthetic HPRT1 and HPRT1R sequences were present on separate chromosomes or inserted at their specific integration sites in the SacCer_April2011/sacCer3 or GRCm38/mm10 genomes (produced using the reform tool; https://gencore.bio.nyu.edu/reform/). Coverage tracks were produced in bigWig format using bamCoverage (deepTools v3.5.0)80 with bin size 1, filtering by strand, normalized using RPGC to an effective genome size of 12,000,000, and visualized using IGV v2.12.3 (ref. 81). Peaks were called using macs2 v2.1.0 (ref. 82) with the parameters: --nomodel -f BAM --keep-dup all -g 1.2e7.
Locus copy number estimation
For copy number estimation in yeast strains, coverage depth was calculated from whole-genome sequencing data for the synthetic HPRT1 and HPRT1R loci as well as the entire yeast genome (excluding chrM) using samtools v1.9 depth86, and the calculated depth of the synthetic loci was divided by the genome average.
Sequencing coverage analysis
Relative coverage analysis was performed for yeast ATAC-seq, RNA-seq, and CUT&RUN experiments. Average coverage depth was calculated over the synthetic HPRT1 and HPRT1R loci, 100-kb sliding windows of yeast genome using samtools v1.9 bedcov86, which reports the total read base count (the sum of per base read depths) per specified region, and then dividing the total read base count by the region size − 100,735 bp for the HPRT1/HPRT1R loci or 100,000 bp for the 100-kb windows. Coverage was corrected for estimated copy numbers of the HPRT1 and HPRT1R episomes. The yeast genome was split into 100-kb sliding windows with 10-kb step size using bedtools v2.29.2 makewindows87. The average of the 100-kb windows was then calculated. The average coverage depth over the synthetic loci was then divided by the relevant genome average to determine relative coverage depth in each context (that is, HPRT1 average coverage/average coverage of yeast 100-kb windows = relative coverage of HPRT1 compared to the yeast genome). For peak analysis, total peaks were counted across the HPRT1 and HPRT1R loci, or averaged over the yeast genome 100-kb windows.
For mouse genome RNA-seq read analysis, the mouse genome was split into 100-kb sliding windows with 10-kb step size using bedtools v2.29.2 makewindows87. The windows were then filtered to exclude ENCODE blacklist regions88, centromeres, telomeres, and annotated transcripts based on Gencode comprehensive gene annotation, release M10 (GRCm38.p4). RNA-seq reads were counted for the synthetic loci and for the 100-kb genomic windows using samtools v1.9 (ref. 86) view with arguments -c -F 2308 -L (reference bed file).
Replicate correlation
Correlation between sequencing assay replicates was assessed using deepTools v3.5.0 (ref. 80) multiBigwigSummary to first calculate average bigWig scores for each dataset across the mouse genome in 10-kb bins, and across the yeast genome in 100-bp bins. Biological and technical replicates were compared using plotCorrelation with the following arguments: --corMethod pearson --whatToPlot scatterplot --skipZeros --removeOutliers --log1p.
Metaplots analysis
TSSs were defined as the 5′ coordinate of the experimentally identified CAGE-seq peaks. Metaplots were produced using deepTools v3.5.0 (ref. 80) computeMatrix and plotProfile, with argument --plotType se. Matrices were computed for ATAC-seq and H3K4me3 CUT&RUN signals and profiles were plotted for TSSs across the HPRT1 and HPRT1R loci and across the rest of the yeast genome.
Motif analysis
Putative promoter regions in the synthetic HPRT1 and HPRT1R loci were defined as 200 bp upstream and 100 bp downstream of the TSSs identified based on CAGE-seq peaks (above). Motif discovery was performed on the putative promoter regions, ATAC-seq peaks, and ATAC-seq peaks that intersect with putative promoters, identified with bedtools v2.29.2 intersect87. Regions of interest were combined from HPRT1 and HPRT1R for motif analysis using MEME v4.102 (ref. 25) with a maximum motif width of 10 bp. This width was determined empirically by observing that increasing widths did not result in the predicting of any more informative motifs. Tomtom27 was performed to scan the identified motifs for matches to motifs in the YEASTRACT database65. GOmo89 was performed to identify gene ontology terms linked to gene promoters containing the identified motifs.
