Abstract
CD4+ T cell differentiation requires metabolic reprogramming to fulfil the bioenergetic demands of proliferation and effector function, and enforce specific transcriptional programmes1,2,3. Mitochondrial membrane dynamics sustains mitochondrial processes4, including respiration and tricarboxylic acid (TCA) cycle metabolism5, but whether mitochondrial membrane remodelling orchestrates CD4+ T cell differentiation remains unclear. Here we show that unlike other CD4+ T cell subsets, T helper 17 (TH17) cells have fused mitochondria with tight cristae. T cell-specific deletion of optic atrophy 1 (OPA1), which regulates inner mitochondrial membrane fusion and cristae morphology6, revealed that TH17 cells require OPA1 for its control of the TCA cycle, rather than respiration. OPA1 deletion amplifies glutamine oxidation, leading to impaired NADH/NAD+ balance and accumulation of TCA cycle metabolites and 2-hydroxyglutarate—a metabolite that influences the epigenetic landscape5,7. Our multi-omics approach revealed that the serine/threonine kinase liver-associated kinase B1 (LKB1) couples mitochondrial function to cytokine expression in TH17 cells by regulating TCA cycle metabolism and transcriptional remodelling. Mitochondrial membrane disruption activates LKB1, which restrains IL-17 expression. LKB1 deletion restores IL-17 expression in TH17 cells with disrupted mitochondrial membranes, rectifying aberrant TCA cycle glutamine flux, balancing NADH/NAD+ and preventing 2-hydroxyglutarate production from the promiscuous activity of the serine biosynthesis enzyme phosphoglycerate dehydrogenase (PHGDH). These findings identify OPA1 as a major determinant of TH17 cell function, and uncover LKB1 as a sensor linking mitochondrial cues to effector programmes in TH17 cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RNA-sequencing and ATAC–seq data have been deposited in the Gene Expression Omnibus (GEO) as the superseries GSE207603. This superseries contains RNA-sequencing datasets with accession numbers GSE156742 and GSE207601 and a single ATAC–seq dataset with accession number GSE207602. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository, with the dataset identifier PXD036162. Source data are provided with this paper.
Code availability
LC–MS and GC–MS metabolomics data were analysed using R code developed in-house, which is publicly available at https://gitlab.gwdg.de/joerg.buescher/metabolomics_scripts.
References
Buck, M. D., O'Sullivan, D. & Pearce, E. L. T cell metabolism drives immunity. J. Exp. Med. 212, 1345–1360 (2015).
Bailis, W. et al. Distinct modes of mitochondrial metabolism uncouple T cell differentiation and function. Nature 571, 403–407 (2019).
Xu, T. et al. Metabolic control of TH17 and induced Treg cell balance by an epigenetic mechanism. Nature 548, 228–233 (2017).
Giacomello, M., Pyakurel, A., Glytsou, C. & Scorrano, L. The cell biology of mitochondrial membrane dynamics. Nat. Rev. Mol. Cell Biol. 21, 204–224 (2020).
Martinez-Reyes, I. & Chandel, N. S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 11, 102 (2020).
Cogliati, S. et al. Mitochondrial cristae shape determines respiratory chain supercomplexes assembly and respiratory efficiency. Cell 155, 160–171 (2013).
Ye, D., Guan, K. L. & Xiong, Y. Metabolism, activity, and targeting of d- and l-2-hydroxyglutarates. Trends Cancer 4, 151–165 (2018).
Chan, D. C. Mitochondrial dynamics and its involvement in disease. Annu. Rev. Pathol. 15, 235–259 (2020).
Youle, R. J. & van der Bliek, A. M. Mitochondrial fission, fusion, and stress. Science 337, 1062–1065 (2012).
Matilainen, O., Quiros, P. M. & Auwerx, J. Mitochondria and epigenetics—crosstalk in homeostasis and stress. Trends Cell Biol. 27, 453–463 (2017).
Herkenne, S. et al. Developmental and tumor angiogenesis requires the mitochondria-shaping protein Opa1. Cell Metab. 31, 987–1003.e1008 (2020).
Song, Z., Chen, H., Fiket, M., Alexander, C. & Chan, D. C. OPA1 processing controls mitochondrial fusion and is regulated by mRNA splicing, membrane potential, and Yme1L. J. Cell Biol. 178, 749–755 (2007).
