Abstract
Protective immunity relies on the interplay of innate and adaptive immune cells with complementary and redundant functions. Innate lymphoid cells (ILCs) have recently emerged as tissue-resident, innate mirror images of the T cell system, with which they share lineage-specifying transcription factors and effector machinery1. Located at barrier surfaces, ILCs are among the first responders against invading pathogens and thus could potentially determine the outcome of the immune response2. However, so far it has not been possible to dissect the unique contributions of ILCs to protective immunity owing to limitations in specific targeting of ILC subsets. Thus, all of the available data have been generated either in mice lacking the adaptive immune system or with tools that also affect other immune cell subsets. In addition, it has been proposed that ILCs might be dispensable for a proper immune response because other immune cells could compensate for their absence3,4,5,6,7. Here we report the generation of a mouse model based on the neuromedin U receptor 1 (Nmur1) promoter as a driver for simultaneous expression of Cre recombinase and green fluorescent protein, which enables gene targeting in group 2 ILCs (ILC2s) without affecting other innate and adaptive immune cells. Using Cre-mediated gene deletion of Id2 and Gata3 in Nmur1-expressing cells, we generated mice with a selective and specific deficiency in ILC2s. ILC2-deficient mice have decreased eosinophil counts at steady state and are unable to recruit eosinophils to the airways in models of allergic asthma. Further, ILC2-deficient mice do not mount an appropriate immune and epithelial type 2 response, resulting in a profound defect in worm expulsion and a non-protective type 3 immune response. In total, our data establish non-redundant functions for ILC2s in the presence of adaptive immune cells at steady state and during disease and argue for a multilayered organization of the immune system on the basis of a spatiotemporal division of labour.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The RNA-sequencing data are deposited in the Gene Expression Omnibus (GEO) repository database under the accession numbers GSE213424, GSE212011 and GSE212940. Source data are provided with this paper.
Code availability
Data were analysed using the standard Seurat 4.0.1 or 4.0.2 pipeline, or with the stated variations. User scripts will be shared upon request.
References
Vivier, E. et al. Innate lymphoid cells: 10 years on. Cell 174, 1054–1066 (2018).
Klose, C. S. & Artis, D. Innate lymphoid cells as regulators of immunity, inflammation and tissue homeostasis. Nat. Immunol. 17, 765–774 (2016).
Vely, F. et al. Evidence of innate lymphoid cell redundancy in humans. Nat. Immunol. 17, 1291–1299 (2016).
Rankin, L. C. et al. Complementarity and redundancy of IL-22-producing innate lymphoid cells. Nat. Immunol. 17, 179–186 (2016).
Colonna, M. Innate lymphoid cells: diversity, plasticity, and unique functions in immunity. Immunity 48, 1104–1117 (2018).
Sonnenberg, G. F. & Hepworth, M. R. Functional interactions between innate lymphoid cells and adaptive immunity. Nat. Rev. Immunol. 19, 599–613 (2019).
Orange, J. S. et al. Comment on: evidence of innate lymphoid cell redundancy in humans. Nat. Immunol. 19, 788–789 (2018).
Klose, C. S. N. et al. The neuropeptide neuromedin U stimulates innate lymphoid cells and type 2 inflammation. Nature 549, 282–286 (2017).
Wallrapp, A. et al. The neuropeptide NMU amplifies ILC2-driven allergic lung inflammation. Nature 549, 351–356 (2017).
Cardoso, V. et al. Neuronal regulation of type 2 innate lymphoid cells via neuromedin U. Nature 549, 277–281 (2017).
Robinette, M. L. et al. Transcriptional programs define molecular characteristics of innate lymphoid cell classes and subsets. Nat. Immunol. 16, 306–317 (2015).
Kondo, M., Weissman, I. L. & Akashi, K. Identification of clonogenic common lymphoid progenitors in mouse bone marrow. Cell 91, 661–672 (1997).
Klose, C. S. N. et al. Differentiation of type 1 ILCs from a common progenitor to all helper-like innate lymphoid cell lineages. Cell 157, 340–356 (2014).
Constantinides, M. G. et al. PLZF expression maps the early stages of ILC1 lineage development. Proc. Natl Acad. Sci. USA 112, 5123–5128 (2015).