Public sequencing data
We obtained UCSC browser data for CpG islands90,91, as well as the following ENCODE data92. DNase-seq from ES-E14 mouse embryonic stem cells, ENCSR000CMW93. Chromatin immunoprecipitation with sequencing (ChIP-seq) from ES-Bruce mouse embryonic stem cells, ENCSR000CBG, ENCSR000CDE, ENCSR000CFN94, ENCSR000CCC. RNA-seq from ES-E14 mouse embryonic stem cells, ENCSR000CWC, ENCSR000CWC. ATAC-seq data from embryonic day (E)11.5 mouse embryonic tissue, ENCSR282YTE, ENCFF936VGM28. ChIP-seq data from E11.5 mouse embryonic tissue, ENCSR427OZM, ENCFF952ZWD, ENCSR531RZS, ENCFF033UPR, ENCSR240OUM, ENCFF179QWF28. DNase-seq from H1 human ES cells ENCSR000EJN, ChIP-seq from H1 human ES cells ENCSR443YAS, ENCSR880SUY, ENCSR928HYM, RNA-seq from H1 human ES cells ENCSR000COU95. Long RNA-seq data from H1 human ES cells, ENCSR000COU, ENCFF563OKS, ENCFF501KFP, ENCFF407PJY, ENCFF761BKF2.
We obtained public sequencing data for yeast from the following datasets (Gene Expression Omnibus (GEO) accession numbers): ATAC-seq (GSM6139041), H3K4me3 ChIP-seq (GSM3193266), RNA-seq (GSM5702033) and yeast CAGE-seq (ref. 96).
DNA reagents
Sequences and identifiers, where applicable, for all DNA reagents used in this study are available as supplementary material, including all oligonucleotides, synthetic DNA segments, plasmids, landing pads, homology arms and yeast strains.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data generated in this study are available in the NCBI GEO database under accession GSE252482.
References
The ENCODE Project Consortium. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Pertea, M. The human transcriptome: an unfinished story. Genes 3, 344–360 (2012).
Struhl, K. Transcriptional noise and the fidelity of initiation by RNA polymerase II. Nat. Struct. Mol. Biol. 14, 103–105 (2007).
Clark, M. B. et al. The reality of pervasive transcription. PLoS Biol. 9, e1000625 (2011). discussion e1001102.
van Bakel, H., Nislow, C., Blencowe, B. J. & Hughes, T. R. Response to “The reality of pervasive transcription”. PLoS Biol. 9, e1001102 (2011).
David, L. et al. A high-resolution map of transcription in the yeast genome. Proc. Natl Acad. Sci. USA 103, 5320–5325 (2006).
Chen, W. H., Wei, W. & Lercher, M. J. Minimal regulatory spaces in yeast genomes. BMC Genomics 12, 320 (2011).
Gherman, A., Wang, R. & Avramopoulos, D. Orientation, distance, regulation and function of neighbouring genes. Hum. Genomics 3, 143–156 (2009).
Eddy, S. R. The ENCODE project: missteps overshadowing a success. Curr. Biol. 23, R259–R261 (2013).
Zhang, W., Mitchell, L. A., Bader, J. S. & Boeke, J. D. Synthetic genomes. Annu. Rev. Biochem. 89, 77–101 (2020).
Venter, J. C., Glass, J. I., Hutchison, C. A. 3rd & Vashee, S. Synthetic chromosomes, genomes, viruses, and cells. Cell 185, 2708–2724 (2022).
Laurent, J. M. et al. Big DNA as a tool to dissect an age-related macular degeneration-associated haplotype. Precis. Clin. Med. 2, 1–7 (2019).
Brosh, R. et al. A versatile platform for locus-scale genome rewriting and verification. Proc. Natl Acad. Sci. USA 118, e2023952118 (2021).
Mitchell, L. A. et al. De novo assembly and delivery to mouse cells of a 101 kb functional human gene. Genetics 218, iyab038 (2021).
Pinglay, S. et al. Synthetic regulatory reconstitution reveals principles of mammalian Hox cluster regulation. Science 377, eabk2820 (2022).
Brosh, R. et al. Synthetic regulatory genomics uncovers enhancer context dependence at the Sox2 locus. Mol. Cell 83, 1140–1152.e1147 (2023).
Agmon, N. et al. Yeast golden gate (yGG) for the efficient assembly of S. cerevisiae transcription units. ACS Synth. Biol. 4, 853–859 (2015).