Ishihara, N., Fujita, Y., Oka, T. & Mihara, K. Regulation of mitochondrial morphology through proteolytic cleavage of OPA1. EMBO J. 25, 2966–2977 (2006).
MacVicar, T. & Langer, T. OPA1 processing in cell death and disease—the long and short of it. J. Cell Sci. 129, 2297–2306 (2016).
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V. K. IL-17 and Th17 Cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Wagner, A. et al. Metabolic modeling of single Th17 cells reveals regulators of autoimmunity. Cell 184, 4168–4185.e4121 (2021).
Ivanov, I. I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498 (2009).
Sano, T. et al. An IL-23R/IL-22 circuit regulates epithelial serum amyloid A to promote local effector Th17 responses. Cell 163, 381–393 (2015).
Hirota, K. et al. Fate mapping of IL-17-producing T cells in inflammatory responses. Nat. Immunol. 12, 255–263 (2011).
Esplugues, E. et al. Control of TH17 cells occurs in the small intestine. Nature 475, 514–518 (2011).
Birsoy, K. et al. Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides. Nature 508, 108–112 (2014).
Arguello, R. J. et al. SCENITH: a flow cytometry-based method to functionally profile energy metabolism with single-cell resolution. Cell Metab. 32, 1063–1075.e1067 (2020).
Gaublomme, J. T. et al. Single-cell genomics unveils critical regulators of Th17 cell pathogenicity. Cell 163, 1400–1412 (2015).
Bao, X. R. et al. Mitochondrial dysfunction remodels one-carbon metabolism in human cells. eLife 5, e10575 (2016).
Nikkanen, J. et al. Mitochondrial DNA replication defects disturb cellular dNTP pools and remodel one-carbon metabolism. Cell Metab. 23, 635–648 (2016).
Birsoy, K. et al. An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell 162, 540–551 (2015).
Sullivan, L. B. et al. Supporting aspartate biosynthesis is an essential function of respiration in proliferating cells. Cell 162, 552–563 (2015).
Wu, L. et al. Niche-selective inhibition of pathogenic Th17 cells by targeting metabolic redundancy. Cell 182, 641–654.e620 (2020).
Berod, L. et al. De novo fatty acid synthesis controls the fate between regulatory T and T helper 17 cells. Nat. Med. 20, 1327–1333 (2014).
Locasale, J. W. et al. Phosphoglycerate dehydrogenase diverts glycolytic flux and contributes to oncogenesis. Nat. Genet. 43, 869–874 (2011).
Blagih, J., Krawczyk, C. M. & Jones, R. G. LKB1 and AMPK: central regulators of lymphocyte metabolism and function. Immunol Rev 249, 59–71 (2012).
MacIver, N. J. et al. The liver kinase B1 is a central regulator of T cell development, activation, and metabolism. J. Immunol. 187, 4187–4198 (2011).
He, N. et al. Metabolic control of regulatory T cell (Treg) survival and function by Lkb1. Proc. Natl Acad. Sci. USA 114, 12542–12547 (2017).
Yang, K. et al. Homeostatic control of metabolic and functional fitness of Treg cells by LKB1 signalling. Nature 548, 602–606 (2017).
Kottakis, F. et al. LKB1 loss links serine metabolism to DNA methylation and tumorigenesis. Nature 539, 390–395 (2016).
Shackelford, D. B. & Shaw, R. J. The LKB1–AMPK pathway: metabolism and growth control in tumour suppression. Nat. Rev. Cancer 9, 563–575 (2009).
Tiainen, M., Ylikorkala, A. & Makela, T. P. Growth suppression by Lkb1 is mediated by a G1 cell cycle arrest. Proc. Natl Acad. Sci. USA 96, 9248–9251 (1999).
Kullmann, L. & Krahn, M. P. Controlling the master-upstream regulation of the tumor suppressor LKB1. Oncogene 37, 3045–3057 (2018).
Fan, J. et al. Human phosphoglycerate dehydrogenase produces the oncometabolite d-2-hydroxyglutarate. ACS Chem. Biol. 10, 510–516 (2015).
Intlekofer, A. M. et al. Hypoxia induces production of l-2-hydroxyglutarate. Cell Metab. 22, 304–311 (2015).
