Abstract
Clostridioides difficile infection (CDI) is a major cause of healthcare-associated gastrointestinal infections1,2. The exaggerated colonic inflammation caused by C. difficile toxins such as toxin B (TcdB) damages tissues and promotes C. difficile colonization3,4,5,6, but how TcdB causes inflammation is unclear. Here we report that TcdB induces neurogenic inflammation by targeting gut-innervating afferent neurons and pericytes through receptors, including the Frizzled receptors (FZD1, FZD2 and FZD7) in neurons and chondroitin sulfate proteoglycan 4 (CSPG4) in pericytes. TcdB stimulates the secretion of the neuropeptides substance P (SP) and calcitonin gene-related peptide (CGRP) from neurons and pro-inflammatory cytokines from pericytes. Targeted delivery of the TcdB enzymatic domain, through fusion with a detoxified diphtheria toxin, into peptidergic sensory neurons that express exogeneous diphtheria toxin receptor (an approach we term toxogenetics) is sufficient to induce neurogenic inflammation and recapitulates major colonic histopathology associated with CDI. Conversely, mice lacking SP, CGRP or the SP receptor (neurokinin 1 receptor) show reduced pathology in both models of caecal TcdB injection and CDI. Blocking SP or CGRP signalling reduces tissue damage and C. difficile burden in mice infected with a standard C. difficile strain or with hypervirulent strains expressing the TcdB2 variant. Thus, targeting neurogenic inflammation provides a host-oriented therapeutic approach for treating CDI.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article (and its supplementary data and source data files). All biological materials are available upon request from the co-corresponding authors. The following sources were utilized for this work: Allen Brain Atlas (https://mousespinal.brain-map.org/) and SILVA (https://www.arb-silva.de/no_cache/download/archive/release_132/). Mouse and human intestinal cell data are available from the Broad Institute Single Cell portal with the accession number SCP1038. The following datasets from the NCBI Expression Omnibus database were used: for stromal data, GSE132465, GSE132257 and GSE144735; for colon-innervating DRG neurons (https://hockley.shinyapps.io/ColonicRNAseq/) GSE102962. Source data are provided with this paper.
References
Smits, W. K., Lyras, D., Lacy, D. B., Wilcox, M. H. & Kuijper, E. J. Clostridium difficile infection. Nat. Rev. Dis. Primers 2, 16020 (2016).
Guh, A. Y. et al. Trends in U.S. burden of Clostridioides difficile infection and outcomes. N. Engl. J. Med. 382, 1320–1330 (2020).
Kelly, C. P., Pothoulakis, C. & LaMont, J. T. Clostridium difficile colitis. N. Engl. J. Med. 330, 257–262 (1994).
El Feghaly, R. E. et al. Markers of intestinal inflammation, not bacterial burden, correlate with clinical outcomes in Clostridium difficile infection. Clin. Infect. Dis. 56, 1713–1721 (2013).
Fletcher, J. R. et al. Clostridioides difficile exploits toxin-mediated inflammation to alter the host nutritional landscape and exclude competitors from the gut microbiota. Nat. Commun. 12, 462 (2021).
Pruss, K. M. & Sonnenburg, J. L. C. difficile exploits a host metabolite produced during toxin-mediated disease. Nature 593, 261–265 (2021).
Yu, H. et al. Cytokines are markers of the Clostridium difficile-induced inflammatory response and predict disease severity. Clin. Vaccine Immunol. 24, e00037-17 (2017).
Solomon, K. et al. Mortality in patients with Clostridium difficile infection correlates with host pro-inflammatory and humoral immune responses. J. Med. Microbiol. 62, 1453–1460 (2013).
Kelly, C. P. et al. Host immune markers distinguish Clostridioides difficile infection from asymptomatic carriage and non-C. difficile diarrhea. Clin. Infect. Dis. 70, 1083–1093 (2020).
Abhyankar, M. M. et al. Immune profiling to predict outcome of Clostridioides difficile infection. mBio 11, e00905–e00920 (2020).
Aktories, K., Schwan, C. & Jank, T. Clostridium difficile toxin biology. Annu. Rev. Microbiol. 71, 281–307 (2017).
Kordus, S. L., Thomas, A. K. & Lacy, D. B. Clostridioides difficile toxins: mechanisms of action and antitoxin therapeutics. Nat. Rev. Microbiol. 20, 285–298 (2021).
Choi, J. E. & Di Nardo, A. Skin neurogenic inflammation. Semin. Immunopathol. 40, 249–259 (2018).