Hoyler, T. et al. The transcription factor GATA-3 controls cell fate and maintenance of type 2 innate lymphoid cells. Immunity 37, 634–648 (2012).
Halim, T. Y. et al. Retinoic-acid-receptor-related orphan nuclear receptor alpha is required for natural helper cell development and allergic inflammation. Immunity 37, 463–474 (2012).
Moro, K. et al. Innate production of TH2 cytokines by adipose tissue-associated c-Kit+Sca-1+ lymphoid cells. Nature 463, 540–544 (2010).
Furusawa, J. et al. Critical role of p38 and GATA3 in natural helper cell function. J. Immunol. 191, 1818–1826 (2013).
Klein Wolterink, R. G. et al. Essential, dose-dependent role for the transcription factor Gata3 in the development of IL-5+ and IL-13+ type 2 innate lymphoid cells. Proc. Natl Acad. Sci. USA 110, 10240–10245 (2013).
Nussbaum, J. C. et al. Type 2 innate lymphoid cells control eosinophil homeostasis. Nature 502, 245–248 (2013).
Kopf, M. et al. IL-5-deficient mice have a developmental defect in CD5+ B-1 cells and lack eosinophilia but have normal antibody and cytotoxic T cell responses. Immunity 4, 15–24 (1996).
Kim, K. et al. Terminally differentiating eosinophils express neutrophil primary granule proteins as well as eosinophil-specific granule proteins in a temporal manner. Immune Netw. 17, 410–423 (2017).
Zhou, L. et al. Innate lymphoid cells support regulatory T cells in the intestine through interleukin-2. Nature 568, 405–409 (2019).
Cupedo, T. et al. Human fetal lymphoid tissue-inducer cells are interleukin 17-producing precursors to RORC+CD127+ natural killer-like cells. Nat. Immunol. 10, 66–74 (2009).
Cella, M. et al. A human natural killer cell subset provides an innate source of IL-22 for mucosal immunity. Nature 457, 722–725 (2009).
Takatori, H. et al. Lymphoid tissue inducer-like cells are an innate source of IL-17 and IL-22. J. Exp. Med. 206, 35–41 (2009).
Sanos, S. L. et al. RORγt and commensal microflora are required for the differentiation of mucosal interleukin 22-producing NKp46+ cells. Nat. Immunol. 10, 83–91 (2009).
Satoh-Takayama, N. et al. Microbial flora drives interleukin 22 production in intestinal NKp46+ cells that provide innate mucosal immune defense. Immunity 29, 958–970 (2008).
Zheng, Y. et al. Interleukin-22 mediates early host defense against attaching and effacing bacterial pathogens. Nat. Med. 14, 282–289 (2008).
Harris, N. L. & Loke, P. Recent advances in type-2-cell-mediated immunity: insights from helminth infection. Immunity 47, 1024–1036 (2017).
Tsou, A. M. et al. Neuropeptide regulation of non-redundant ILC2 responses at barrier surfaces. Nature https://doi.org/10.1038/s41586-022-05297-6 (2022).
Niola, F. et al. Id proteins synchronize stemness and anchorage to the niche of neural stem cells. Nat. Cell Biol. 14, 477–487 (2012).
Grote, D., Souabni, A., Busslinger, M. & Bouchard, M. Pax 2/8-regulated Gata 3 expression is necessary for morphogenesis and guidance of the nephric duct in the developing kidney. Development 133, 53–61 (2006).
Luche, H., Weber, O., Nageswara Rao, T., Blum, C. & Fehling, H. J. Faithful activation of an extra-bright red fluorescent protein in “knock-in” Cre-reporter mice ideally suited for lineage tracing studies. Eur. J. Immunol. 37, 43–53 (2007).
Halim, T. Y. F. et al. Group 2 innate lymphoid cells are critical for the initiation of adaptive T helper 2 cell-mediated allergic lung inflammation. Immunity 40, 425–435 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Ferreira-Gomes, M. et al. SARS-CoV-2 in severe COVID-19 induces a TGF-β-dominated chronic immune response that does not target itself. Nat. Commun. 12, 1961 (2021).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
Meyerholz, D. K., Griffin, M. A., Castilow, E. M. & Varga, S. M. Comparison of histochemical methods for murine eosinophil detection in an RSV vaccine-enhanced inflammation model. Toxicol. Pathol. 37, 249–255 (2009).