Intlekofer, A. M. et al. l-2-Hydroxyglutarate production arises from noncanonical enzyme function at acidic pH. Nat. Chem. Biol. 13, 494–500 (2017).
Yang, L. et al. Serine catabolism feeds NADH when respiration is impaired. Cell Metab. 31, 809–821.e806 (2020).
Buck, M. D. et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell 166, 63–76 (2016).
Faubert, B. et al. Loss of the tumor suppressor LKB1 promotes metabolic reprogramming of cancer cells via HIF-1α. Proc. Natl Acad. Sci. USA 111, 2554–2559 (2014).
Johnson, M. O. et al. Distinct regulation of Th17 and Th1 cell differentiation by glutaminase-dependent metabolism. Cell 175, 1780–1795.e1719 (2018).
Baksh, S. C. & Finley, L. W. S. Metabolic coordination of cell fate by α-ketoglutarate-dependent dioxygenases. Trends Cell Biol. 31, 24–36 (2021).
Buck, M. D., Sowell, R. T., Kaech, S. M. & Pearce, E. L. Metabolic instruction of immunity. Cell 169, 570–586 (2017).
Reinhardt, R. L., Liang, H. E. & Locksley, R. M. Cytokine-secreting follicular T cells shape the antibody repertoire. Nat. Immunol. 10, 385–393 (2009).
Mohrs, K., Wakil, A. E., Killeen, N., Locksley, R. M. & Mohrs, M. A two-step process for cytokine production revealed by IL-4 dual-reporter mice. Immunity 23, 419–429 (2005).
Wan, Y. Y. & Flavell, R. A. Identifying Foxp3-expressing suppressor T cells with a bicistronic reporter. Proc. Natl Acad. Sci. USA 102, 5126–5131 (2005).
Sinclair, L. V., Neyens, D., Ramsay, G., Taylor, P. M. & Cantrell, D. A. Single cell analysis of kynurenine and system L amino acid transport in T cells. Nat. Commun. 9, 1981 (2018).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC–seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.21–21.29.29 (2015).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Lun, A. T. & Smyth, G. K. De novo detection of differentially bound regions for ChIP-seq data using peaks and windows: controlling error rates correctly. Nucleic Acids Res. 42, e95 (2014).
Lun, A. T. & Smyth, G. K. csaw: a Bioconductor package for differential binding analysis of ChIP-seq data using sliding windows. Nucleic Acids Res. 44, e45 (2016).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Musa, Y. R., Boller, S., Puchalska, M., Grosschedl, R. & Mittler, G. Comprehensive proteomic investigation of Ebf1 heterozygosity in pro-B lymphocytes utlizing data independent acquisition. J. Proteome Res. 17, 76–85 (2018).
Acknowledgements
We thank the members of the Immunometabolism Department at the MPI-IE for helpful discussion; the Electron Microscopy Laboratory at the University of Padova for the invaluable help; and the Imaging, Metabolomics, FACS, Sequencing and Animal facilities at the MPI-IE for their excellent technical support. This work was supported in part by the Max Planck Society, the Leibniz Prize (to E.L.P.), two Bloomberg Distinguished Professorships from Johns Hopkins University (E.L.P. and E.J.P.), the National Institutes of Health R01AI156274 (to E.L.P.) and R35GM144103 (to H.S.), a Marie Sklodowska-Curie actions Individual Fellowship (MSCA-IF) (to F.B.), a Sir Henry Wellcome Fellowship (to D.J.P.) and an Alexander von Humboldt Postdoctoral Fellowship (to M.V.).
Author information
Authors and Affiliations
Contributions
F.B., K.P., D.J.P., M.V., C.S.F., L.J.F., A.Q., J.E.-H., M.A.S., K.M.G., A.M.K., B.K., M.F., M.M., G.C., M.C., K.J.D., H.S.,T.J., J.M.B., H.S., D.O., E.J.P. and E.L.P. designed and/or performed experiments. F.B., K.P., D.J.P., C.S.F., L.J.F., M.V., A.Q., J.E.-H., M.A.S., K.M.G., A.M.K., M.F., M.M., G.C., M.C., K.J.D., T.J., J.M.B., D.O., E.J.P. and E.L.P. analysed data. D.E.S. and N.R. performed all bioinformatics analysis. Y.M. and G.M. performed proteomics analysis. F.B. and E.L.P. designed the study and wrote the manuscript. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
E.L.P. is member of the scientific advisory boards of ImmunoMet Therapeutics. E.L.P. and E.J.P. are founders of and scientific advisors to Rheos Medicines.