Chiu, I. M., von Hehn, C. A. & Woolf, C. J. Neurogenic inflammation and the peripheral nervous system in host defense and immunopathology. Nat. Neurosci. 15, 1063–1067 (2012).
Huang, J. et al. Clostridium difficile toxins induce VEGF-A and vascular permeability to promote disease pathogenesis. Nat. Microbiol. 4, 269–279 (2019).
Castagliuolo, I. et al. Neuronal involvement in the intestinal effects of Clostridium difficile toxin A and Vibrio cholerae enterotoxin in rat ileum. Gastroenterology 107, 657–665 (1994).
Kelly, C. P. et al. Neutrophil recruitment in Clostridium difficile toxin A enteritis in the rabbit. J. Clin. Invest. 93, 1257–1265 (1994).
Pothoulakis, C. et al. CP-96,345, a substance P antagonist, inhibits rat intestinal responses to Clostridium difficile toxin A but not cholera toxin. Proc. Natl Acad. Sci. USA 91, 947–951 (1994).
Castagliuolo, I. et al. Increased substance P responses in dorsal root ganglia and intestinal macrophages during Clostridium difficile toxin A enteritis in rats. Proc. Natl Acad. Sci. USA 94, 4788–4793 (1997).
Castagliuolo, I. et al. Neurokinin-1 (NK-1) receptor is required in Clostridium difficile-induced enteritis. J. Clin. Invest. 101, 1547–1550 (1998).
Keates, A. C. et al. CGRP upregulation in dorsal root ganglia and ileal mucosa during Clostridium difficile toxin A-induced enteritis. Am. J. Physiol. 274, G196–G202 (1998).
Mantyh, C. R., McVey, D. C. & Vigna, S. R. Extrinsic surgical denervation inhibits Clostridium difficile toxin A-induced enteritis in rats. Neurosci. Lett. 292, 95–98 (2000).
Lyras, D. et al. Toxin B is essential for virulence of Clostridium difficile. Nature 458, 1176–1179 (2009).
Kuehne, S. A. et al. The role of toxin A and toxin B in Clostridium difficile infection. Nature 467, 711–713 (2010).
Carter, G. P. et al. Defining the roles of TcdA and TcdB in localized gastrointestinal disease, systemic organ damage, and the host response during Clostridium difficile infections. mBio 6, e00551 (2015).
Russell, F. A., King, R., Smillie, S. J., Kodji, X. & Brain, S. D. Calcitonin gene-related peptide: physiology and pathophysiology. Physiol. Rev. 94, 1099–1142 (2014).
Tao, L. et al. Frizzled proteins are colonic epithelial receptors for C. difficile toxin B. Nature 538, 350–355 (2016).
Chen, P. et al. Structural basis for recognition of frizzled proteins by Clostridium difficile toxin B. Science 360, 664–669 (2018).
Chen, P. et al. Structural basis for CSPG4 as a receptor for TcdB and a therapeutic target in Clostridioides difficile infection. Nat. Commun. 12, 3748 (2021).
Yuan, P. et al. Chondroitin sulfate proteoglycan 4 functions as the cellular receptor for Clostridium difficile toxin B. Cell Res. 25, 157–168 (2015).
Simonetti, M. et al. Wnt–Fzd signaling sensitizes peripheral sensory neurons via distinct noncanonical pathways. Neuron 83, 104–121 (2014).
Liu, Z. et al. Structural basis for selective modification of Rho and Ras GTPases by Clostridioides difficile toxin B. Sci. Adv. 7, eabi4582 (2021).
Terada, N. et al. Immunohistochemical study of NG2 chondroitin sulfate proteoglycan expression in the small and large intestines. Histochem. Cell Biol. 126, 483–490 (2006).
Lee, H. O. et al. Lineage-dependent gene expression programs influence the immune landscape of colorectal cancer. Nat. Genet. 52, 594–603 (2020).
Sweeney, M. D., Ayyadurai, S. & Zlokovic, B. V. Pericytes of the neurovascular unit: key functions and signaling pathways. Nat. Neurosci. 19, 771–783 (2016).
Muhl, L. et al. Single-cell analysis uncovers fibroblast heterogeneity and criteria for fibroblast and mural cell identification and discrimination. Nat. Commun. 11, 3953 (2020).
Proebstl, D. et al. Pericytes support neutrophil subendothelial cell crawling and breaching of venular walls in vivo. J. Exp. Med. 209, 1219–1234 (2012).
Hartmann, D. A. et al. Brain capillary pericytes exert a substantial but slow influence on blood flow. Nat. Neurosci. 24, 633–645 (2021).
