Abstract
The cerebellum is thought to help detect and correct errors between intended and executed commands1,2 and is critical for social behaviours, cognition and emotion3,4,5,6. Computations for motor control must be performed quickly to correct errors in real time and should be sensitive to small differences between patterns for fine error correction while being resilient to noise7. Influential theories of cerebellar information processing have largely assumed random network connectivity, which increases the encoding capacity of the network’s first layer8,9,10,11,12,13. However, maximizing encoding capacity reduces the resilience to noise7. To understand how neuronal circuits address this fundamental trade-off, we mapped the feedforward connectivity in the mouse cerebellar cortex using automated large-scale transmission electron microscopy and convolutional neural network-based image segmentation. We found that both the input and output layers of the circuit exhibit redundant and selective connectivity motifs, which contrast with prevailing models. Numerical simulations suggest that these redundant, non-random connectivity motifs increase the resilience to noise at a negligible cost to the overall encoding capacity. This work reveals how neuronal network structure can support a trade-off between encoding capacity and redundancy, unveiling principles of biological network architecture with implications for the design of artificial neural networks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The electron microscopy dataset is publicly available at the BossDB (https://bossdb.org/) repository https://bossdb.org/project/nguyen_thomas2022. Further directions for accessing neuron 3D renderings and derived neuron connectivity graphs are available at https://github.com/htem/cb2_project_analysis. Source data are provided with this paper.
Code availability
Software developed in this work is open-source and available in the following repositories. Daisy: https://github.com/funkelab/daisy. MD-Seg front-end: https://github.com/htem/neuroglancer_mdseg/tree/segway_pr_v2 MD-Seg back-end: https://github.com/htem/segway.mdseg. Segmentation and synapse prediction scripts: https://github.com/htem/segway. Analysis code: https://github.com/htem/cb2_project_analysis. Other software packages used in this work are described in ‘Software and code’ in the Reporting Summary.
Change history
11 January 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41586-023-05703-7
References
Wolpert, D. M., Miall, R. C. & Kawato, M. Internal models in the cerebellum. Trends Cogn. Sci. 2, 338–347 (1998).
Ebner, T. J. & Pasalar, S. Cerebellum predicts the future motor state. Cerebellum 7, 583–588 (2008).
Strick, P. L., Dum, R. P. & Fiez, J. A. Cerebellum and nonmotor function. Annu. Rev. Neurosci. 32, 413–434 (2009).
Koziol, L. F. et al. Consensus paper: the cerebellum’s role in movement and cognition. Cerebellum 13, 151–177 (2014).
Schmahmann, J. D. Disorders of the cerebellum: ataxia, dysmetria of thought, and the cerebellar cognitive affective syndrome. J. Neuropsychiatry Clin. Neurosci. 16, 367–378 (2004).
Wagner, M. J. & Luo, L. Neocortex–cerebellum circuits for cognitive processing. Trends Neurosci. 43, 42–54 (2020).
Cayco-Gajic, N. A. & Silver, R. A. Re-evaluating circuit mechanisms underlying pattern separation. Neuron 101, 584–602 (2019).
Marr, D. A theory of cerebellar cortex. J. Physiol. 202, 437–470 (1969).
Albus, J. S. A theory of cerebellar function. Math. Biosci. 10, 25–61 (1971).
Babadi, B. & Sompolinsky, H. Sparseness and expansion in sensory representations. Neuron 83, 1213–1226 (2014).
Billings, G., Piasini, E., Lőrincz, A., Nusser, Z. & Silver, R. A. Network structure within the cerebellar input layer enables lossless sparse encoding. Neuron 83, 960–974 (2014).
Cayco-Gajic, N. A., Clopath, C. & Silver, R. A. Sparse synaptic connectivity is required for decorrelation and pattern separation in feedforward networks. Nat. Commun. 8, 1116 (2017).
Litwin-Kumar, A., Harris, K. D., Axel, R., Sompolinsky, H. & Abbott, L. F. Optimal degrees of synaptic connectivity. Neuron 93, 1153–1164.e7 (2017).
Eccles, J. C., Ito, M. & Szentagothai, J. The Cerebellum as a Neuronal Machine (Springer–Verlag, 1967).