Szybalska, E. H. & Szybalski, W. Genetics of human cell line. IV. DNA-mediated heritable transformation of a biochemical trait. Proc. Natl Acad. Sci. USA 48, 2026–2034 (1962).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Murata, M. et al. Detecting expressed genes using CAGE. Methods Mol. Biol. 1164, 67–85 (2014).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Zhang, Z. & Dietrich, F. S. Mapping of transcription start sites in Saccharomyces cerevisiae using 5′ SAGE. Nucleic Acids Res. 33, 2838–2851 (2005).
Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Gorkin, D. U. et al. An atlas of dynamic chromatin landscapes in mouse fetal development. Nature 583, 744–751 (2020).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Ku, M. et al. Genomewide analysis of PRC1 and PRC2 occupancy identifies two classes of bivalent domains. PLoS Genet. 4, e1000242 (2008).
Mendenhall, E. M. et al. GC-rich sequence elements recruit PRC2 in mammalian ES cells. PLoS Genet. 6, e1001244 (2010).
Lynch, M. D. et al. An interspecies analysis reveals a key role for unmethylated CpG dinucleotides in vertebrate Polycomb complex recruitment. EMBO J. 31, 317–329 (2012).
Jermann, P., Hoerner, L., Burger, L. & Schubeler, D. Short sequences can efficiently recruit histone H3 lysine 27 trimethylation in the absence of enhancer activity and DNA methylation. Proc. Natl Acad. Sci. USA 111, E3415–E3421 (2014).
Wachter, E. et al. Synthetic CpG islands reveal DNA sequence determinants of chromatin structure. eLife 3, e03397 (2014).
Li, H. et al. Polycomb-like proteins link the PRC2 complex to CpG islands. Nature 549, 287–291 (2017).
Xu, Z. et al. Bidirectional promoters generate pervasive transcription in yeast. Nature 457, 1033–1037 (2009).
Neil, H. et al. Widespread bidirectional promoters are the major source of cryptic transcripts in yeast. Nature 457, 1038–1042 (2009).
Tisseur, M., Kwapisz, M. & Morillon, A. Pervasive transcription—lessons from yeast. Biochimie 93, 1889–1896 (2011).
Lu, Z. & Lin, Z. Pervasive and dynamic transcription initiation in Saccharomyces cerevisiae. Genome Res. 29, 1198–1210 (2019).
Gvozdenov, Z., Barcutean, Z. & Struhl, K. Functional analysis of a random-sequence chromosome reveals a high level and the molecular nature of transcriptional noise in yeast cells. Mol. Cell 83, 1786–1797.e1785 (2023).
Zhou, J. et al. Exogenous artificial DNA forms chromatin structure with active transcription in yeast. Sci. China Life Sci. 65, 851–860 (2022).
Luthra, I. et al. Regulatory activity is the default DNA state in eukaryotes. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-024-01235-4 (2024).
Chapard, C. et al. Exogenous chromosomes reveal how sequence composition drives chromatin assembly, activity, folding and compartmentalization. Preprint at bioRxiv https://doi.org/10.1101/2022.12.21.520625 (2023).
Kordis, D. & Gubensek, F. Horizontal SINE transfer between vertebrate classes. Nat. Genet. 10, 131–132 (1995).
Pace, J. K. 2nd, Gilbert, C., Clark, M. S. & Feschotte, C. Repeated horizontal transfer of a DNA transposon in mammals and other tetrapods. Proc. Natl Acad. Sci. USA 105, 17023–17028 (2008).
Husnik, F. & McCutcheon, J. P. Functional horizontal gene transfer from bacteria to eukaryotes. Nat. Rev. Microbiol. 16, 67–79 (2018).
Kambayashi, C. et al. Geography-dependent horizontal gene transfer from vertebrate predators to their prey. Mol. Biol. Evol. 39, msac052 (2022).
McLysaght, A. & Hurst, L. D. Open questions in the study of de novo genes: what, how and why. Nat. Rev. Genet. 17, 567–578 (2016).
Kudla, G., Lipinski, L., Caffin, F., Helwak, A. & Zylicz, M. High guanine and cytosine content increases mRNA levels in mammalian cells. PLoS Biol. 4, e180 (2006).
Neymotin, B., Ettorre, V. & Gresham, D. Multiple transcript properties related to translation affect mRNA degradation rates in Saccharomyces cerevisiae. G3 6, 3475–3483 (2016).