Stark, K. et al. Capillary and arteriolar pericytes attract innate leukocytes exiting through venules and ‘instruct’ them with pattern-recognition and motility programs. Nat. Immunol. 14, 41–51 (2013).
Saito, M. et al. Diphtheria toxin receptor-mediated conditional and targeted cell ablation in transgenic mice. Nat. Biotechnol. 19, 746–750 (2001).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Lai, N. Y. et al. Gut-innervating nociceptor neurons regulate Peyer’s patch microfold cells and SFB levels to mediate Salmonella host defense. Cell 180, 33–49 e22 (2020).
He, M. et al. Emergence and global spread of epidemic healthcare-associated Clostridium difficile. Nat. Genet. 45, 109–113 (2013).
Lanis, J. M., Heinlen, L. D., James, J. A. & Ballard, J. D. Clostridium difficile 027/BI/NAP1 encodes a hypertoxic and antigenically variable form of TcdB. PLoS Pathog. 9, e1003523 (2013).
Mansfield, M. J. et al. Phylogenomics of 8,839 Clostridioides difficile genomes reveals recombination-driven evolution and diversification of toxin A and B. PLoS Pathog. 16, e1009181 (2020).
Shen, E. et al. Subtyping analysis reveals new variants and accelerated evolution of Clostridioides difficile toxin B. Commun. Biol. 3, 347 (2020).
Lopez-Urena, D. et al. Toxin B variants from Clostridium difficile strains VPI 10463 and NAP1/027 share similar substrate profile and cellular intoxication kinetics but use different host cell entry factors. Toxins 11, 348 (2019).
Chung, S. Y. et al. The conserved Cys-2232 in Clostridioides difficile toxin B modulates receptor binding. Front. Microbiol. 9, 2314 (2018).
Luo, J. et al. TFPI is a colonic crypt receptor for TcdB from hypervirulent clade 2 C. difficile. Cell 185, 980–994.e15 (2022).
Schottelndreier, D., Langejurgen, A., Lindner, R. & Genth, H. Low density lipoprotein receptor-related protein-1 (LRP1) is involved in the uptake of Clostridioides difficile toxin A and serves as an internalizing receptor. Front. Cell Infect. Microbiol. 10, 565465 (2020).
Tao, L. et al. Sulfated glycosaminoglycans and low-density lipoprotein receptor contribute to Clostridium difficile toxin A entry into cells. Nat. Microbiol. 4, 1760–1769 (2019).
Mileto, S. J. et al. Clostridioides difficile infection damages colonic stem cells via TcdB, impairing epithelial repair and recovery from disease. Proc. Natl Acad. Sci. USA 117, 8064–8073 (2020).
Porat-Shliom, N., Milberg, O., Masedunskas, A. & Weigert, R. Multiple roles for the actin cytoskeleton during regulated exocytosis. Cell. Mol. Life Sci. 70, 2099–2121 (2013).
Meyer, G. K. et al. Clostridium difficile toxins A and B directly stimulate human mast cells. Infect. Immun. 75, 3868–3876 (2007).
Geissler, B., Tungekar, R. & Satchell, K. J. Identification of a conserved membrane localization domain within numerous large bacterial protein toxins. Proc. Natl Acad. Sci. USA 107, 5581–5586 (2010).
Brescia, P. & Rescigno, M. The gut vascular barrier: a new player in the gut–liver–brain axis. Trends Mol. Med. 27, 844–855 (2021).
Spadoni, I. et al. A gut–vascular barrier controls the systemic dissemination of bacteria. Science 350, 830–834 (2015).
Drokhlyansky, E. et al. The human and mouse enteric nervous system at single-cell resolution. Cell 182, 1606–1622.e23 (2020).
Thompson, B. J. et al. Protective roles of α-calcitonin and β-calcitonin gene-related peptide in spontaneous and experimentally induced colitis. Dig. Dis. Sci. 53, 229–241 (2008).
Song, H. et al. Functional characterization of pulmonary neuroendocrine cells in lung development, injury, and tumorigenesis. Proc. Natl Acad. Sci. USA 109, 17531–17536 (2012).
Zhu, X., Bergles, D. E. & Nishiyama, A. NG2 cells generate both oligodendrocytes and gray matter astrocytes. Development 135, 145–157 (2008).
Harris, J. A. et al. Anatomical characterization of Cre driver mice for neural circuit mapping and manipulation. Front. Neural Circuits 8, 76 (2014).
