Abstract
Solid cancers exhibit a dynamic balance between cell death and proliferation ensuring continuous tumour maintenance and growth1,2. Increasing evidence links enhanced cancer cell apoptosis to paracrine activation of cells in the tumour microenvironment initiating tissue repair programs that support tumour growth3,4, yet the direct effects of dying cancer cells on neighbouring tumour epithelia and how this paracrine effect potentially contributes to therapy resistance are unclear. Here we demonstrate that chemotherapy-induced tumour cell death in patient-derived colorectal tumour organoids causes ATP release triggering P2X4 (also known as P2RX4) to mediate an mTOR-dependent pro-survival program in neighbouring cancer cells, which renders surviving tumour epithelia sensitive to mTOR inhibition. The induced mTOR addiction in persisting epithelial cells is due to elevated production of reactive oxygen species and subsequent increased DNA damage in response to the death of neighbouring cells. Accordingly, inhibition of the P2X4 receptor or direct mTOR blockade prevents induction of S6 phosphorylation and synergizes with chemotherapy to cause massive cell death induced by reactive oxygen species and marked tumour regression that is not seen when individually applied. Conversely, scavenging of reactive oxygen species prevents cancer cells from becoming reliant on mTOR activation. Collectively, our findings show that dying cancer cells establish a new dependency on anti-apoptotic programs in their surviving neighbours, thereby creating an opportunity for combination therapy in P2X4-expressing epithelial tumours.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated and/or analysed during this study are included in the Article and its Supplementary Information. Original scans of the kinase activation assay shown in Fig. 1c and the immunoblot experiments shown in Figs. 1d,e, 2a,g, 3a,c,e–g,j,l and 4f and Extended Data Figs. 1a,i, 2b and 3c–i are provided in the Supplementary Information. Tumour measurements for Figs. 1j, 2k, 3p and 4i are included in the Source Data. The FACS gating strategy for Fig. 4e is shown in the Supplementary Information. All oligonucleotide sequences, guide sequences for shRNAs and siRNA sequences are described in Supplementary Tables 1–3. Source data are provided with this paper.
References
Endo, H. & Inoue, M. Dormancy in cancer. Cancer Sci. 110, 474–480 (2019).
Gudipaty, S. A., Conner, C. M., Rosenblatt, J. & Montell, D. J. Unconventional ways to live and die: cell death and survival in development, homeostasis, and disease. Annu. Rev. Cell Dev. Biol. 34, 311–332 (2018).
van Schaik, T. A., Chen, K. S. & Shah, K. Therapy-induced tumor cell death: friend or foe of immunotherapy? Front. Oncol. 11, 678562 (2021).
Diwanji, N. & Bergmann, A. Two sides of the same coin - compensatory proliferation in regeneration and cancer. Adv. Exp. Med. Biol. 1167, 65–85 (2019).
Fonseca, B. D., Smith, E. M., Lee, V. H., MacKintosh, C. & Proud, C. G. PRAS40 is a target for mammalian target of rapamycin complex 1 and is required for signaling downstream of this complex. J. Biol. Chem. 282, 24514–24524 (2007).
Price, D. J., Grove, J. R., Calvo, V., Avruch, J. & Bierer, B. E. Rapamycin-induced inhibition of the 70-kilodalton S6 protein kinase. Science 257, 973–977 (1992).
Oliver Metzig, M. et al. Inhibition of caspases primes colon cancer cells for 5-fluorouracil-induced TNF-α-dependent necroptosis driven by RIP1 kinase and NF-κB. Oncogene 35, 3399–3409 (2016).
Sui, X. et al. JNK confers 5-fluorouracil resistance in p53-deficient and mutant p53-expressing colon cancer cells by inducing survival autophagy. Sci. Rep. 4, 4694 (2014).
Tian, H. et al. A reserve stem cell population in small intestine renders Lgr5-positive cells dispensable. Nature 478, 255–259 (2011).
de Sousa e Melo, F. et al. A distinct role for Lgr5+ stem cells in primary and metastatic colon cancer. Nature 543, 676–680 (2017).
Chen, J. et al. Inosine released from dying or dead cells stimulates cell proliferation via adenosine receptors. Front. Immunol. 8, 504 (2017).
Gregory, C. D. & Pound, J. D. Cell death in the neighbourhood: direct microenvironmental effects of apoptosis in normal and neoplastic tissues. J. Pathol. 223, 177–194 (2011).