Courel, M. et al. GC content shapes mRNA storage and decay in human cells. eLife 8, e49708 (2019).
Vakirlis, N. et al. A molecular portrait of de novo genes in yeasts. Mol. Biol. Evol. 35, 631–645 (2018).
Schlotterer, C. Genes from scratch—the evolutionary fate of de novo genes. Trends Genet. 31, 215–219 (2015).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Zhao, Z. & Zhang, F. Sequence context analysis in the mouse genome: single nucleotide polymorphisms and CpG island sequences. Genomics 87, 68–74 (2006).
Galupa, R. et al. Enhancer architecture and chromatin accessibility constrain phenotypic space during Drosophila development. Dev. Cell 58, 51–62 e54 (2023).
Pich, O. et al. Somatic and germline mutation periodicity follow the orientation of the DNA minor groove around nucleosomes. Cell 175, 1074–1087.e1018 (2018).
Hon, C. C. et al. An atlas of human long non-coding RNAs with accurate 5′ ends. Nature 543, 199–204 (2017).
Iyer, M. K. et al. The landscape of long noncoding RNAs in the human transcriptome. Nat. Genet. 47, 199–208 (2015).
Ahmad, K. & Henikoff, S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell 9, 1191–1200 (2002).
Rando, O. J. & Ahmad, K. Rules and regulation in the primary structure of chromatin. Curr. Opin. Cell Biol. 19, 250–256 (2007).
Truong, D. M. & Boeke, J. D. Resetting the yeast epigenome with human nucleosomes. Cell 171, 1508–1519.e1513 (2017).
Lazar-Stefanita, L., Haase, M. A. B. & Boeke, J. D. Humanized nucleosomes reshape replication initiation and rDNA/nucleolar integrity in yeast. Preprint at bioRxiv https://doi.org/10.1101/2023.05.06.539710 (2023).
Haase, M. A. B. et al. Human macroH2A1 drives nucleosome dephasing and genome instability in histone-humanized yeast. Preprint at bioRxiv https://doi.org/10.1101/2023.05.06.538725 (2023).
Monteiro, P. T. et al. YEASTRACT+: a portal for cross-species comparative genomics of transcription regulation in yeasts. Nucleic Acids Res. 48, D642–D649 (2020).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Castro-Mondragon, J. A. et al. JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 50, D165–D173 (2022).
Madeira, F. et al. Search and sequence analysis tools services from EMBL–EBI in 2022. Nucleic Acids Res. 50, W276–W279 (2022).
German, S., Mitchell, L. A., Vela Gartner, A., Fenyö, D. & Boeke, J. D. MenDEL: PCR primer design as constrained optimization process. Preprint at bioRxiv https://doi.org/10.1101/2022.06.26.496474 (2022).
Gietz, R. D. & Schiestl, R. H. Large-scale high-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 38–41 (2007).
Zhao, Y. et al. CREEPY: CRISPR-mediated editing of synthetic episomes in yeast. Nucleic Acids Res. 51, e72 (2023).
Gueldener, U., Heinisch, J., Koehler, G. J., Voss, D. & Hegemann, J. H. A second set of loxP marker cassettes for Cre-mediated multiple gene knockouts in budding yeast. Nucleic Acids Res. 30, e23 (2002).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Faust, G. G. & Hall, I. M. SAMBLASTER: fast duplicate marking and structural variant read extraction. Bioinformatics 30, 2503–2505 (2014).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21–29 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–165 (2016).
Robinson, J. T. et al. Integrative Genomics Viewer. Nat. Biotechnol. 29, 24–26 (2011).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Orsi, G. A., Kasinathan, S., Zentner, G. E., Henikoff, S. & Ahmad, K. Mapping regulatory factors by immunoprecipitation from native chromatin. Curr. Protoc. Mol. Biol. 110, 21–25 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Buske, F. A., Kundaje, A. & Boyle, A. P. The ENCODE Blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 9354 (2019).
Buske, F. A., Boden, M., Bauer, D. C. & Bailey, T. L. Assigning roles to DNA regulatory motifs using comparative genomics. Bioinformatics 26, 860–866 (2010).
Gardiner-Garden, M. & Frommer, M. CpG islands in vertebrate genomes. J. Mol. Biol. 196, 261–282 (1987).
Rhead, B. et al. The UCSC Genome Browser database: update 2010. Nucleic Acids Res. 38, D613–D619 (2010).