Palay, S. L. & Chan-Palay, V. Cerebellar Cortex: Cytology and Organization (Springer–Verlag, 1974).
Liu, A. & Regehr, W. G. Normalization of input patterns in an associative network. J. Neurophysiol. 111, 544–551 (2014).
Walter, J. T. & Khodakhah, K. The advantages of linear information processing for cerebellar computation. Proc. Natl Acad. Sci. USA 106, 4471–4476 (2009).
Ohmae, S. & Medina, J. F. Climbing fibers encode a temporal-difference prediction error during cerebellar learning in mice. Nat. Neurosci. 18, 1798–1803 (2015).
Heffley, W. et al. Coordinated cerebellar climbing fiber activity signals learned sensorimotor predictions. Nat. Neurosci. 21, 1431–1441 (2018).
Sanger, T. D., Yamashita, O. & Kawato, M. Expansion coding and computation in the cerebellum: 50 years after the Marr–Albus codon theory. J. Physiol. 598, 913–928 (2020).
Cao, Y., Geddes, T. A., Yang, J. Y. H. & Yang, P. Ensemble deep learning in bioinformatics. Nat. Mach. Intell. 2, 500–508 (2020).
Bell, C. C., Han, V. & Sawtell, N. B. Cerebellum-like structures and their implications for cerebellar function. Annu. Rev. Neurosci. 31, 1–24 (2008).
Stevens, C. F. What the fly’s nose tells the fly’s brain. Proc. Natl Acad. Sci. USA 112, 9460–9465 (2015).
Eichler, K. et al. The complete connectome of a learning and memory centre in an insect brain. Nature 548, 175–182 (2017).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Houston, C. M. et al. Exploring the significance of morphological diversity for cerebellar granule cell excitability. Sci. Rep. 7, 46147 (2017).
Sultan, F. Distribution of mossy fibre rosettes in the cerebellum of cat and mice: evidence for a parasagittal organization at the single fibre level. Eur. J. Neurosci. 13, 2123–2130 (2001).
Gilmer, J. I. & Person, A. L. Morphological constraints on cerebellar granule cell combinatorial diversity. J. Neurosci. 37, 12153–12166 (2017).
Knogler, L. D., Markov, D. A., Dragomir, E. I., Štih, V. & Portugues, R. Sensorimotor representations in cerebellar granule cells in larval zebrafish are dense, spatially organized, and non-temporally patterned. Curr. Biol. 27, 1288–1302 (2017).
Wagner, M. J., Kim, T. H., Savall, J., Schnitzer, M. J. & Luo, L. Cerebellar granule cells encode the expectation of reward. Nature 544, 96–100 (2017).
Giovannucci, A. et al. Cerebellar granule cells acquire a widespread predictive feedback signal during motor learning. Nat. Neurosci. 20, 727–734 (2017).
Heck, D. H., Thach, W. T. & Keating, J. G. On-beam synchrony in the cerebellum as the mechanism for the timing and coordination of movement. Proc. Natl Acad. Sci. USA 104, 7658–7663 (2007).
de Solages, C. et al. High-frequency organization and synchrony of activity in the purkinje cell layer of the cerebellum. Neuron 58, 775–788 (2008).
Wise, A. K., Cerminara, N. L., Marple-Horvat, D. E. & Apps, R. Mechanisms of synchronous activity in cerebellar Purkinje cells. J. Physiol. 588, 2373–2390 (2010).
Harvey, R. J. & Napper, R. M. A. Quantitatives studies on the mammalian cerebellum. Prog. Neurobiol. 36, 437–463 (1991).
Napper, R. M. & Harvey, R. J. Number of parallel fiber synapses on an individual Purkinje cell in the cerebellum of the rat. J. Comp. Neurol. 274, 168–177 (1988).
Isope, P. & Barbour, B. Properties of unitary granule cell→Purkinje cell synapses in adult rat cerebellar slices. J. Neurosci. 22, 9668–9678 (2002).
Lanore, F., Cayco-Gajic, N. A., Gurnani, H., Coyle, D. & Silver, R. A. Cerebellar granule cell axons support high-dimensional representations. Nat. Neurosci. 24, 1142–1150 (2021).