Cao, Y. Q. et al. Primary afferent tachykinins are required to experience moderate to intense pain. Nature 392, 390–394 (1998).
Grako, K. A., Ochiya, T., Barritt, D., Nishiyama, A. & Stallcup, W. B. PDGFα-receptor is unresponsive to PDGF-AA in aortic smooth muscle cells from the NG2 knockout mouse. J. Cell Sci. 112, 905–915 (1999).
Huang, H. et al. Generation of a NK1R-CreER knockin mouse strain to study cells involved in neurokinin 1 receptor signaling. Genesis 54, 593–601 (2016).
Nassar, M. A. et al. Nociceptor-specific gene deletion reveals a major role for Nav1.7 (PN1) in acute and inflammatory pain. Proc. Natl Acad. Sci. USA 101, 12706–12711 (2004).
Buch, T. et al. A Cre-inducible diphtheria toxin receptor mediates cell lineage ablation after toxin administration. Nat. Methods 2, 419–426 (2005).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Lyon, M. F. & Glenister, P. H. A new allele sash (Wsh) at the W-locus and a spontaneous recessive lethal in mice. Genet. Res. 39, 315–322 (1982).
Zhang, Y. et al. The role of purified Clostridium difficile glucosylating toxins in disease pathogenesis utilizing a murine cecum injection model. Anaerobe 48, 249–256 (2017).
Chen, X. et al. A mouse model of Clostridium difficile-associated disease. Gastroenterology 135, 1984–1992 (2008).
Hockley, J. R. F. et al. Single-cell RNAseq reveals seven classes of colonic sensory neuron. Gut 68, 633–644 (2019).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Rao, M. et al. Enteric glia express proteolipid protein 1 and are a transcriptionally unique population of glia in the mammalian nervous system. Glia 63, 2040–2057 (2015).
Lau, M. T. et al. Molecular dissection of box jellyfish venom cytotoxicity highlights an effective venom antidote. Nat. Commun. 10, 1655 (2019).
Davies, A. J. et al. Natural killer cells degenerate intact sensory afferents following nerve injury. Cell 176, 716–728.e18 (2019).
Jank, T., Giesemann, T. & Aktories, K. Clostridium difficile glucosyltransferase toxin B-essential amino acids for substrate binding. J. Biol. Chem. 282, 35222–35231 (2007).
Wilson, K. H., Kennedy, M. J. & Fekety, F. R. Use of sodium taurocholate to enhance spore recovery on a medium selective for Clostridium difficile. J. Clin. Microbiol. 15, 443–446 (1982).
Rao, C. et al. Multi-kingdom ecological drivers of microbiota assembly in preterm infants. Nature 591, 633–638 (2021).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina paired-end read merger. Bioinformatics 30, 614–620 (2014).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Acknowledgements
We thank members of the Dong and Rao Laboratories for technical assistance and suggestions. This study was partially supported by grants from the National Institutes of Health (NIH): R01NS080833, R01NS117626, R01AI132387 and R01AI139087 to M.D.; K08DK110532, R01DK135707 and R01DK130836 to M.R.; DP2GM136652 to S.R.-N.; T32DK007477 for supporting M.A.M.; and R01HL150106 to K.Y. G.A.K. was supported by a NDSEG Fellowship and A.S. by a Schmidt Science Fellowship. We acknowledge support from the NIH-funded Harvard Digestive Disease Center (P30DK034854) and the Boston Children’s Hospital Intellectual and Developmental Disabilities Research Center (P30HD18655). M.R. was supported by an Odyssey Award from the Richard and Susan Smith Family Foundation. M.D. holds the Investigator in the Pathogenesis of Infectious Disease Award from the Burroughs Wellcome Fund. S.R.-N. acknowledges support from Mathers Foundation, Career Award for Medical Scientists from the Burroughs Wellcome Fund, and a Pew Biomedical Scholarship. Figure 3c was created using BioRender (https://biorender.com).