Luo, Y. et al. New developments on the Encyclopedia of DNA Elements (ENCODE) data portal. Nucleic Acids Res. 48, D882–D889 (2020).
Sethi, A. et al. Supervised enhancer prediction with epigenetic pattern recognition and targeted validation. Nat. Methods 17, 807–814 (2020).
He, Y. et al. Spatiotemporal DNA methylome dynamics of the developing mouse fetus. Nature 583, 752–759 (2020).
Lee, D., Zhang, J., Liu, J. & Gerstein, M. Epigenome-based splicing prediction using a recurrent neural network. PLoS Comput. Biol. 16, e1008006 (2020).
McMillan, J., Lu, Z., Rodriguez, J. S., Ahn, T. H. & Lin, Z. YeasTSS: an integrative web database of yeast transcription start sites. Database 2019, baz048 (2019).
Acknowledgements
This work was supported by NIH/NHGRI grant 1RM1HG009491. The authors thank G. Ellis for her help with BAC DNA preparation, cell culture work and next-generation sequencing and A. Chess for helpful early discussions about the behaviour of random sequences.
Author information
Authors and Affiliations
Contributions
B.R.C. and J.D.B. conceptualized the study. R.B. and H.J.A. produced some cell lines and performed quality control. H.J.A. and M.T.M. ran next-generation sequencing assays and initial processing of the resulting data. B.R.C. built all synthetic constructs in yeast, produced some cell lines, performed all assays in yeast and mouse ES cells, performed final sequencing data processing, analysis and visualization, and created figures. B.R.C., J.D.B., and R.B. wrote the manuscript. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.D.B. is a founder and director of CDI Labs Inc., a founder of and consultant to Neochromosome Inc, a founder, scientific advisory board member of and consultant to ReOpen Diagnostics LLC, and serves or served on the scientific advisory board of the following: Sangamo Inc., Logomix Inc., Modern Meadow Inc., Rome Therapeutics Inc., Sample6 Inc., Tessera Therapeutics Inc. and the Wyss Institute, unrelated to this work. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Sean Eddy, Yo Suzuki, Martin Taylor and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Synthetic locus assembly and cell line engineering.
a,b, Landing pad installed on the X chromosome (a) or chromosome 3 (b) of the mouse genome. Presence of the landing pad was verified by Capture-seq (black track) on DNA from mESCs, as was the deletion of the endogenous Hprt locus, or one Sox2 locus (grey track), prior to synthetic locus delivery. Components of the landing pad are indicated above the sequencing track. c, Strategy for integrating synthetic loci into the yeast (Sc) genome. A landing pad containing the URA3 selectable/counterselectable marker cassette and flanked by heterotypic lox sites was integrated at YKL162C-A on chrXI by integrative homologous recombination in strains already harboring an episomal HPRT1 or HPRT1R assemblon. The synthetic loci were then recombined into this locus using transient Cre expression, overwriting the URA3 landing pad. d, Strategy for integrating synthetic loci into the mouse (Mm) genome. A landing pad containing a selectable/counterselectable cassette flanked by heterotypic lox sites was installed on chrX or chr3, overwriting the endogenous Hprt or one Sox2 allele, respectively. The synthetic loci were then delivered to these loci using Cre-mediated cassette exchange, overwriting the landing pad. e, Assembly of HPRT1R in two parts, the first containing segments 1-15 and the second segments 15-28. An example junction-qPCR verification is shown, reporting Ct values for qPCR reactions using primers for the indicated junctions (i.e., junction 1/2 is the junction between segments 1 and 2) performed on whole yeast DNA following assembly transformation and selection. The last column represents a positive control reaction detecting a yeast genomic marker. Sequencing coverage plots for the two half-assemblies from yeast whole genome sequencing is shown below. f, The eSwAP-In18 (extrachromosomal Switching Auxotrophies for Progressive Integration) strategy for combining the two HPRT1R half-assemblies. Segments 15-28 and the LEU2 marker are combined with segments 1-15 and the rest of the vector backbone through homologous recombination using the common segment 15 and common sequence downstream of the selection markers as homology arms to promote recombination. Positive recombinants are selected as Leu+ and 5-FOAr (Ura–). g, Estimated copy number of HPRT1 and HPRT1R in yeast when present episomally (Epi) or integrated (chrXI). Each data point represents the ratio of average whole genome sequencing coverage depth over the synthetic locus divided by the average coverage depth over the yeast genome for independently-sequenced yeast clones.