Dittman, J. S., Kreitzer, A. C. & Regehr, W. G. Interplay between facilitation, depression, and residual calcium at three presynaptic terminals. J. Neurosci. 20, 1374–1385 (2000).
Huang, C.-C. et al. Convergence of pontine and proprioceptive streams onto multimodal cerebellar granule cells. eLife 2, e00400 (2013).
Chabrol, F. P., Arenz, A., Wiechert, M. T., Margrie, T. W. & DiGregorio, D. A. Synaptic diversity enables temporal coding of coincident multisensory inputs in single neurons. Nat. Neurosci. 18, 718–727 (2015).
Wagner, M. J. et al. Shared cortex–cerebellum dynamics in the execution and learning of a motor task. Cell 177, 669–682.e24 (2019).
Kuan, A. T. et al. Dense neuronal reconstruction through X-ray holographic nano-tomography. Nat. Neurosci. 23, 1637–1643 (2020).
Zavitz, D., Amematsro, E. A., Borisyuk, A. & Caron, S. J. C. Connectivity patterns that shape olfactory representation in a mushroom body network model. Preprint at bioRxiv https://doi.org/10.1101/2021.02.10.430647 (2021).
Zheng, Z. et al. Structured sampling of olfactory input by the fly mushroom body. Curr. Biol. 32, 3334–3349.e6 (2022).
Guo, C. et al. Purkinje cells directly inhibit granule cells in specialized regions of the cerebellar cortex. Neuron 91, 1330–1341 (2016).
Rowan, M. J. M. et al. Graded control of climbing-fiber-mediated plasticity and learning by inhibition in the cerebellum. Neuron 99, 999–1015.e6 (2018).
Kozareva, V. et al. A transcriptomic atlas of mouse cerebellar cortex comprehensively defines cell types. Nature 598, 214–219 (2021).
Osorno, T. et al. Candelabrum cells are ubiquitous cerebellar cortex interneurons with specialized circuit properties. Nat. Neurosci. 25, 702–713 (2022).
Gao, Z. et al. A cortico-cerebellar loop for motor planning. Nature 563, 113–116 (2018).
Hua, Y., Laserstein, P. & Helmstaedter, M. Large-volume en-bloc staining for electron microscopy-based connectomics. Nat. Commun. 6, 7923 (2015).
Phelps, J. S. et al. Reconstruction of motor control circuits in adult Drosophila using automated transmission electron microscopy. Cell 184, 759–774.e18 (2021).
Deerinck, T. J., Bushong, E. A., Thor, A. & Ellisman, M. H. NCMIR methods for 3D EM: a new protocol for preparation of biological specimens for serial block face scanning electron microscopy. Microscopy 1, 6–8 (2010).
Saalfeld, S., Cardona, A., Hartenstein, V. & Tomancak, P. CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25, 1984–1986 (2009).
Funke, J. et al. Large scale image segmentation with structured loss based deep learning for connectome reconstruction. IEEE Trans. Pattern Anal. Mach. Intell. 41, 1669–1680 (2018).
Sheridan, A. et al. Local shape descriptors for neuron segmentation. Preprint at bioRxiv https://doi.org/10.1101/2021.01.18.427039 (2021).
Falk, T. et al. U-Net: deep learning for cell counting, detection, and morphometry. Nat. Methods 16, 67–70 (2018).
Lee, K., Zung, J., Li, P., Jain, V. & Seung, H. S. Superhuman accuracy on the SNEMI3D connectomics challenge. Preprint at https://doi.org/10.48550/arXiv.1706.00120 (2017).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
Nguyen, T., Malin-Mayor, C., Patton, W. & Funke, J. Daisy: block-wise task dependencies for luigi. https://github.com/funkelab/daisy (2022).
Buhmann, J. et al. Automatic detection of synaptic partners in a whole-brain Drosophila electron microscopy data set. Nat. Methods 18, 771–774 (2021).
Boergens, K. M. et al. webKnossos: efficient online 3D data annotation for connectomics. Nat. Methods 14, 691–694 (2017).
Scheffer, L. K. et al. A connectome and analysis of the adult Drosophila central brain. eLife 9, e57443 (2020).
Ester, M., Kriegel, H.-P., Sander, J. & Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. In Proc. of the Second International Conference on Knowledge Discovery and Data Mining 226–231 (AAAI Press, 1996).