Rock, K. L. & Kono, H. The inflammatory response to cell death. Annu. Rev. Pathol. 3, 99–126 (2008).
Greten, F. R. & Grivennikov, S. I. Inflammation and cancer: triggers, mechanisms, and consequences. Immunity 51, 27–41 (2019).
Chen, Q., Sun, L. & Chen, Z. J. Regulation and function of the cGAS-STING pathway of cytosolic DNA sensing. Nat. Immunol. 17, 1142–1149 (2016).
Ahn, J. et al. Inflammation-driven carcinogenesis is mediated through STING. Nat. Commun. 5, 5166 (2014).
Bakhoum, S. F. et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature 553, 467–472 (2018).
Chen, Q. et al. Carcinoma–astrocyte gap junctions promote brain metastasis by cGAMP transfer. Nature 533, 493–498 (2016).
Lemos, H. et al. STING promotes the growth of tumors characterized by low antigenicity via IDO activation. Cancer Res. 76, 2076–2081 (2016).
Paludan, S. R., Reinert, L. S. & Hornung, V. DNA-stimulated cell death: implications for host defence, inflammatory diseases and cancer. Nat. Rev. Immunol. 19, 141–153 (2019).
Haag, S. M. et al. Targeting STING with covalent small-molecule inhibitors. Nature 559, 269–273 (2018).
Di Virgilio, F. & Adinolfi, E. Extracellular purines, purinergic receptors and tumor growth. Oncogene 36, 293–303 (2017).
Burnstock, G. Purinergic signalling: therapeutic developments. Front. Pharmacol. 8, 661 (2017).
Balazs, B. et al. Investigation of the inhibitory effects of the benzodiazepine derivative, 5-BDBD on P2X4 purinergic receptors by two complementary methods. Cell. Physiol. Biochem. 32, 11–24 (2013).
Hansen, M. R., Krabbe, S. & Novak, I. Purinergic receptors and calcium signalling in human pancreatic duct cell lines. Cell. Physiol. Biochem. 22, 157–168 (2008).
Cekic, C. & Linden, J. Purinergic regulation of the immune system. Nat. Rev. Immunol. 16, 177–192 (2016).
Yang, J. W., Zhang, Q. H. & Liu, T. Autophagy facilitates anticancer effect of 5-fluorouracil in HCT-116 cells. J. Cancer Res. Ther. 14, S1141–S1147 (2018).
Zou, Z., Tao, T., Li, H. & Zhu, X. mTOR signaling pathway and mTOR inhibitors in cancer: progress and challenges. Cell Biosci. 10, 31 (2020).
Faller, W. J. et al. mTORC1-mediated translational elongation limits intestinal tumour initiation and growth. Nature 517, 497–500 (2015).
Gupta, J. et al. Dual function of p38alpha MAPK in colon cancer: suppression of colitis-associated tumor initiation but requirement for cancer cell survival. Cancer Cell 25, 484–500 (2014).
Canli, O. et al. Myeloid cell-derived reactive oxygen species induce epithelial mutagenesis. Cancer Cell 32, 869–883 (2017).
Qin, S. et al. Role of HMGB1 in apoptosis-mediated sepsis lethality. J. Exp. Med. 203, 1637–1642 (2006).
Boj, S. F. et al. Organoid models of human and mouse ductal pancreatic cancer. Cell 160, 324–338 (2015).
Pallangyo, C. K., Ziegler, P. K. & Greten, F. R. IKKbeta acts as a tumor suppressor in cancer-associated fibroblasts during intestinal tumorigenesis. J. Exp. Med. 212, 2253–2266 (2015).
Fellmann, C. et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 5, 1704–1713 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Drost, J. et al. Sequential cancer mutations in cultured human intestinal stem cells. Nature 521, 43–47 (2015).
Acknowledgements
We thank H. Kunkel, S. Bösser, K. Mohs, P. Gupta, E. Rudolf and C. Danneil for expert technical assistance as well as the staff at the Animal Facility, the Histology Core Facility and the Flow Cytometry Core Facility at Georg-Speyer-Haus. Work in the laboratory of F.R.G. is supported by institutional funds from Georg-Speyer-Haus, and by the LOEWE Center Frankfurt Cancer Institute financed by the Hessen State Ministry for Higher Education, Research and the Arts (III L 5 - 519/03/03.001 - (0015)), Deutsche Forschungsgemeinschaft (FOR2438: Gr1916/11-1; SFB1292-Project ID: 318346496-TP16; SFB1479-Project ID: 441891347-P02; GRK2336) and the ERC (Advanced Grant PLASTICAN-101021078). The Institute for Tumor Biology and Experimental Therapy, Georg-Speyer-Haus is financed jointly by the German Federal Ministry of Health and the Ministry of Higher Education, Research and the Arts of the State of Hessen.