Extended Data Fig. 2 Yeast sequencing assay replicate tracks.
Sequencing tracks for ATAC-seq (a,b), H3K4me3 CUT&RUN (c,d), RNA-seq (e,f), and CAGE-seq (g,h) at HPRT1 and HPRT1R in yeast. Tracks are shown for episomal (Epi, a,c,e,g) and chromosomally integrated (chrXI, b,d,f,h) synthetic loci in two biological replicates for each strain. The HPRT1 coding sequence is indicated for the HPRT1 locus, and the relative position corresponding to the reversed coding sequence is indicated for the HPRT1R locus. The synthetic locus region is shaded for chromosomally integrated loci, and ~50 kb of flanking yeast genome is included, with annotated yeast genes indicated. RNA-seq (e,f) and CAGE-seq (g, h) tracks are stranded, displayed with reverse strand reads inverted and below forward strand reads. For CAGE-seq (g,h), “(Zoom)” tracks show the same data as the tracks above for each strain with a smaller y-axis scale to better visualize smaller peaks.
Extended Data Fig. 3 Yeast sequencing assay replicate correlates.
Scatterplots showing correlation between ATAC-seq (a), H3K4me3 CUT&RUN (b), RNA-seq (c), and CAGE-seq (d) signal (bigWig scores) over 100 bp windows for the combined synthetic locus and yeast genome (Sc + HPRT1/HPRT1R) and for the synthetic locus only (HPRT1/HPRT1R-only). Both replicates for each of the episomal (Epi) and chromosomally integrated (chrXI) synthetic loci are compared, and Pearson’s correlation (p) between each replicate is reported.
Extended Data Fig. 4 Yeast sequencing assays supplement.
a, Snapshot of a region on S. cerevisiae chromosome X showing sequencing data for ATAC-seq, H3K4me3 CUT&RUN, RNA-seq, and CAGE-seq from this study (purple) and data for ATAC-seq, H3K4me3 ChIP-seq, RNA-seq, and CAGE-seq from publicly available, published datasets (grey). Data from this study is from one replicate yeast strain with HPRT1 integrated on chrXI. GEO accession numbers for the public data: ATAC-seq GSM6139041, H3K4me3 ChIP-seq GSM3193266, RNA-seq GSM5702033, yeast CAGE-seq96. b, Sequencing tracks for ATAC-seq, H3K4me3 CUT&RUN, RNA-seq, and CAGE-seq for the region in the yeast genome just upstream of the HPRT1/HPRT1R integration site. Annotated yeast genes are shown below. Data is from one replicate yeast strain with HPRT1 integrated on chrXI. c, Metaplots of ATAC-seq and H3K4me3 CUT&RUN signal at the transcription start site (TSS, defined by experimental CAGE-seq peaks) +/− 0.5 kb for synthetic HPRT1 and HPRT1R loci integrated into the yeast genome on chrXI. Shaded region shows standard error. d,f,h, Relative coverage depth for ATAC-seq (d), H3K4me3 CUT&RUN (f), and RNA-seq (h) of the synthetic loci as episomes (Epi) and chromosomally integrated (chrXI) compared to the genome average. For episomes, coverage was corrected for the estimated copy number of the episomes. e,g,i, Peak counts per 100 kb for ATAC-seq (e), H3K4me3 CUT&RUN (g), and CAGE-seq (i) for the synthetic loci as episomes (Epi) and chromosomally integrated (chrXI) as well as the genome average. Bars show mean of biological replicates +/- SD. * p < 0.05, ** p < 0.01, two-tailed paired t-test comparing HPRT1/HPRT1R to the yeast genome average.
Extended Data Fig. 5 Sequencing motifs analysis for HPRT1 and HPRT1R in yeast.
a-c, The top 10 motifs identified across both synthetic HPRT1 and HPRT1R loci, within predicted promoter regions, defined as 200 bp upstream and 100 bp downstream of the experimentally identified CAGE-seq peaks (a), ATAC-seq peaks (b), and ATAC-seq peaks that overlap with predicted promoters (c). d, The only two motifs identified genome-wide in ATAC-seq peaks overlapping with CAGE-predicted promoters. Motifs were identified using MEME with a max width of 10 bp. For each set of motifs, the number of identified sites is reported, as well as the total number of sites analyzed, the E-value, and predicted transcription factor binding sites (TFBSs), identified using Tomtom in the MEME suite.