Hamming, R. W. Error detecting and error correcting codes. Bell Syst. Tech. J. 29, 147–160 (1950).
Dhar, M., Hantman, A. W. & Nishiyama, H. Developmental pattern and structural factors of dendritic survival in cerebellar granule cells in vivo. Sci Rep. 8, 17561 (2018).
Tyrrell, T. & Willshaw, D. Cerebellar cortex: its simulation and the relevance of Marr’s theory. Philos. Trans. R. Soc. Lond. B 336, 239–257 (1992).
Gundappa-Sulur, G., De Schutter, E. & Bower, J. M. Ascending granule cell axon: an important component of cerebellar cortical circuitry. J. Comp. Neurol. 408, 580–596 (1999).
Huang, C.-M., Wang, L. & Huang, R. H. Cerebellar granule cell: ascending axon and parallel fiber. Eur. J. Neurosci. 23, 1731–1737 (2006).
Walter, J. T., Dizon, M.-J. & Khodakhah, K. The functional equivalence of ascending and parallel fiber inputs in cerebellar computation. J. Neurosci. 29, 8462–8473 (2009).
Bradbury, J. et al. JAX: composable transformations of Python+NumPy programs. https://github.com/google/jax (2018).
Hennigan, T., Cai, T., Norman, T. & Babuschkin, I. Haiku: Sonnet for JAX. https://github.com/deepmind/dm-haiku (2020).
Babuschkin, I. et al. The DeepMind JAX Ecosystem. http://github.com/deepmind (2020).
Ito, M. The molecular organization of cerebellar long-term depression. Nat. Rev. Neurosci. 3, 896–902 (2002).
Acknowledgements
We thank X. Guan, Y. Hu, M. Liu, E. Mayette, M. Narwani, M. Osman, D. Patel, E. Phalen, R. Singh and K. Yu for neuron reconstruction and manual annotation of ground truth for cell segmentation and synapse predictions; J. Rozowsky for neuron reconstruction, analysis, and the initial finding of redundant MF→GrC connectivity; P. Li and V. Jain for access to and support with Google Armitage/BrainMaps software; Y. Hu and M. Osman for contributing code for segmentation accuracy measurements; M. Narwani, K. Yu and X. Guan for contributing code for the proofreading platform; J. Buhmann and N. Eckstein for discussions and machine learning advice; C. Guo, T. Osorno, S. Rudolph and L. Witter, for discussion, advice, and help with electron microscopy preparations; C. Bolger for electron microscopy sample preparation; T. Ayers, R. Smith and Luxel Corporation for coating GridTape; O. Rhoades for help with illustrations; D. Bock, J. Drugowitsch, A. Handler, C. Ott, R. Wilson, A. Kuan and members of the Lee laboratory for comments on the manuscript. This work was supported by the NIH (R21NS085320 and RF1MH114047 to W.-C.A.L. and R01MH122570 and R35NS097284 to W.G.R.), the Bertarelli Program in Translational Neuroscience and Neuroengineering, Stanley and Theodora Feldberg Fund, and the Edward R. and Anne G. Lefler Center. Portions of this research were conducted on the O2 High Performance Compute Cluster at Harvard Medical School partially provided through NIH NCRR (1S10RR028832-01) and a Foundry Award for the HMS Connectomics Core.
Author information
Authors and Affiliations
Contributions
T.M.N., L.A.T., W.G.R. and W.-C.A.L. conceptualized the project and designed experiments. D.G.C.H. sectioned samples. L.A.T. imaged and aligned the electron microscopy dataset. A.S., T.M.N. and J.F. developed the segmentation and synapse prediction networks. T.M.N. and J.F. developed Daisy. T.M.N. developed the MD-Seg proofreading platform. T.M.N. and L.A.T. developed and applied the segmentation pipeline. T.M.N., J.L.R., I.R. and X.C.Y. developed and applied the synapse prediction pipeline. T.M.N., L.A.T., J.L.R., I.R. and X.C.Y. performed reconstructions and analysed the data. T.M.N., L.A.T., J.L.R., W.G.R. and W.-C.A.L. wrote the paper with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
Harvard University filed a patent application regarding GridTape (WO2017184621A1) on behalf of the inventors including W.-C.A.L. and D.G.C.H., and negotiated licensing agreements with interested partners. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Albert Cardona, Mackenzie Mathis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Similarity between a convolutional neural network and the cerebellar feedforward network.