Acknowledgements
We thank C. Braeuning, T. Borodina, T. Conrad and the other members of the Genomics Platform at MDC Berlin–Mitte for carrying out the RNA-seq experiments; A. Branco and M. Fernandes of the Benjamin Franklin Flow Cytometry Facility for cell sorting (supported by DFG Instrument Grants INST 335/597-1 FUGG and INST 335/777-1 FUGG); J. Fehling (Rosa26LSL-RFP), A. Lasorella (Id2fl/fl) and M. Busslinger (Gata3fl/fl) for providing mice; L. Paul for technical assistance; I. Mattiola for discussion and M. Witkowski for reading the manuscript. This work was supported by grants from the European Research Council Starting Grant (ERCEA; 803087 to C.S.N.K.), the German Research Foundation (DFG; Project-ID 259373024—CRC/TRR 167, FOR2599 project 5, B05—CRC/TRR 241, —KL 2963/5-2, SPP1937—KL 2963/2-1 and KL 2963/3-1 to C.S.N.K., SPP1937 DI764/9-2; SFB-TRR156/A02; SFB-TRR241/A01; SFB1444/P11 to A.D., SPP1937 DU1295/4-1 to C.U.D, TRR241 project Z03 and SFB1340 project B06 to A.A.K.), the Swiss National Science foundation (grant ID: 184425 to M.O.J.), the Leibniz Association (Leibniz Collaborative Excellence, TargArt) to C.R. and M.-F.M., the Crohn’s and Colitis Foundation Research Fellowship Award (award no. 527125 to A.M.T., award no. 851136 to M.A., and award no. 937437 to H.Y.), the WCM Department of Pediatrics Junior Faculty Pilot Award, and The Jill Roberts Center Pilot Award for Research in IBD (to A.M.T.); the Sackler Brain and Spine Institute Research (to C.C.), the US National Institutes of Health (DK126871, AI151599, AI095466, AI095608, AI142213, AR070116, AI172027 and DK132244 to D.A.), LEO foundation, Cure for IBD, Jill Roberts Institute, the Sanders Family, the Rosanne H. Silbermann Foundation (all to D.A.) and the state of Berlin and the European Regional Development Fund to M.-F.M. (ERDF 2014-2020, EFRE 1.8/11, 563 Deutsches Rheuma-Forschungszentrum).
Author information
Authors and Affiliations
Contributions
K.J.J., P.M.T., M.O.J., H.Y., M.A., X.G. and C.S.N.K. carried out most of the experiments and analysed the data. S.B., V.S.-T., P.L., A.P., Z.A.R., A.S., A.M.T., C.C., C.S., S.H. and A.A.K. helped with performing the experiments. A.I., D.B., F.R.H., G.M.G., P.D. and M.-F.M. performed RNA-sequencing analysis. C.U.D., C.R., A.D. and D.A. provided crucial input and tools for the study. K.J.J. and C.S.N.K. conceived the project and wrote the manuscript with input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
D.A. has contributed to scientific advisory boards at Pfizer, Takeda, FARE and the KRF. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Pentao Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 (related to Fig 1): Gating strategy for flow cytometry.
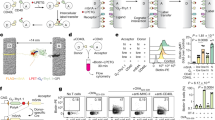
a, Generation of genetic construct of the Nmur1iCre–eGFP BAC transgene. 70th bp of exon 2 to 29th bp downstream of the TGA stop codon of exon 3 was replaced by a cassette containing iCre and eGFP linked with P2A and T2A. (86,388,138-86,386,296) were replaced through homologous recombination. The FRT-flanked kanamycin selection cassette was deleted after FLP-mediated recombination. Removal of LoxP and LoxP511 sites on BAC vector backbone. b, Expression of the Nmur1 gene in NMUR1–eGFP+ and NMUR1–eGFP− cells purified by FACsorting from the intestinal lamina propria. c, Gating strategy of bone marrow precursors. Lineage: CD3, CD5, CD19, Ly6G, FcεRIa. d, Gating strategy of intestinal lymphoid populations. e, Gating strategy of intestinal myeloid populations. Lineage: CD3, CD5, CD19, FcεRIa. f, Gating strategy of intestinal eosinophils, 2 alternative gates shown. Lineage: CD3, CD5, CD19, Ly6G. g, Gating strategy of γδ T cells, regulatory T cells (Treg) and TH2 cells from the mesenteric lymph node. h, Gating strategy of mast cells and basophils from the peritoneal lavage. Lineage: CD3, CD5, CD19, Ly6G.