Author information
Authors and Affiliations
Contributions
J.M. and M.D. conceived the project. J.M., M.A.M., G.A.K., A.S., M.L., S.W., M.R. and M.D. participated in experimental design, execution and data analyses. G.A.K., M.L., A.S. and S.W. contributed equally. Cultured DRG studies were performed by J.M. and L.Z. Cultured pericyte studies were performed by J.M. and M.L. J.Z. provided essential training and established surgical models. M.A.M. and M.R. performed most of the immunohistochemistry and confocal imaging studies with assistance from J.M. and A.S. J.M., G.A.K. and S.R.-N. performed microbiome analyses and in vitro analyses of C. difficile. J.M. conceived the toxogenetic approaches. P.-G.L. designed and purified the toxogenetic proteins. K.Y. helped with pericyte culture and analysis. J.D.G. provided human tissue samples. R.G. and R.J. designed and provided essential reagents. R.K.R.M. and J.G.H. prepared C. difficile spores and strains. M.R. and M.D. supervised and coordinated the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Histopathology score analysis and time course of TcdB-induced inflammation in a mouse cecum injection model.
a, A list of criteria for assessing histopathology scores of colonic tissues. b-e, H&E micrographs illustrating oedema (b, black line indicates oedema extent); immune cell infiltration (c); epithelial disruption (d); haemorrhage or congestion (e). f, Representative histopathology of human CDI colon tissues compared to a normal control, illustrating neutrophilia, oedematous inflammation, and pseudomembranes. g-j, The subcategories for the histopathology scoring in Fig. 1b: oedema (g), immune cell infiltration (h), epithelial disruption (i), and haemorrhage or congestion (j). n = 5, 6, 9, 5 mice. k-l, SP (k) and CGRP (l) levels in colonic explants from the mouse cecal TcdB injection model at the indicated time (min) following TcdB injection. Vehicle was at 120 min. n = 6, 7, 7, 7, 7 mice. m-n, The levels of SP (m) and CGRP (n) in colonic explants from CDI mouse models infected with either C. difficile 630 or a toxin-null strain (630 A- B-) at 23 h (n = 5), 37 h (n = 5), and 48 h (n = 3), normalized to mice treated with antibiotics but gavaged with saline (time point 0, n = 5). g-j, assessed by one-way ANOVA with post hoc Dunnett’s test. k-l, assessed by Kruskall-Wallis test with post-hoc Dunn’s test. Centre line, mean; error bars reflect the standard error of the mean (SEM); n.s., not significant, p values are exact. Scale bar is 50 µm in b, c and 20 µm in d and e. n is described from left to right.
Extended Data Fig. 2 Histological scoring for cecum injections of TcdB in the indicated mouse models as described in Fig. 1c,d.
a-e, Histopathology total scores and subscores of TcdB injected Tac1 KO mice, n = 7, 8 mice. f-j, Histopathology total scores and subscores of TcdB injected Nk1r KO mice, n = 7, 7 mice. k-o, Histopathology total scores and subscores of TcdB injected Calca KO mice, n = 8, 10 mice. p-t, Histopathology total scores and subscores of TcdB injected Calcb KO mice, n = 3, 5 mice. u, Representative histopathology of Tac1 KO mice injected with TcdB compared to Tac1 KO mice injected with TcdB and SP in the cecal TcdB injection model (120 min incubation). v, Histopathology scores of experiments described in u, n = 3, 4 mice. Assessed by Student’s T-test, two tailed, relative to WT control mice. Centre line: mean; error bars: SEM; n.s., not significant, p values are exact. n is described from left to right.
Extended Data Fig. 3 Histological scoring for CDI in the indicated KO mouse models as described in Fig. 1e,f.
a-e, Histopathology total scores and subscores of CDI in Tac1 KO mice, n = 11, 20 mice. f-j, Histopathology total scores and subscores of CDI in Nk1r KO mice, n = 7, 10 mice. k-o, Histopathology total scores and subscores of CDI in Calcb KO mice, n = 6, 9, 7 mice. a-j, assessed by Student’s T-test, two tailed, relative to WT control mice. k-o, assessed by one-way ANOVA with post hoc Dunnett’s test. Centre line: mean; error bars: SEM; n.s., not significant, p values are exact. n is described from left to right.
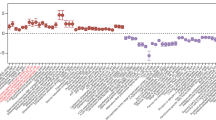
Extended Data Fig. 4 FZD1/2/7 are expressed in DRG and enteric neurons.
a, Single cell RNA-seq analysis of FZD1, 2, and 7 expression in different cell types within colonic tissues in mice58. Size of dot indicates proportion expressing whereas the colour indicates expression level (red high, blue low). Putative excitatory motor neuron (PEMN), putative inhibitory motor neuron (PIMN), putative sensory neuron (PSN), putative sudomotor and vasodilator neurons (PSVN), putative interneuron (PIN). b, FZD1, 2, and 7 expression in DRG neurons that innervate colonic tissues are displayed as violin plots72. c, In situ hybridization of spinal cord sections shows enriched expression of FZD1/2/7 in dorsal root ganglia areas. Data are from Allen Brain Atlas73. d, Phase-contrast images show that DRG neurons exposed to TcdB (1 pM, 60 min) exhibit no morphological changes. Scale bar is 50 µm. n = 3 independent replicates. e, RNAscope in situ hybridization showing localization of Pdgfrb expression (magenta) within Cspg4+ cells (cyan, Cspg4-dsRed mice) in colonic tissues. Panels in bottom row show regions in yellow boxed area at higher magnification. Scale bar: 50 µm and lower panel is 20 µm.