Author information
Authors and Affiliations
Contributions
M. Schmitt, F.C., J.G. and F.R.G. designed the experiments. M. Schmitt, F.C., J.G., E.E., K.B.K., Y.D. and M. Schewe performed and analysed animal and organoid experiments. A.M.N. performed immunostaining and RT–PCR. M.P. generated mutant organoids and shRNA plasmids and contributed to animal experiments. A.C.K. performed and M.C.A. analysed Seahorse experiments. J.V. initiated the project. M.R. helped with FACS experiments. M.K. helped in generation of Sting-knockout organoids. T.W.B. performed siRNA experiments. V.P. performed ROS analysis and ATP measurements. A.A., T.S., M.C.A. and F.J.d.S. provided essential material. F.R.G. conceptualized and conceived the project. M. Schmitt, J.G. and F.R.G. wrote the manuscript. All of the authors commented on the manuscript draft.
Corresponding author
Ethics declarations
Competing interests
M. Schmitt, J.G. and F.R.G. have filed a patent regarding the use of P2X4 inhibitors in combination with cytotoxic compounds. F.J.d.S. is an employee of Genentech and owns Roche shares. F.R.G. is a consultant for Amazentis, which is a company not related to this study. All other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Sebastien Roger, Louis Vermeulen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 (Related to Figure 1): CRC resistance to 5-FU depends on mTORC1 but not mTORC2.
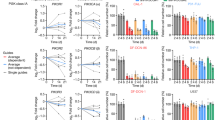
a, Immunoblot analysis of two different human tumor organoid lines derived from sporadic CRC treated as indicated (n = 3). b, Reseeding capacity of hCRC organoid lines treated with 5-FU, rapamycin or 5-FU/rapamycin (n = 3, one of three biological replicates). c, Reseeding capacity of doxycycline-inducible hCRCshRAPTORr#2 tumor organoids treated as indicated (n = 4, one of three biological replicates). d, Raptor qRT-PCR analysis of doxycycline treated hCRCshRAPTOR#2 tumor organoids (n = 3 biological replicates). e, Reseeding capacity of doxycycline-inducible hCRCshRICTOR tumor organoids treated as indicated (n = 4, one of three biological replicates). f, Rictor qRT-PCR analysis of doxycycline treated hCRCshRictor#1 tumor organoids (n=3 biological replicates). g, Reseeding capacity of doxycycline inducible hCRCshRICTOR#2 tumor organoids treated as indicated (n = 4, one of three biological replicates). h, Rictor qRT-PCR analysis of doxycycline treated hCRCshRictor#2 tumor organoids (n=3 biological replicates). i, Immunoblot analysis of Lgr5EGFP-DTR tumor organoids treated as indicated (n = 3). j) Profiles for Extracellular Acidification Rate (ECAR) of AOM/DSSATKN tumor organoids treated as indicated for 24h (n=4 for rapamycin treated organoids in all other conditions n=5 biological replicates of one experiment). k) Profiles for Oxygen Consumption Rate (OCR) of AOM/DSSATKN tumor organoids described in (j) (n = 4 for rapamycin treated organoids in all other conditions n = 5 biological replicates of one experiment). l) Percentage of metabolic potential over baseline in ECAR in organoids described in (j) (n = 4 for rapamycin treated organoids in all other conditions n = 5 biological replicates of one experiment). m) Percentage of metabolic potential over baseline in OCR in organoids treated described in (j) (n = 4 for rapamycin treated organoids in all other conditions n = 5 biological replicates of one experiment). All data are mean ±SD (except for j,k where data are mean ±SEM) and analysed by two-tailed Student’s t-test (d,f,h) or 1-way ANOVA with Bonferroni’s multiple comparison (b, c,e,g,l,m).