Extended Data Fig. 2 (related to Fig 1): Nmur1iCre-eGFP specifically labels ILC2s in steady state and during type 2 inflammation.
a, Flow cytometry plots of GFP and ST2 in lung ILC2s (left) and TH2 cells (right) of Nmur1iCre-eGFP mice. Numbers denote percentage of cells inside the gate. b, GFP expression in TH2 cells and ILC2s of Nmur1iCre-eGFP mice, quantification of a. c, GFP expression in mLN CD45− cells (black) and ILC2s (gray). d, Quantification of c. e, Quantification of GFP expression across organs and cell types as displayed in the heat map of Fig. 1d. f, RFP and ST2 in lung ILC2s (left) and TH2 cells (right) of Nmur1iCre-eGFP x Rosa26LSL-RFP F1 offspring. Numbers denote percentage of cells inside the gate. g, RFP expression in TH2 cells and ILC2s, quantification of f. h, RFP expression in mLN CD45− cells (black) and ILC2s (gray) of Nmur1iCre-eGFP x Rosa26LSL-RFP F1 offspring. i, Quantification of h. j, RFP expression across organs and cell types as displayed in the heat map of Fig. 1h. k, Heat map of flow-cytometric quantification of GFP and RFP expression in TH2 and Treg cells. Values represent median percentage of positive cells of parental gate, n = 3 mice per group. l–q, Immunofluorescence micrographs of RFP expression in Nmur1iCre-eGFP x Rosa26LSL-RFP F1 offspring stained for alpha smooth muscle actin (α-SMA), CD45 and DAPI in the small intestine (l), mesenteric lymph nodes (m), lung (n), mesenteric fat (o), femoral muscle (p) and large intestine (q). r, GFP and Cre (RFP) expression seven days after Nippostrongylus brasiliensis (N.b.) helminth infection across immune cell populations in Nmur1iCre-eGFP x Rosa26LFL-RFP F1 offspring. s,t, GFP (s) and RFP (t) expression in Nmur1iCre-eGFP Rosa26LSL-RFP mice, seven days after infection with N.b.. Data in (r-t) are representative of two independent experiments with 3 mice per group.
Extended Data Fig. 3 (related to Fig 2): Nmur1iCre-eGFP Gata3fl/fl and Nmur1iCre-eGFP Id2fl/fl mice lack ILC2s in all tissues investigated.
a, Flow cytometry plots of ILC2s gated from live CD45+ Lin− (CD3, CD5, CD19) cells as CD127+ KLRG1+ or CD127+ ST2+ across organs of Nmur1iCre-eGFP Gata3fl/fl mice and littermate controls. Numbers denote percentage of cells inside the gate. b, Quantification of a. Mean ± s.d., Student’s t-test. c-d, Heat maps displaying Log2(fold change) of the median cell frequencies among live CD45+ cells across different populations and organs of Nmur1iCre-eGFP Gata3fl/fl (c) and Nmur1iCre-eGFP Id2fl/fl (d) mice, compared with the respective littermate controls. Results are representative of two independent experiments with 3-4 mice per group. Littermate controls for a-c were Gata3fl/+ or Nmur1iCre-eGFP Gata3fl/+, for d Id2fl/+ or Nmur1iCre-eGFP Id2fl/+. e, Quantification of relative TH2 (top) and Treg (bottom) numbers in Nmur1iCre-eGFP Id2fl/fl mice and Id2fl/fl littermate controls. Results are representative of three independent experiments with 3–6 mice per group. Mean ± s.d., Student’s t-test. f, Cytokine secretion by cultured Nmur1iCre-eGFP Id2fl/fl immune cells isolated from lung and intestine, stimulated with IL-2, IL-7, IL-25 and IL-33 or PMA/Ionomycin for 8 h, measured in the supernatant by cytometric bead assay. Mean ± s.d., two-way ANOVA with Šídák‘s multiple comparison tests, data are representative of two independent experiments with 3-4 mice per group. g, Serum IL-5 in untreated Id2fl/fl and Nmur1iCre-eGFP Id2fl/fl animals as determined by the BD Cytometric Bead Array Mouse IL-5 Enhanced Sensitivity Flex Set, Student’s t-test. NS not significant, *p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001. a-g One symbol represents data from one mouse.