Extended Data Fig. 5 Cspg4+ pericytes wrap around blood vessels and form close contact with CGRP-containing afferent neurons.
a, Colon tissues from Cspg4-DsRed mice were harvested and subjected to whole mount immunofluorescence analysis, using an antibody against CD31 (a marker for blood vessel endothelial cells; magenta). DsRed (cyan) signals were detected directly. DsRed-positive cells surround CD31-positive capillaries in colonic crypts. Maximum intensity projection of the whole stack is presented in Fig. 2e. Scale bar is 50 µm. b, Experiments were carried out as in panel a, except that lymphatic endothelial cells were labeled with an antibody against Lyve-1 (magenta) and DsRed signals were detected using an antibody against DsRed. DsRed-positive cells are clearly separate from lymphatic endothelial cells. Scale bar is 50 µm. c-f, Experiments were carried out as described in panel a. Enteric glial cells were visualized with GFP fluorescence in examining proteolipid protein 1 (PLP1)-GFP/-Cspg4-dsRed dual reporter mice (panel c). Fibroblasts were labeled using an antibody against platelet-derived growth factor receptor alpha (PGDFRA, magenta, panel d). Myofibroblasts were labeled using an antibody against smooth muscle actin (SMA, magenta, panel e). Mast cells and interstitial cells of Cajal were labeled using an antibody against cKit (magenta, panel f). Scale bar: 10 µm. g, TcdB induced cell-rounding of cultured human brain vascular pericytes. Scale bar: 25 µm. h, The indicated TcdB mutants were injected into mouse ears via intradermal injection37. Pericytes surrounding ear arterioles were labeled and visualized through DsRed. TcdB and TcdB-FzM induced morphological changes of Cspg4+ pericytes surrounding ear arterioles, whereas TcdB-Cspg4M showed no effect. Scale bar, 50 µm. n = 3 mice/group. i, Experiments were carried out as described in panel h, except that WT mice were utilized and pericytes were detected by immunostaining using an antibody against smooth muscle actin (labeling pericytes). TcdB-FzM disrupted pericytes around ear arterioles, whereas TcdB-Cspg4M has no effect. Scale bar: 50 µm. n = 3 mice/group. j, Experiments were carried out as in panel a, except that neuronal processes were labeled using an antibody against Tubb3 (magenta, a marker for neuronal processes), showing that neuronal processes are extending alongside DsRed-positive pericytes. Scale bar: 10 µm. k, Experiments were carried out as in panel a, except that an antibody against CGRP was added to detect CGRP-positive nerve terminals. 3-dimensional reconstruction of images showed that DsRed+ pericytes (cyan) surround the vasculature (CD31 endothelial marker; magenta) and contact CGRP-expressing nerve terminals (CGRP; yellow). Scale bar: 10 µm. l, Immunoblot analysis of cell lysates showed expression of CGRP receptors (CALCRL: calcitonin receptor like receptor; and RAMP1: receptor activity modifying protein 1) in primary cultured human brain vascular pericytes. Two human cell lines, HeLa and U87, were analyzed in parallel as controls, which do not express detectable levels of CGRP receptors. Total protein staining with Coomassie blue was used as a control for protein loading. For gel source data, see Supplementary Fig. 1. N = 2 replicates.
Extended Data Fig. 6 AAV-DTR mediates selective expression of DTR in ENS and DRG neurons in Tac1-Cre mice.