Peer review
Peer review information
Nature thanks Navdeep Chandel, Luca Scorrano and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mitochondrial membrane remodeling in CD4+ TN cells under T-helper (Th) and regulatory (Treg) conditions in vitro and in vivo.
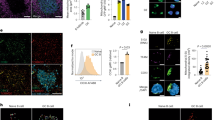
a) Mitochondrial sphericity in live cell spinning disk microscopy images in CD4+ TN cells (PhAM mice) cultured in Th and Treg conditions (TN: n = 1673, Th1: n = 8406, Th2: n = 5485, Th17: n = 7672 and Treg: n = 4824 segmented objects (mitochondria), 2 independent experiments. Violin plot shows median with quartiles. b) EM images (left) and cristae width (right) in TN cells cultured in Th and Treg conditions (TN: n = 362, Th1: n = 479, Th2: n = 369, Th17: n = 319, and Treg: n = 294 cristae, 3 biological replicates). (c) Intracellular expression in TN, Th and Treg cells (n = 4 biological replicates). Representative graph summarize results of two independent experiments. d) Immunoblot of CD4+ TN, Th and Treg cells, day 4, representative experiment out of two. ACTIN: sample processing control (e) Quantification of protein levels from immunoblots as in Fig. 1b (n = 4 biological replicates, 2 independent experiments). f) Immunoblot of proteins that control mitochondrial dynamics in CD4+ TN cells cultured for the indicated times in Th and Treg conditions. TUBULIN: sample processing control. Right, protein levels (n = 4 biological replicates, 2 independent experiments). g) EM images (left) and cristae width (right) in CD4+ TN and effector T cells from reporter mice subjected to immune challenges (naive: n = 410, Th1: n = 262, Th2: n = 675, Th17: n = 461 and Treg: n = 402 cristae, 3 mice per condition). Data b, g are geometric mean, c mean and e, f means ± s.e.m. Two-sided Tukey’s test (b) or Dunnett’s test (a, c, e, f, g). Exact P values are indicated.
Extended Data Fig. 2 Mitochondrial dysfunction and proliferation defects in Opa1CD4Cre T cells.
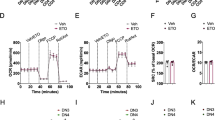
CD4+ TN cells from control and Opa1CD4Cre mice were cultured 4 days in Th and Treg conditions. a) Immunoblot for OPA1 at the end of the culture (2 biological replicates per condition shown). ACTIN: loading control. b) Spinning disk microscopy images from mitochondria (MitoTracker Deep Red, purple) and nuclei (Hoechst, blue). Scale bar: 2 μm. Representative images from 3 biological replicates. c) Baseline OCR (left) and ECAR (right) (n = 4 biological replicates). d) Mitochondrial membrane potential determined by Tetramethylrhodamine methyl ester (TMRM) (n = 3 biological replicates)e) Cell proliferation, day 3 (n = 8 biological replicates, 2 independent experiments). f) Cell viability determined by 7-aminoactinomycin D (7-AAD) and annexin-V staining, day 4 (n = 7 biological replicates, 2 independent experiments). g) Cell-cycle analysis determined by Ki-67 and DAPI staining (n = 3 biological replicates). h) Surface expression of CD25, CD44 and CD69, and intracellular expression of Nur77 in CD4+ TN cells from control and Opa1CD4Cre mice cultured 3 days in Th and Treg conditions (n = 4 biological replicates). i) IL-17A expression and proliferation in control and Opa1CD4Cre Th17 cells, day 3. Graph shows IL-17A expression per each cell division state gated on live CD4+ T cells (n = 4 biological replicates). j) IL-17A expression in CD4+ TN cells cultured in Th17 conditions with increasing concentration of plate bound anti-CD3 or IL-6 for three days and restimulated (n = 4 biological replicates). Representative plots and a graph summarizing the results of at least two independent experiments are shown, except where noted otherwise. Data are means ± s.e.m. Two-sided Šidák’s test (d, e, f, h, i, j) or unpaired two-tailed t-test (c). Exact P values are indicated.