Extended Data Fig. 4 (related to Fig 3): ILC2s regulate eosinophils in tissue.
Analysis of eosinophils across organs in the steady state. a, Eosinophils were gated from live CD45+ Lin− (CD3, CD5, CD19, Ly6G) cells using SiglecF and CCR3, across organs of Nmur1iCre-eGFP Id2fl/fl mice and littermate controls. b, Relative ILC2 and eosinophil numbers in bone marrow chimera of Nmur1iCre-eGFP Id2fl/fl (KO) and Id2fl/fl (WT) recipients transplanted with bone marrow of Nmur1iCre-eGFP Id2fl/fl (KO) or Id2fl/fl (WT) donors. c, Quantification of eosinophils in IL5−/− mice and respective IL5+/− littermate controls. d, Flow cytometry plots of granulocyte/macrophage progenitor (GMP) gated from live CD45+ Lin− (CD3, CD5, CD19, Gr1, B220) SCA-1− c-Kithi cells as CD34+ CD16/CD32hi and relative quantification in bone marrow of Nmur1iCre-eGFP Id2fl/fl mice and Id2fl/fl littermate controls. e, Quantification of relative Eosinophil precursors (EoP) and GMPs in the bone marrow of IL-5−/− compared with WT controls. f, Flow-cytometric quantification of relative numbers of bone marrow-derived eosinophils gated from live CD45+ Lin− (CD3, CD5, CD19) CD11b+ cells as SSChi and SiglecF+. BM cells were cultured with either SCF and FLT3L, IL-5, ILC2P or supernatant from ILC2s for 10 days. g,h Mean fluorescence intensity of CCR3 on eosinophils in Nmur1Cre-T2A-GFP Id2flox/flox (g) and Il5−/− (h) mice and respective littermate controls. b-e, g-h One symbol represents data from one mouse (b-e, g-h) or one well (f), data are mean ± s.d., c—e, g-h Student‘s t-test. b, f One-way ANOVA with Tukey’s multiple comparisons, NS not significant, * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Extended Data Fig. 5 (related to Fig 3): ILC2 deletion regulates myeloid and epithelial populations in the lung.
a, Bubble plot of marker genes of global cell populations in single-cell RNA-sequencing of myeloid and epithelial cells in the steady state lung of Nmur1iCre-eGFP Id2fl/fl mice and littermates. b, Marker genes of subclusters of interstitial macrophages. c, Marker genes of DC subclusters. d, Marker genes in subclusters of alveolar type 2 epithelial cells. e—g, Subclusters in e macrophages, f dendritic cells and g alveolar type 2 epithelial cells. h, Macrophage UMAPs color-coded gray-to-red with genes downregulated in Macrophages of Nmur1iCre-eGFP Id2fl/fl mice. i, Macrophage UMAPs color-coded with genes upregulated in macrophages of Nmur1iCre-eGFP Id2fl/fl mice. j, DC UMAP color-coded by Ccl24 expression. k, UMAPs of alveolar type 2 epithelial cells color-coded with genes downregulated in alveolar type 2 epithelial cells in Nmur1iCre-eGFP Id2fl/fl mice. l, Statistical evaluation of genes displayed in f—h. Cells from four mice per group were pooled. Littermate controls for (a—l) were Id2fl/fl or Id2fl/+.