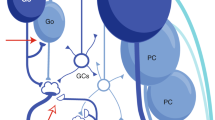
a, Diagram of a simple convolutional neural network with one convolutional layer (input→hidden) and one fully connected layer (hidden→output). The input (left) is made up of a single-channel 2D grid of neurons. The convolutional layer (middle) is made up of neurons each sampling a small local grid of the input (e.g., nine inputs when a 3x3 filter is used, cyan colored circles). This is notably different from a multi-layer perceptron network where the input and the hidden layer are fully connected - the convolution allows an increase in features while decreasing computational cost. Due to the small field of view of each convolutional layer neuron, adjacent neurons share a significant amount of inputs with each other. To increase capacity of the hidden layer, the convolutional neurons can be replicated by n times (typically parameterized as n features). Finally, the output neurons (right) are fully connected with neurons in the preceding convolutional layer. For a classification network, each label (class) is associated with a single binary output neuron for both training and inference. b, Diagram of the cerebellar feedforward network. Mossy fibers (MFs; left) can be considered a 2D grid of sensory and afferent command inputs typically of mixed modalities40,41. Granule cells (GrCs; middle) sample only ~4 MF inputs each. The total number of GrCs is estimated to be hundreds of times more than the number of MFs (Fig. 1b), represented by an expansion factor m. Finally, Purkinje cells (PCs; right) - output neurons of the cerebellar cortex - receive input from tens to hundreds of thousands of GrC axons that pass by PC dendrites.
Extended Data Fig. 2 Automated segmentation and synapse prediction.
a, Serial-section electron microscopy (EM) dataset from lobule V from the cyan boxed region in Fig. 1d. b, The 3D reconstruction segmentation pipeline. (i) EM image data, (ii) boundary affinities, and (iii) automated segmentation output. c, Parallelized volume processing using Daisy. The input dataset is divided into small blocks, on which multiple workers can dynamically query and work.Block completion status and output data are efficiently stored into a persistent database or on disk directly from the workers without going through the centralized scheduler process. d, Example view of targeted neuron reconstruction using merge-deferred segmentation (MD-Seg). Neurons are first segmented as small blocks, and inter-block merge decisions are deferred to proofreaders. This is illustrated by the different colored segments of the displayed neuron. The user interface is based on Neuroglancer, modified to provide the segment "grow" functionality, and to integrate an interface to the database keeping track of neuron name, cell type, completion status, notes, and which agglomeration threshold to use for "growing", as well as searching for neurons based on different criteria and recoloring segments of a single neuron to a single color ("Search DB" and "Color" tabs, not shown). e, Automated segmentation evaluation; plot points denote agglomeration thresholds. Average number of merge and split errors of (n = 9) 6 μm3 test volumes. We used a threshold (star) with 2.33 merges and 27 splits per 6 μm3 for proofreading. f, Automated synapse prediction evaluation; plot points denote connected component thresholds. Precision and recall curve for the synapse inference network. We achieved high synapse prediction accuracy with precision: 95.4% and recall: 92.2%, and an f-score: 93.8% (star).
Extended Data Fig. 3 MF→GrC wiring, convergence, and null models.
a,b, 3D plot of the locations of GrC somas and centers of MF boutons reconstructed in the 320 × 80 × 50 μm subvolume. Blue and orange dots indicate the GrCs and MF boutons respectively in the center 10 μm in the mediolateral axis, as plotted in Fig. 2b. c, Distribution of the number of dendrites per GrC (n = 542). d, Distribution of GrC dendrite lengths (n = 1093). e, Anisotropic positioning of MF bouton→GrC inputs (claws), showing elongated distribution in the dorsal-ventral axis (X) relative to both anterior-posterior (Y) and medio-lateral (Z) axes. Contour lines represent 10% intervals in the distribution. f, MF→GrC random models used for comparison with the reconstructed connectivity. (Methods). g, Similar to Fig. 2c, but with random models from f added. h, Similar to g, but with Radius models of different dendrite lengths. i, Cumulative distribution of MF bouton input redundancy counting the number of GrC pairs sharing 1 MF bouton (similar to Fig. 2c). j, Similar to i, but for 2 MF boutons. k, Similar to i, but for 3 MF boutons. l, Average number of GrC pairs sharing 1, 2, or 3 common MF bouton inputs, comparing reconstructed against the Radius random connectivity model described in d. m, Average sharing of GrCs in the reconstructed network as in l, but normalized to random networks. n, Average fractional distribution of inputs to GrCs from different MF bouton types (categorized as the top-third, middle-third, and bottom third most connected boutons) as a function of GrC sampling size. GrCs were randomly subsampled to produce input composition distributions, with error shadings representing SD. o, Same as Fig. 2e, but with random models from f added.