Extended Data Fig. 2 (related to Figure 2): Dying tumor cells activate mTORC1 in surrounding tumor cells in vivo.
a, Immunofluorescence analysis of p-S6 (red) and EGFP (Lgr5, green) expression in tumor tissues of vehicle or DT treated AOM/DSS mice 24h after injection. Nuclei were counterstained with DAPI. Representative images are shown (n = 3 biological replicates, scale bar = 50 µm). b, Immunoblot analysis of AOM/DSSATKN tumor organoids treated as indicated. For p-γH2AX and the corresponding α-tubulin loading samples were run on a separate gel (see SI). Representative results are shown (n = 3).
Extended Data Fig. 3 (related to Figure 3): Dying tumor cells activate mTORC1 via ATP/P2X4 in human CRC and PDAC.
a, Relative mRNA expression levels of the indicated genes in the colon tumors from Lgr5EGFP-DTR(+) mice 24h after injection of DT injection or vehicle determined by qRT-PCR. (n = 10 tumors for controls and n = 9 tumors for DT). b, Relative mRNA expression levels of the indicated genes in Lgr5EGFP-DTR(+) tumor organoids at 24h after DT treatment determined by qRT-PCR. Expression of vehicle treated organoids was set to 100 arbitrary units (n = 4 biological replicates). c, Immunoblot analysis of Lgr5EGFP-DTR(+) tumor organoids treated as indicated for 30 min (n = 3). d, Immunoblot analysis of Lgr5EGFP-DTR(+) tumor organoids treated as indicated for 16h (n = 3). e, Immunoblot analysis of Sting WT and Sting KO Lgr5EGFP-DTR(+) tumor organoids treated as indicated (n = 4). f, Immunoblot analysis of mouse (CMT93 and CT26, left panel) and human CRC cells (HCT116 and RKO, right panel) treated as indicated (n = 2). g, Immunoblot analysis of Lgr5EGFP-DTR(+) tumor organoids treated as indicated for 12h (n = 3). h, Left: Immunoblot analysis of ATP (100 µM) treated CT26 cells transfected with control or Rheb siRNA (n = 3). Right: Relative mRNA expression levels of Rheb in siRNA transfected cells determined by qRT-PCR (n = 3 biological replicates). i, Left: Immunoblot analysis of ATP (100 µM) treated CT26 cells transfected with control or a combination of RagA and RagB siRNA (n = 3). Right: Relative mRNA expression levels RagA and RagB in siRNA transfected cells determined by qRT-PCR (n = 3 biological replicates). j, Relative mRNA expression levels of P2x and P2y receptors in mouse colon tumor organoids tissues determined by qRT-PCR (n = 3 biological replicates). k, Immunohistochemistry analysis of P2X4 expression in healthy mouse colon or colon tumors of AOM/DSS treated mice. Images from one representative staining (n = 3, scale bar = 300 µm). l, Reseeding capacity of hCRC tumor organoid line 2 treated as indicated (n = 3, one of three biological replicates). m, Reseeding capacity of doxycycline-inducible hCRCshP2X4#2 tumor organoids treated as indicated (n = 4, one of three biological replicates). n, P24 qRT-PCR analysis of doxycycline treated hCRCshP2X4#2 tumor organoids (n = 3 biological replicates). o, Reseeding capacity of human pancreatic tumor organoids treated as indicated (n = 3, one of three biological replicates). p, Reseeding capacity of human pancreatic tumor organoids treated as indicated (n = 3, one of three biological replicates) All data are mean ±SD and analysed by two-tailed Student’s t-test (a,b,h,i,n,o,p) or by 1-way ANOVA with Bonferroni’s multiple comparison (l,m).
Supplementary information
Supplementary Information
This file contains Supplementary Figs. 1–15 and Tables 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Schmitt, M., Ceteci, F., Gupta, J. et al. Colon tumour cell death causes mTOR dependence by paracrine P2X4 stimulation. Nature 612, 347–353 (2022). https://doi.org/10.1038/s41586-022-05426-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05426-1
This article is cited by
-
ATP released from dying cancer cells stimulates P2X4 receptors and mTOR in their neighbors
Purinergic Signalling (2024)
-
Artesunate attenuates the tumorigenesis of choroidal melanoma via inhibiting EFNA3 through Stat3/Akt signaling pathway
Journal of Cancer Research and Clinical Oncology (2024)
-
Therapeutic targeting of P2X4 receptor and mitochondrial metabolism in clear cell renal carcinoma models
Journal of Experimental & Clinical Cancer Research (2023)
-
Nanomedicine for autophagy modulation in cancer therapy: a clinical perspective
Cell & Bioscience (2023)
-
The role of organoids in cancer research
Experimental Hematology & Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.