Extended Data Fig. 6 (related to Fig 3): ILC2s regulate myeloid cells in type-2 inflammatory allergy models of the lung.
a—d, Flow-cytometric quantification of cell population in Nmur1iCre-eGFP Id2fl/fl mice and littermate controls sensitized with A. alternata extract for three consecutive days, analysed 7 days after the first dose. a, Absolute numbers of lung ILC2s, lung and BAL eosinophils and total CD45+ cells in the BAL. b, Relative and absolute numbers of TH2 cells in the lung. c,d, relative (c) and absolute (d) numbers of myeloid populations from the lung interstitium, and alveolar macrophages from the BAL. Data in a—d are representative of two independent experiments with 4–6 animals per group. e—i, Analysis of Nmur1iCre-eGFP Id2fl/fl mice and littermate controls sensitized with papain for three consecutive days, analysed 7 days after the first dose. e, Absolute numbers of ILC2s in the lung interstitium, and eosinophils in the lung interstitium and BAL. f, Eosinophils are visualized by modified Sirius red stain in the lung. g, Relative and absolute numbers of TH2 cells in the lung. h,i, relative (h) and absolute (i) numbers of myeloid populations from the lung interstitium, and alveolar macrophages from the BAL. Data in e and g-i are pooled from two independent experiments with 4-5 mice per group. Untreated controls for (e-i) were Id2fl/fl, papain-treated littermate controls were Id2fl/fl, Id2fl/+ or Nmur1iCre-eGFP Id2fl/+. One symbol represents data from one mouse, data are mean ± s.d., One-way ANOVA with Tukey’s multiple comparison tests, NS: not significant, * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001. BAL, bronchoalveolar lavage.
Extended Data Fig. 7 (related to Fig 3): ILC2s regulate myeloid cells in type-2 inflammatory allergy models of the lung.
a—e, Analysis of Nmur1iCre-eGFP Id2fl/fl mice (cKO) and littermate controls (Ctrl) sensitized and challenged with house dust mite. a, Lung ILC2s (top), BAL eosinophils (middle) and Lung TH2 cells (bottom) were gated from live CD45+ Lin− (CD11b−CD11c− FcεR1a−B220−CD3−5−) cells. The second gate in the eosinophil plots marks alveolar macrophages. b, Quantification of a. c, Periodic-acid-Schiff staining of lungs. d, Gating of cytokine-expressing T helper cells isolated from the lung. e, Quantification of d. f-j, Resident and inflammatory eosinophils from the lungs of papain- and A. alternata-treated mice. f, Flow cytometry plots showing resident and inflammatory eosinophils from lung after Papain treatment of Nmur1iCre-eGFP Id2fl/fl mice (cKO) and littermate controls (Ctrl). g, Relative and absolute numbers of resident and inflammatory eosinophils. h, Flow cytometry plots showing resident and inflammatory eosinophils from lung after A. alternata treatment of Nmur1iCre-eGFP Id2fl/fl mice (cKO) and littermate controls (Ctrl). i, Relative and absolute numbers of resident and inflammatory eosinophils. Data are representative of two independent experiments with 3–5 mice per group (f-i). Symbols represent data from one mouse, data are mean ± s.d., One-way ANOVA with Tukey’s multiple comparisons (b—e) or Student’s t-test (g,i), NS not significant, * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Extended Data Fig. 8 (related to Fig 3): Eosinophils are altered in ILC2-deficient mice.
a—f, Flow-cytometric analysis of eosinophils and ILC2s in the BAL and lung interstitium of Id2fl/fl, Nmur1iCre-eGFP Id2fl/fl mice treated with either PBS or recombinant IL-5, seven days after allergy induction by intranasal application of A. alternata extract. a,b Eosinophils in the lung interstitium, c,d Eosinophils in the BAL, e,f ILC2s in the lung interstitium. Symbols represent data from one mouse, data are pooled from two independent experiments. Data are mean ± s.d., Student’s t-test, NS not significant, * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001. g Flow cytometry plots showing the gating strategy for the analysis and sorting of eosinophils from lung after Papain treatment of Nmur1iCre-eGFP Id2fl/fl mice (cKO) and littermate controls (Ctrl). Data are representative of one experiment with 5 mice per group. h, Pathways from tmod gene set enrichment analysis (GSEA) of bulk RNA sequencing from eosinophils of 4 Nmur1iCre-eGFP Id2fl/fl mice vs 4 WT control animals. Bars are color-coded according to Databases: Gene Ontology (GO, red), Reactome (gray), Hallmark (white), tmod (purple), KEGG (pink). Genes differentially expressed in eosinophils after papain treatment of Nmur1iCre-eGFP Id2fl/fl mice (cKO) and littermate controls (Ctrl) were used for the analysis. i, Volcano plot showing differentially expressed genes of interest in eosinophils after papain treatment of Nmur1iCre-eGFP Id2fl/fl mice (cKO) and littermate controls (Ctrl) j, Heat maps of genes differentially expressed in eosinophils after papain treatment of Nmur1iCre-eGFP Id2fl/fl mice (cKO) and littermate controls (Ctrl). Each heat map corresponds to one pathway shown in h.