Extended Data Fig. 4 MF→GrC and GrC→PC synaptic connectivity.
a, Example EM micrographs of MF (red) to GrC (blue) synapses. b, Example EM micrographs of GrC (blue) to PC (green) synapses. c, Distributions of the number of synapses per connection of the two synapse types. We analyzed (n = 9012) MF→GrC and (n = 19761) GrC→PC synapses in one volumetric EM dataset from one animal. The median of MF→GrC is nine synapses, while GrC→PC is one. Over 97% of GrC→PC unitary connections have 1 to 2 synapses, though instances of 3 to 6 synapses per connection, while rare, do occur.
Extended Data Fig. 5 MF collateral axons connectivity to GrCs.
a, 3D rendering of a MF (red) bouton, with an axon collateral making synapses onto a GrC (blue) dendrite near a claw. Asterisks denote the location of synapses. b, Rendering of a MF bouton with no axon collaterals. c, Distribution of number of collaterals per MF bouton (n = 63); a MF bouton can have multiple axon collaterals, and each collateral may or may not make synapses onto GrCs. d, Box plot (25th, 50th and 75th percentiles with whiskers extended to points within 1.5 IQRs) of the number of MF-GrC connections: per MF bouton (n = 63), per axon collateral (n = 8), and per each bouton in a collateral (n = 16). e, Box plot of the number of synapses in each MF-GrC connection: per MF bouton (n = 978), per collateral (n = 28), and per bouton in collateral axons (n = 51). Due to the low frequency of axon collaterals, GrC targets, and the number of synapses per GrC target, it is unlikely that MF axon collaterals to GrCs represent a major route of signal propagation. f, Example of an axon collateral making synapses to a GrC on the trunk, and with more synapses formed on the claw. g, Example of a connection on the trunk of the dendrite, with no claw. h, Joint probability distribution of the synapse location of MF axon collaterals onto GrCs (on claw vs. trunk) and whether or not the dendrite made a claw onto the same MF bouton, or did not have claws (unformed claws). These examples of MF axon collateral connections to GrCs could represent different states of MF→GrC rewiring, supporting the hypothesis that MF→GrC wiring adapts to changing MF input representation6,42. Scale bars are 10 µm.
Extended Data Fig. 6 MF→GrC oversharing and convergence vs null models.
a, GrC input sharing relative to random connectivity. The matrix shows the degree of input sharing between GrCs (centermost n = 550, sorted by soma position dorsoventrally). The color scale for each cell in the matrix uses the z-score (reconstructed # of sharing minus random mean divided by the SD). b, MF bouton output convergence relative to random connectivity. The matrix shows the degree of output convergence between MF boutons (centermost n = 234, sorted by soma position dorsoventrally). The color scale uses the z-score as in a.
Extended Data Fig. 7 Synapse prediction sensitivity analysis.
a, Cumulative distributions of MF bouton input redundancy as in Fig. 2c but across synapse prediction accuracies ranging from 90% to 10%. We artificially added false positives (FPs) and false negatives (FNs) to the network (Extended Data Fig. 2f) to achieve different accuracies (Methods). b, Cumulative distribution of postsynaptic GrCs per MF bouton as in Fig. 2e but across synapse prediction accuracies. As shown in a and b, we found that the results were consistent across models and only changed substantially when the FP/FN rates increased past 60%. We propose two reasons our results are robust across model prediction accuracies. First, MF-GrC connections are typically composed of multiple synapses (10 on average, Extended Data Fig. 4c). Since we used at least 3 synapses as a threshold for determining connectivity, even with significant missing, undetected synapses (eg. 50%), the remaining synapses are still reliable to reflect binary connectivity. Second, random and spurious false positive predictions are unlikely to coincide to cross the 3-synapse threshold. One interesting implication is that strongly selective features do not require perfect synapse prediction. This is consistent with connectomes in Drosophila where synapse prediction accuracy is ~60% and that connections are typically consist of multiple synapses which means that even with significant missing, undetected synapses (eg. 50%), the remaining synapses are sufficiently reliable to indicate connectivity.