Extended Data Fig. 3 Role of inner and outer MM remodeling in IL-17A production.
Mfn1, Mfn2 or Dnml1 floxed mice were crossed with CD4Cre mice (Mfn1CD4Cre, Mfn2CD4Cre and Drp1CD4Cre). a, b, c, d, e, CD4+ TN cells across genotypes were cultured in Th and Treg conditions. a) Immunoblot of MFN1, MFN2 and DRP1 in a representative experiment out of two. ACTIN: loading control. b) Cytokine and TF expression in Th- and Treg-reestimulated cells across genotypes (Th1 conditions: Control n =15; Mfn1, Mfn2 n =11; Drp1 n =8; Th2, Th17 and Treg conditions; Control n =11; Mfn1, Mfn2 n =7; Drp1 n =4, biological replicates). c) Intracellular levels of T-bet and ROR-γt (Control n = 15; Mfn1CD4Cre n =11; Mfn2CD4Cre n =11; Drp1CD4Cre n =8, biological replicates). d) Proliferation as determined by Cell Trace Violet dilution staining (n = 4 biological replicates, except control; n = 6). e) Relative baseline OCR, ATP-coupled respiration, and maximal respiration in Th17 cells across genotypes (n = 4 biological replicates per group). f, g, h, i, Control and Opa1CD4Cre CD4+ TN cells were transfected with Cas9 and indicated gRNAs. f) Immunoblot for DRP1 and OPA1. ACTIN: loading control. g) Representative spinning disk confocal images from mitochondria (Mitotracker Deepred, green) and nuclei (Hoechst, blue) from 3 biological and transfection replicates. h) OCR upon oligomycin (Oligo), fccp and rotenone + Antimycin A (ROT+AA) addition (left) and relative baseline OCR (right) (n =4 biological and transfection replicates). i) Intracellular IL-17A expression (n =7 biological and transfection replicates). Representative plots and a graph summarizing the results of at least two independent experiments are shown, except where noted otherwise. Data are means ± s.e.m. Two-sided Tukey’s test (b, c, d, e, h, i). Exact P values are indicated.
Extended Data Fig. 4 OPA1-requirement for IL-17A expression across cell culture and in vivo conditions.
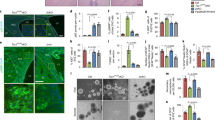
TN cells cultured in Th17 non-pathogenic (np) and pathogenic (p) conditions. a) EM images and cristae width (Control np: n = 528, p: n = 343; Opa1CD4Cre np: n = 321, p: n = 267 cristae, 3 biological replicates). Graph shows geometric mean, black. (b) Cytokine (c) TF expression, d) supernatant cytokines, and e) proliferation, n = 4 biological replicates. f) Cell viability (n = 8, 2 independent experiments). g) Gating strategy to identify CD4+TCRβ+eYFP+ (Th17) cells. h) Ki-67 expression in LP Th17 cells (Control: n = 6; Opa1IL17aCre: n = 3), PP (Control: n = 7; Opa1IL17aCre: n = 4) and MLN (Control: n = 6; Opa1IL17aCre: n = 3), 2 independent experiments. i) Th17 cell numbers (LP: n = 3; PP; Control: n = 3; Opa1IL17aCre: n = 4, MLN: n = 4). Mice were anti-CD3 immunized. j) LP Th17 cell numbers (n = 4), k) ROR-γT expression (n = 4), l) surface expression (CCR6, CXCR3: Control: n = 4; Opa1IL17aCre: n = 3; ICOS, CD27, CD103: n = 4) and m) Ki-67 levels (Control: n = 9; Opa1IL17aCre: n = 8, 2 independent experiments). Mice were MOG immunized. n) Disease incidence, onset and peak disease (days, controlIL17aCre: n = 16; Opa1IL17aCre: n = 14, 2 independent experiments. o) Cell numbers (n = 8). p) Neutrophil percentage (CD45+CD11b+Ly6G+, n = 8). q) CCR6 and CXCR3 levels (n = 8). r) Phosphorylated S6 and 4E-BP1 (CD4+TCRβ+eYFP+ cells). Plots and graphs summarize results of at least two independent experiments, except noted otherwise. Data are means ± s.e.m. Two-sided Tukey’s test (a) or Šidák’s test (b, c, e, i, l, q) or unpaired two-tailed t-test (h, j, k, m, o, p). Exact P values are indicated.