Extended Data Fig. 9 (related to Fig 4): ILC2-deficient mice are highly susceptible to N. brasiliensis infection.
7 (a-h) or 11 days (i—m) after infection with N.b., Nmur1iCre-GFP Gata3fl/fl (a,b,f-h) or Nmur1iCre-eGFP Id2fl/fl (c-e, i—m) mice (cKO) and littermate controls (Ctrl) were analysed alongside untreated WT controls. a,b, Nmur1iCre-GFP Gata3fl/fl mice (cKO) and littermate controls (Ctrl) 7 days after N. b. infection alongside untreated WT controls. ILC2s (a) and eosinophils (b) in the mLN and lung, flow cytometry plots depicting mLN. c, TH2 cells in mLN and lung. d,e Neutrophils in the lung. Littermate controls for c—e were Id2fl/+ or Nmur1iCre-eGFP Id2fl/+. f, Worm burden g, Goblet- and tuft cells per villus, mucus area quantified from UEA-I stain, all as shown in h. h, left: Periodic acid-Schiff staining; middle: intestinal tuft cell staining with Dclk1 (red), DAPI (blue); right: intestinal mucus stained with UEA-I (green), DAPI (blue). Littermate controls for a,b,f-h were Gata3fl/+. i,j, ILC2s and eosinophils in the mLN (i) or mLN and lung (j) 11 days after infection. k, TH2 cells. l, Goblet- and tuft cells per villus, mucus area quantified from UEA-I stain as shown in m. m, intestine as shown in h, 11 days after infection. n-r Infection of BM chimeras of C57BL/6 recipients transplanted with bone marrow of Id2fl/fl (Ctrl), Nmur1iCre-eGFP Id2fl/fl (cKO) donors, or bone marrow of cKO donors supplemented with sorted ILC2 precursors (cKO + ILC2p) from C57BL/6 donors, with N.b. n, Periodic acid-Schiff staining of small intestine seven days after N. b. infection. o, Goblet cells per villus. p,q, Neutrophils in the lung. r, TH2 cells. s, ILC3s, LTi-like ILC3s and NCR+ ILC3s 7 days after N.b. infection of Nmur1iCre-eGFP Id2fl/fl mice (cKO) and littermate controls (Ctrl). a—s One symbol represents data from one mouse, data are mean ± s.d. one-way ANOVA with Tukey’s multiple comparisons, NS not significant, * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Extended Data Fig. 10 (related to Fig 4): Gene expression in epithelial cells of ILC2-deficient mice during N. brasiliensis infection.
a, Heat map showing scaled expression of marker genes for colonic cell populations in single-cellRNA sequencing Data shown in Fig. 4m. b, Violin plots showing normalized count distributions of differentially expressed genes Reg3b, Reg3g, Muc-2, Clca1 and Ang4 in colonic epithelial cells between infected Nmur1iCre-eGFP Id2fl/fl mice (cKO) and littermate controls (Ctrl) in all clusters.
Supplementary information
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jarick, K.J., Topczewska, P.M., Jakob, M.O. et al. Non-redundant functions of group 2 innate lymphoid cells. Nature 611, 794–800 (2022). https://doi.org/10.1038/s41586-022-05395-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05395-5
This article is cited by
-
Group 3 innate lymphoid cells in intestinal health and disease
Nature Reviews Gastroenterology & Hepatology (2024)
-
Cooperation of ILC2s and TH2 cells in the expulsion of intestinal helminth parasites
Nature Reviews Immunology (2024)
-
Pinning down unique ILC2 functions
Nature Reviews Immunology (2022)
-
Neuropeptide regulation of non-redundant ILC2 responses at barrier surfaces
Nature (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.