Extended Data Fig. 8 GrC→PC wiring and similarity of inputs to PCs.
a, Plot of density GrC axons and GrC→PC connectivity rate as a function of height in the molecular layer between the pial surface and PC layer. Across molecular heights, the average axon density is 3.73 ± 1.23 per μm2 (mean ± SD), and the average connection rate is 49.12 ± 4.39% (mean ± SD). Using these numbers and the average area of PC dendrites, we calculated ~125,000 GrC axons pass through the dendritic arbor of each reconstructed PC. At an average connectivity rate of 49%, only about 60,000 GrC axons were connected to each PC, 3-5× less than typically assumed in models of the cerebellar cortex17,67,74. b, Box plot (25th, 50th and 75th percentiles with whiskers extended to points within 1.5 IQRs) of pairwise Hamming similarity between PC postsynaptic targets of non-local GrC axons, and local GrC axons with different numbers of shared MF bouton inputs. Across local GrCs sharing 0, 1, 2, and 3 MF boutons, Hamming similarity means = p = 0.0001137, Kruskal-Wallis H-test. 0-shared vs 1-shared p = 0.0132, 0-shared vs 3-shared p = 0.00797, 1-shared vs 3-shared p = 0.0186, 2-shared vs 3-shared p = 0.0309, other pairings p > 0.05, Dunn’s post hoc tests, Bonferroni corrected for multiple comparisons. c, 3D rendering of EM reconstructed PCs arbitrarily colored.
Extended Data Fig. 9 MF-GrC-PC simulations.
a, Normalized dimensionality of GrCs as a function of input variability using a continuous model of spike frequency13. Noise was modeled as the degree of variation of spiking frequency across all MF inputs (Methods). b, Modeled learned signal size (Methods) as a function of variability between MF input patterns, comparing pattern separation performance between overrepresented (top-third most connected) and underrepresented (bottom-third) MF boutons. Signal size from the reconstructed network is normalized by the random connectivity model for each population separately. c-d, SNR analyses of modeled MF-GrC networks, measuring noise robustness with (Modeled) or without (Random) redundant oversharing of MF inputs (Fig. 2c, d). SNR was computed across GrC subpopulations ranked by robustness (Methods) at a 40% noise level in c, and across GrC subpopulations and noise levels in d (normalized to SNRs of the "random" model at each noise level and subpopulation). The white box in d denotes the noise level shown in c. Redundant oversharing helps PCs learn more reliably by encoding the most robust signals in a subset of more correlated GrCs. e, Binary GrC→PC selective subsampling increases SNR. Left: PCs (green) randomly subsample GrCs (blue) with MF (red) inputs containing signal (S) or noise (N). Right: PCs connect to GrCs encoding signal-relevant MFs, leading to a higher SNR (Fig. 4d). f, Prediction accuracy of a linear neural network trained on output patterns of the GrCs as a function of MF input variability, comparing performance of MF-GrC networks between models that were fully connected, randomly subsampled with 50% connectivity, and selective subsampled with 50% connectivity, all as a function of MF input variability. g, Dimensionality of the GrC population as a function of the percentage of GrCs randomly removed, normalized to the dimensionality with 100% of the population. h, Prediction accuracy as in f comparing performance of MF-GrC networks between randomly and selectively subsampled models as a function of a percentage of randomly removed GrCs.
Supplementary information
Supplementary Information
This file contains Supplementary Table of Contents; Supplementary Data 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nguyen, T.M., Thomas, L.A., Rhoades, J.L. et al. Structured cerebellar connectivity supports resilient pattern separation. Nature 613, 543–549 (2023). https://doi.org/10.1038/s41586-022-05471-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05471-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.