Extended Data Fig. 5 Th17 cells rely on OPA1 independent of metabolic activity.
a, b, c, d, CD4+ TN cells were cultured in Th and Treg conditions. a) Bioenergetic profile (baseline OCR/ECAR, n = 4 independent experiments). b) Glucose and kynurenine uptake, phosphorylated S6 (n = 4) and 4E-BP1 (Th1: n = 3, Th2,Th17,Treg: n = 4 biological replicates). c) de novo protein and nascent RNA synthesis by O-propargyl puromycin (OPP) and ribonucleoside ethynyl uridine (EU) incorporation, respectively (n = 4 biological replicates). Cyclohexamide (CHX) and actinomycin D (ActD): negative controls. d) Protein translation dependence on mitochondria (left, Th1, Th2: n = 6; Th17,Treg: n = 7 biological replicates) or glycolysis (right, Th1, Th2: n = 7: Th17, Treg: n = 8 biological replicates), 2 independent experiments. (e) OCR and ECAR (10 mM: n = 8; 1 mM and 0 mM n = 4 biological replicates) and (f) IFN-γ expression (n = 3 biological replicates) in cells cultured in Th1 conditions 3 days before overnight culture in RPMI with 10% non-dialyzed FBS and decreasing glucose concentrations. g, h, i, CD4+ TN cells cultured in Th17 np/p conditions. g) Bioenergetic profile (n =4 biological replicates). h) de novo protein synthesis (left, n = 7 biological replicates, 2 independent experiments) and nascent RNA synthesis (right, np/p: n = 4, ActD: n = 2 biological replicates). i) Protein translation dependence on mitochondria (left, np: n = 8: p: n = 3 biological replicates) or glycolysis (right, np: n = 8: p: n = 7 biological replicates, 2 independent experiments). Representative plots and graphs summarize results of at least two independent experiments, except where noted otherwise. Data are means ± s.e.m. Two-sided Tukey’s test (b–e), Šidák’s test (f) or unpaired two-tailed t-test (h, i). Exact P values are indicated.
Extended Data Fig. 6 Multi-OMICS analysis of Th17 cells upon OPA1 deletion.
Control and Opa1CD4Cre CD4+ TN cells were cultured in Th17 conditions and the RNA extracted and sequenced. a) Volcano plot shows differentially expressed genes (1518: log10 P value < 3, log2 FC > 0.5). b) DAVID KEGG pathway analysis from differentially up- (779) or downregulated (739) genes (FDR, false discovery rate). c) Control and Opa1CD4Cre CD4+ TN cells were cultured in Th17 conditions and proteins from whole cell lysates extracted and analyzed (LC-MS). Left, volcano plot from differentially expressed proteins (562: log10 P value<1.3, log2 FC > 0.58). Middle and right volcano plots, protein expression in relation to OXPHOS and serine and glycine biosynthesis pathways, respectively. d) Cellular ATP, AMP and AMP/ATP ratio by LC-MS (n= 4 per genotype, day 4). e) Schematic of [U-13C]-Glucose and [U-13C]-Glutamine carbon allocation into central carbon metabolites. f) Fractional labelling of palmitate with [U-13C]-Glucose (left) or [U-13C]-Glutamine (right) in control and Opa1CD4Cre Th17 cells cultured 20 h on day 3 with fully labelled subsrates (n = 4 per group). g) Control and Opa1CD4Cre CD4+ TN cells were cultured in Th17 conditions 4 days and lipids extracted and analyzed (n = 4 biological replicates). Graph, FC of distinct lipid classes in Opa1CD4Cre vs. control (HEXCER: hexosylceramide, TG: triacylglycerades, LPI: lysophosphatidylinositol, DG: diacylglycerol, PA: phosphatidic acid, LPE: lysophosphatidylethanolamine, PG: phosphatidylglycerol, Cholesterol, PC: phosphatidylcholines, CER: ceramides, SM: sphingomyelin, PS: phosphatidylserine, LPC: lysophosphatidylcholines, PE: phosphatidylethanolamine, PI: phosphatidylinositol, Sphingosine, LPS: lipopolysaccharide, CE: cholesterol esters, CL: cardiolipins). Transcriptomics, proteomics and lipidomics were performed once (a,b: n = 3; g: n = 4 biological replicates). Graphs summarize results of at least two independent experiments in d, f. Data are means ± s.e.m. Unpaired two-tailed t-test (a, c, d, f, g). Exact P values are indicated.