Extended Data Fig. 6 Insertion and characterization of Sphis5 at transcription start sites in synthetic HPRT1 and HPRT1R loci.
a, Identification of Sphis5 insertion sites based on experimental RNA-seq and CAGE-seq. Sequencing tracks for HPRT1 (top, blue) and HPRT1R (bottom, pink) are shown with forward reads on the top and reverse reads underneath and inverted. Sphis5 insertion sites are indicated with black dashed lines across the sequencing tracks. The precise insertion position, in base-pairs, and direction of transcription is indicated below the sequencing tracks. b, Example strategy for insertion of the Sphis5 coding sequence at a third experimentally-identified transcription start site. RNA-seq and CAGE-seq sequencing tracks are shown, as well as CAGE-seq peaks. The Sphis5 coding sequence (green arrow) is inserted with the 5’UTR at the 5’ boundary of the CAGE-seq peak. c, PCR verification of on-target Sphis5 insertion. PCRs were performed using a forward primer outside of the Sphis5 coding sequence and a reverse primer inside the Sphis5 coding sequence (as indicated by red arrows in b) for each insertion position. PCRs were performed on the parental yeast strain with just the HPRT1 or HPRT1R episome (Epi, top panel) and on the derivative clones with Sphis5 inserted at the indicated position, in base-pairs (bottom panel). d, Spot assays of parental HPRT1 and HPRT1R episome-containing yeast strains, and their derivative Sphis5 insertion strains, with the position of the Sphis5 insertion is indicated in base-pairs. Yeast were spotted on YPD, SC–Leu, and SC–Leu–His with 3-amino-1,2,4-triazole (3-AT), a competitive inhibitor of the Sphis5 gene product, added at the indicated concentrations. e, Spot assays of the HPRT1 strain with Sphis5 inserted at 13493 bp and derivatives in which transcription factors were knocked out by URA3 integration. Yeast were spotted on SC–Leu–Ura and SC–Leu–His with 3-AT added at the indicated concentrations. f, Insertion of the Sphis5 coding sequence into the yeast genome at the YKL162C-A locus, not adjacent to a transcription start site, as a negative control. g, PCR verification of on-target Sphis5 insertion into the genome using a primer pair as indicated by red arrows in f. h, Spot assays of the WT parental yeast strain, that strain with Sphis5 inserted into the genome, with the HPRT1 episome, and with Sphis5 inserted into the HPRT1 episome at 13493 bp. Yeast were spotted on YPD, SC–Leu, and SC–His. Residual background growth of the strains with Sphis5 inserted into the genome and in cells with the HPRT1 episome (lacking Sphis5) reflects the fact that yeast cells contain large stores of vacuolar amino acids which allow for a limited number of cell divisions on selective medium; the slightly higher growth in the former presumably reflects exceedingly low levels of Sphis5 transcription.
Extended Data Fig. 7 HPRT1, HPRT1R in mESCs replicate tracks.
Sequencing tracks for ATAC-seq, H3K4me3, RNAP2, H3K27ac, H3K27me3, and RNA-seq at HPRT1 integrated on chrX (a, b), and at HPRT1R integrated on chrX (c, d) and on chr3 (e, f), in mESCs. Two biological replicates (Rep. 1, Rep. 2) are shown for two clones (denoted (1) and (2)) derived from independent integrations. The synthetic locus region is shaded, and the HPRT1 coding sequence is indicated in a and b. ~50 kb of flanking mouse genome is included for each position, with annotated mouse genes indicated.
Extended Data Fig. 8 HPRT1, HPRT1R in mESCs replicate correlates.
Scatterplots showing correlation between sequencing assays for HPRT1 on chrX (a), HPRT1R on chrX (b), HPRT1R on chr3 (c), HPRT1RnoCpG on chrX (d), and HPRT1RnoCpG on chr3 (e). Plots compare signal, as bigWig scores, over 10 kb windows for the combined synthetic locus and mouse genome, for ATAC-seq, H3K4me3, RNAP2, H3K27ac, and H3K27me3 CUT&RUN, and RNA-seq. Both replicates (Rep. 1, Rep. 2) for each of the independent clonal isolates (Clone 1, Clone 2) are compared, and Pearson’s correlation (p) between each replicate is reported.