a, AAV-DTR was administered to Tac1-Cre mice via tail vein injection. Colon tissues were harvested 3-weeks later and subjected to whole mount immunofluorescence analysis. DRG were harvested and their cryosections were analyzed. HuC/D (magenta) marks neuron soma in colon tissues. An antibody against β3-tubulin (TUBB3, magenta) was utilized to label DRG neurons. DTR-GFP was labeled using an antibody against GFP (cyan). Scale for DRG: 100 µm; for myenteric plexus: 50 µm. Images were collected using a confocal microscope and maximal intensity projection is presented. GFP is directly fused to the C-terminus of DTR, thus cyan color represents DTR expression. n = 4 mice. b, Experiments were carried out as described in upper panel. 5-HT (magenta) labels enterochromaffin cells in colonic tissues, showing no co-expression of DTR-GFP (cyan). Scale bars represent 100 µm (50 µm for right panel, which shows boxed region at higher magnification). c-d, Experiments were carried out as described in Fig. 3e. Colonic tissues (panel c) and DRG (panel d) were isolated and subjected to analysis as described in panel a. WT mice not exposed to AAV-DTR were analyzed in parallel as a negative control (right panels). HuC/D labels myenteric plexus neurons within the proximal colon and TUBB3 labels DRG neurons of a wildtype mouse injected with AAV-DTR. No GFP positive cell bodies or fibers were detected in WT mice. Scale bar is 50 µm. e-h, Sub-scoring of histopathology, related to Fig. 3f, n = 7, 6, 4 mice. One-way ANOVAs were applied with post hoc Tukey’s test as indicated. Centre line: mean; error bars: SEM; n.s., not significant, p values are exact. n is described from left to right.
Extended Data Fig. 7 Extrinsic afferent neuron-dependent secretion of SP is necessary and sufficient for TcdB mediated pathology.
a-d; Histopathology subscoring of experiments following TcdB administration to RTX treated mice. Related to Fig. 3h, n = 10, 10 mice. e-j, Histopathology of RTX treated/vehicle mice treated with TcdB alone or TcdB and SP for 120 min, demonstrating that SP can restore the effects of TcdB in RTX-treated mice. Representative histopathology in e, and histopathology scores f-j, n = 4, 4, 6, and 6 mice. The scale bar is 50 µm. k, CGRP secretion from RTX treated mice is reduced in the cecal TcdB injection model, and is restored by co-administration of SP with TcdB. n = 2, 4, 6, 4, and 6 mice. l-m, Nav1.8Cre/+ Rosa-LSL-Tdtomato mice have no evident Tomato expression (magenta) in the ENS (myenteric neurons marked with HuC/D, cyan) with only sparse Tomato+ nerve fibers from extrinsic neurons labeled in the colon (l). In contrast, there are many Tomato+ neurons in DRG (m). Scale bar: 50 µm. n-q, Histopathology subscores of Nav1.8Cre/+, Rosa-LSL-DTR mice treated with GTD-iDT related to Fig 3i, j; n = 7, 9 mice. For a-d and n-q, significance was assessed by student’s two-tailed t-test. For f-k, significance was assessed by one-way ANOVA with post hoc Dunnett’s test. Centre line: mean; error bars: SEM. p values are exact. n is described from left to right.
Extended Data Fig. 8 Inhibiting SP or CGRP signaling reduces severity of C. difficile colitis.
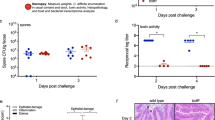
a, CDI models were carried out on WT versus Tac1 KO mice, using either C. difficile 630 Δerm or a toxin-null mutant strain (A-/B-). C. difficile CFUs from faeces were analyzed at 2 days post infection. The toxin-null strain showed reduced colonization on WT mice compared with the standard 630 Δerm, suggesting that the presence of toxins facilitates C. difficile colonization. In contrast, 630 Δerm showed similar levels of CFUs as the toxin-null strain in Tac1 KO mice, suggesting that the toxin’s contribution to C. difficile colonization relies on intact SP signaling. n = 7, 6, 4, 3 mice. b, CDI models were carried out on WT versus Calcb KO mice (using C. difficile 630 Δerm). C. difficile CFUs from faeces showed reduced colonization in Calcb KO mice compared with WT mice, n = 4, 5 mice. c, Representative H&E images for the experiments described in Fig. 4c. Scale bar is 50 µm. d-g, Sub-categories of histopathological scoring of Fig. 4c; n = 15, 12, 11, 3. h-l, Histopathology scoring of mice administered vehicle or aprepitant at the indicated times (0, 30 or 60 min following cecum injection and wound closure); n = 7, 6, 6, 7 mice. Significance was assessed for a – Two-Way ANOVA with post hoc Sidak’s test and by T-test for b. One Way ANOVA with post hoc Dunnett’s test was used for d-l. Centre line: mean; error bars: SEM. p values are exact. n is described from left to right.