Extended Data Fig. 7 OPA1 controls chromatin accessibility in Th17 cells.
a) CD4+ TN cells from control and Opa1CD4Cre mice were cultured in Th17 conditions for 1 day followed by 48 h incubation with inhibitors for histone acetylation and methylation, and DNA methylation. IL-17A expression was analyzed by flow cytometry and results show log2 FC of IL-17A MFI in inhibitor-treated vs vehicle-treated cells (n = 7 biological replicates for all conditions except GSK: n = 4 biological replicates). Graph combines data from two independent experiments. Data are mean ± s.e.m. Two sided Šidák’s test. Exact P values are indicated. b, c, d, e, CD4+ TN cells from control and Opa1CD4Cre mice were cultured in Th17 conditions 4 days and the DNA extracted and processed for ATAC-seq (n = 3 biological replicates). b) Number of differentially accessible chromatin regions (FC > 2, p < 0.01). c) Genomic localization of the differentially regulated accessible chromatin regions. d) HOMER motif TF analysis on regulated chromatin regions. e) GO term pathway enrichment analysis from the differentially regulated accessible chromatin regions between Opa1CD4Cre and control Th17 cells (FC > 2, p < 0.01).
Extended Data Fig. 8 LKB1 deletion in OPA1 reestablishes IL-17A expression.
LKB1 phosphorylation kinetics in CD4+ TN cells cultured in Th and Treg (a) and np/p Th17 (b) conditions (Control: n = 4; Opa1CD4Cre: n = 3 biological replicates). c, d, e, f, g, h Control and Opa1CD4Cre CD4+ TN cells transfected with Cas9 and indicated gRNAs. (c) Immunoblot for LKB1 and OPA1. ACTIN: loading control, (d) IL-17A mRNA expression (RT-PCR, n = 7), (e) CD25, CD44,CD69 and Nur77 expression (n = 3), (f) ROR-γT expression (n = 4), (g) cell proliferation (n = 4), (h) and cell-cycle analysis (n = 4, biological and transfection replicates). i) Left, clinical disease score upon MOG immunization and at day 16 (Right), (control: n = 8; Opa1IL17aCre: n = 7; and Opa1Stk11hetIL17aCre: n = 6 mice). j) CD4+TCRβ+eYFP+ cell frequencies, day 20 post-immunization (n = 6 mice). k) Immunoblot for LKB1 and OPA1 in CD4+TCRβ+eYFP+ cells from control, Opa1IL17aCre, Stk11IL17aCre and Opa1Stk11 IL17aCre mice. ACTIN: loading control. l) Left, clinical disease score upon MOG immunization and at day 16 (Right) (control: n = 9; Opa1IL17aCre: n = 7; Stk11IL17aCre: n = 6 and Opa1Stk11 IL17aCre: n = 6 mice). m) CD4+TCRβ+eYFP+ SC cell numbers (n = 6 mice per genotype except Stk11IL17aCre: n = 5 mice) and n) IL-17A and IFN-γ expression, day 20 post-immunization (control: n = 6 mice; Opa1IL17Cre: n = 5 mice; Stk11IL17Cre: n = 5 mice; Opa1Stk11 IL17Cre: n = 6 mice). Representative plots and graphs summarize results of at least two independent experiments, except noted otherwise. Data are means ± s.e.m. Two-sided Šidák’s test (a, b, h, n) or Tukey’s test (d, e, f, g, i, j, l, m). Exact P values are indicated.