Extended Data Fig. 9 mESC sequencing assays compared to public genomic data.
a, Snapshot of a region on M. musculus chromosome 3 showing sequencing data for ATAC-seq/DNase-seq, H3K4me3 ChIP-seq, H3K27ac ChIP-seq, H3K27me3 ChIP-seq, RNAP2 ChIP-seq, and RNA-seq, from this study (purple) and from publicly available, published datasets (grey). Data from this study is from one replicate mESC clone with HPRT1 integrated on chrX. b, ENCODE data for DNase-seq, H3K4me3, RNAP2, H3K27ac, and H3K27me3 ChIP-seq, and RNA-seq from mouse embryonic stem cells. The browser shot shows the region on chrX used for integration of synthetic loci, including the native mouse Hprt gene. Predicted CpG islands are shown below. c, Data from this study for a clone in which HPRT1R in integrated on chromosome 3 (HPRT1R chr3 (1)) and so has an intact Hprt locus. d, ENCODE data for DNase-seq, H3K4me3, H3K27ac, and H3K27me3 ChiP-seq, and RNA-seq from the H1 human embryonic stem cell line. Browser shot is from the hg38 genome and shows the region on human chrX containing the native HPRT1 gene. Predicted CpG islands are shown below. The region that was cloned as the synthetic HPRT1 locus is indicated with a grey bar. Mouse datasets are: DNase-seq from ES-E14 mouse embryonic stem cells, ENCSR000CMW. ChIP-seq from ES-Bruce mouse embryonic stem cells, ENCSR000CBG, ENCSR000CDE, ENCSR000CFN, ENCSR000CCC. RNA-seq from ES-E14 mouse embryonic stem cells, ENCSR000CWC. Human datasets are: DNase-seq from H1-hESC ENCSR000EJN, ChIP-seq from H1-hESC ENCSR443YAS, ENCSR880SUY, ENCSR928HYM, RNA-seq from H1-hESC ENCSR000COU.
Extended Data Fig. 10 HPRT1RnoCpG design, assembly, and sequencing replicates.
a, b, GC-content (%GC) overlaid with coverage tracks for episomal HPRT1 RNA- and CAGE-seq (a), and episomal HPRT1R RNA- and CAGE-seq (b). %GC was calculated over 5 kb windows, and the range from 35-49% is indicated as a black line overlaying the coverage tracks. RNA-seq and CAGE-seq tracks are stranded, displayed with reverse strand reads inverted and below forward strand reads. c, A CpG-less version of the HPRT1R locus (HPRT1RnoCpG) was assembled from 36 segments into the same assemblon vector as HPRT1 and HPRT1R. The assemblon was purified and delivered to mESCs, integrating on chrX and on chr3. d, PCRs across the segment junctions for the HPRT1RnoCpG assembly. e, DNA sequencing coverage plots from next generation sequencing verification of assembled and integrated synthetic loci. The yeast assembly was whole genome sequenced and mESC samples were capture-sequenced. Assembly, episomal assemblon in yeast; Mm chrX, integrated on M. musculus chrX; Mm chr3, integrated on M. musculus chr3 – (1) and (2) indicate two independent mESC clones; %GC, GC-content as a line plot and color-scaled. f–i, Sequencing tracks for ATAC-seq, H3K4me3, RNAP2, H3K27ac, H3K27me3, and RNA-seq at HPRT1RnoCpG integrated on chrX (f, g) and on chr3 (h, i), in mESCs. Two biological replicates (Rep. 1, Rep. 2) are shown for two clones (denoted (1) and (2)) derived from independent integrations. The synthetic locus region is shaded. ~50 kb of flanking mouse genome is included for each position, with annotated mouse genes indicated.
Supplementary information
Supplementary Information
A full guide to supplementary Tables 1–6 (tables supplied separately).
Supplementary Tables
Supplementary Tables 1–6 – see Supplementary Information document for full descriptions.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Camellato, B.R., Brosh, R., Ashe, H.J. et al. Synthetic reversed sequences reveal default genomic states. Nature 628, 373–380 (2024). https://doi.org/10.1038/s41586-024-07128-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07128-2
This article is cited by
-
Mammalian cells repress random DNA that yeast transcribes
Nature (2024)
-
Musings on art and science
Nature Structural & Molecular Biology (2024)
-
New insights shed light on the enigma of genetic diversity and species complexity
Science China Life Sciences (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.