Extended Data Fig. 9 Inhibiting SP or CGRP reduces histopathological scores in cecum injection assays and in CDI models.
a, Representative H&E images for the experiment described in Fig. 4d. Scale bar is 50 µm. b-e, Sub-categories of histopathological scoring of Fig. 4d, n = 13,7, 8, 8 mice. f, C. difficile 630 Δerm fitness (measured by OD600) was not affected by treatment with aprepitant or olcegepant (20 µM) in vitro. Representative of 2 independent experiments (3 wells/group). g-h, CDI experiments (with C. difficile 630) were carried out on WT and Tac1 KO mice (co-housed from weaning), followed by treatment with vehicle or aprepitant. C. difficile colonization (panel g) and histopathological scores (panel h) were shown, n = 7, 7, 6, 7 mice. i-j, CDI experiments (with C. difficile 630) were carried out on WT and Calcb KO mice (littermates, co-housed from birth), followed by treatment with vehicle or olcegepant. C. difficile colonization (panel i) and histopathological scores (panel j) were shown. n = 4, 4, 5, 7 mice. k, TcdA induced oedema in footpad injection assays in mice, which is prevented by pre-treatment with aprepitant. Boiled TcdA solution did not induce any oedema. n = 7,4, 3 mice. Significance was assessed by one-way ANOVA with post hoc Dunnett’s test. Centre line: mean; error bars: SEM. n.s., not significant, p values are exact. n is described from left to right.
Extended Data Fig. 10 Inhibiting SP or CGRP reduces histopathological scores and colonization of hypervirulent strains that express TcdB2.
a, Representative H&E images for the experiments described in Fig. 4i. Scale bar is 50 µm. b-e, Sub-categories of histopathological scoring of Fig. 4i, n = 3, 9, 7 mice. f, Representative H&E images for the experiment described in Fig. 4j. Scale bar is 50 µm. g-j, Sub-categories of histopathological scoring of Fig. 4j, n = 21, 10, 10, 12, 10, 5 mice. Significance was assessed by one-way ANOVA with post hoc Dunnett’s test. Centre line: mean; error bars: SEM; n.s., not significant; p values are exact. n is described from left to right.
Supplementary information
Supplementary Information
Antibody information (Supplementary Table 1) and source western blot scans and Coomassie staining related to Extended Data Fig. 5l (Supplementary Fig. 1).
Supplementary Video 1
Pericytes envelope the colonic vascular network. Mouse colonic tissue from CSPG4–DsRed mice was collected, processed by whole mount and subjected to immunostaining with BABB optical clearing of tissue. Z-stack images were captured on a Zeiss LSM 700 confocal microscope with Zen Black software. Z-stack images were processed and rendered into 3D image movies using Aivia image analysis software. In the 3D image movie, an initial view is seen peering down at epithelium lining bottom of intestinal crypts (background nonspecific green fluorescence signal). Removal of epithelial background reveals CD31-immunolabelled capillary networks and a deeper arteriole (green). DsRed antibody immunolabelled CSPG4–DsRed reporter pericytes (magenta) are found in close juxtaposition to the vascular network throughout.
Supplementary Video 2
Pericytes wrap around mucosal capillaries. Mouse colonic tissue from CSPG4–DsRed mice was collected, processed by cryosection and subjected to immunostaining. Z-stack images were captured on a Zeiss LSM 880 confocal microscope with Zen Black software. Z-stack images were processed and rendered into 3D image movies using Aivia image analysis software. The movie demonstrates a 3D view of two CSPG4-reporter labelled pericytes (magenta). CD31-immunolabelled capillaries (green) are found nestled within the pericyte bodies and wrapped within pericyte cellular processes.
Supplementary Video 3
Colonic neurovascular unit comprised pericyte, capillary and CGRP+ neuron processes. Mouse colonic tissue from CSPG4–DsRed mice was collected, processed by cryosection and subjected to immunostaining. Z-stack images were captured on a Zeiss LSM 880 confocal microscope with Zen Black software. Z-stack images were processed and rendered into 3D image movies using Aivia image analysis software. The movie demonstrates a 3D image of a colonic neurovascular unit. A single CSPG4–DsRed reporter-labelled pericyte (magenta) is shown. A CD31-immunolabelled vessel (green) is surrounded by the pericyte body and its processes. CGRP-immunolabelled neuron processes (grey) appear to interact with the pericyte at multiple points.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Manion, J., Musser, M.A., Kuziel, G.A. et al. C. difficile intoxicates neurons and pericytes to drive neurogenic inflammation. Nature 622, 611–618 (2023). https://doi.org/10.1038/s41586-023-06607-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06607-2
This article is cited by
-
Clostridioides difficile infection drives neuronal inflammation
Nature (2023)
-
C. difficile toxin strikes a nerve in the gut
Nature Reviews Gastroenterology & Hepatology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.