Extended Data Fig. 9 LKB1 antagonizes OPA1 deficiency independent of mTOR and AMPK.
a) Immunoblot for phosphorylated and total AMPK and ACC in control and Opa1CD4Cre Th17 cells. ACTIN: sample processing control. Graph, relative signal intensity (n = 4, 2 independent experiments). b) IL-17A expression in control and Opa1CD4Cre CD4+ TN cells cultured in Th17 conditions and Compound C from day 0 (d0) or from day 3 to 4 (d3) and restimulated (n = 3). c) Left, immunoblot for AMPK and OPA1 in T cells transfected with Cas9 and indicated gRNAs and cultured 5 days in Th17 conditions. ACTIN: sample processing control. Right, IL-17A expression in restimulated cells (n = 3 biological and transfection replicates). d) Expression of phosphorylated S6 (n = 8) and 4E-BP1 (n = 7 biological and transfection replicates), 2 independent experiments. e) IL-17A expression in T cells transfected with Cas9 and indicated gRNAs and cultured in Th17 conditions 2 days followed by 48 h rapamycin treatment (n = 3 biological and transfection replicates). f) Phosphorylated H2Ax expression (n = 6, 2 independent experiments). g) Cyclic AMP (cAMP) levels (LC-MS, n = 8, 2 independent experiments). h) Basal calcium levels (n = 8, 2 independent experiments). i) Mitochondrial ROS (mitoSOX, n = 3). j) Control and Opa1CD4Cre CD4+ TN cells cultured in Th17 conditions 1 day followed by 2 day inhibitor-treatment. Graph, IL-17A MFI log2 FC inhibitor-treated vs. vehicle-treated cells (n = 7 biological replicates, except Tempol: n = 4 and NAC: n = 3, 2 independent experiments). Representative plots and graphs summarize results of at least two independent experiments, except noted otherwise. Data (a–i) are means ± s.e.m. Unpaired two-tailed t-test (a, f, g, h, i), two-sided Tukey’s test (b, c, d) or Šidák’s test (e, j). Exact P values are indicated.
Extended Data Fig. 10 LKB1 couples TCA cycle and serine metabolism to Th17 effector function.
a) Principal component analysis (PCA) of the transcriptome of control and Opa1CD4Cre CD4+ TN cells transfected with Cas9 and indicated gRNAs, cultured 4 days in Th17 conditions (n = 3 biological and transfection replicates). b) Volcano plots of the differentially expressed genes (DEG, FC > 1, p-value < 0.001) for the comparisons: control gStk11 vs. control gCtl (left) and Opa1CD4Cre gStk11 vs. Opa1CD4Cre gCtl (right). Blue dots annotate the highest 40 DEG. c, d, KEGG pathway and Gene Ontology (GO) annotation analysis from DEG (FC>1, p-value<0.001) between (c) control gStk11 vs. control gCtl or (d) Opa1CD4Cre gStk11 vs. Opa1CD4Cre gCtl. e) Intracellular serine in control and Opa1CD4Cre CD4+ TN cells transfected with Cas9 and indicated gRNAs (n = 6 transfection replicates, 2 independent experiments) by LC-MS. f) Left, schematic shows 2-HG production from primary and promiscuous enzymatic reactions and the inhibitors used for targeting its production. Right, CD4+ TN cells from control and Opa1CD4Cre mice were cultured under Th17 conditions for 1 day followed by 48 h inhibitor treatment. IL-17A expression was analyzed by flow cytometry in restimulated cells and expressed as log2 FC inhibitor-treated vs. vehicle-treated cells (CBR, AGI, NHI-2, GSK864: n = 7 per group; BAY, LW6: n = 3 per group). Graph summarizes data from 2 experiments. g) Kinetics of fractional labelling of glutamate (m + 5), α-ketoglutarate (m + 4), malate (m + 4) and fumarate (m+4) in control and Opa1CD4Cre CD4+ T cells transfected with Cas9 and indicated gRNAs cultured for 4 days in Th17 conditions and incubated with [U-13C]-Glutamine for the indicated times (n = 3 biological and transfection replicates). Data are means ± s.e.m. Two-sided Šidák’s test (f) or Tukey’s test (e, g). Exact P values are indicated.
Supplementary information
Source data
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Baixauli, F., Piletic, K., Puleston, D.J. et al. An LKB1–mitochondria axis controls TH17 effector function. Nature 610, 555–561 (2022). https://doi.org/10.1038/s41586-022-05264-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05264-1
This article is cited by
-
TBC1D23 mediates Golgi-specific LKB1 signaling
Nature Communications (2024)
-
Small molecule metabolites: discovery of biomarkers and therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
Drugs targeting TGF-β/Notch interaction attenuate hypertrophic scar formation by optic atrophy 1-mediated mitochondrial fusion
Molecular and Cellular Biochemistry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.