Abstract
Stomata exert considerable effects on global carbon and water cycles by mediating gas exchange and water vapour1,2. Stomatal closure prevents water loss in response to dehydration and limits pathogen entry3,4. However, prolonged stomatal closure reduces photosynthesis and transpiration and creates aqueous apoplasts that promote colonization by pathogens. How plants dynamically regulate stomatal reopening in a changing climate is unclear. Here we show that the secreted peptides SMALL PHYTOCYTOKINES REGULATING DEFENSE AND WATER LOSS (SCREWs) and the cognate receptor kinase PLANT SCREW UNRESPONSIVE RECEPTOR (NUT) counter-regulate phytohormone abscisic acid (ABA)- and microbe-associated molecular pattern (MAMP)-induced stomatal closure. SCREWs sensed by NUT function as immunomodulatory phytocytokines and recruit SOMATIC EMBRYOGENESIS RECEPTOR-LIKE KINASE (SERK) co-receptors to relay immune signalling. SCREWs trigger the NUT-dependent phosphorylation of ABA INSENSITIVE 1 (ABI1) and ABI2, which leads to an increase in the activity of ABI phosphatases towards OPEN STOMATA 1 (OST1)—a key kinase that mediates ABA- and MAMP-induced stomatal closure5,6—and a reduction in the activity of S-type anion channels. After induction by dehydration and pathogen infection, SCREW–NUT signalling promotes apoplastic water loss and disrupts microorganism-rich aqueous habitats to limit pathogen colonization. The SCREW–NUT system is widely distributed across land plants, which suggests that it has an important role in preventing uncontrolled stomatal closure caused by abiotic and biotic stresses to optimize plant fitness.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information files (uncropped blots and gel images, primers, peptides, and exact P values). The sequences of proteins were obtained from TAIR (https://www.arabidopsis.org/), UniProt (https://www.uniprot.org/), NCBI (https://www.ncbi.nlm.nih.gov/) and Phytozome (https://phytozome-next.jgi.doe.gov/). The protein structures were obtained from AlphaFold Protein Structure Database (https://alphafold.ebi.ac.uk/). Sequence data from this article can be found in the Arabidopsis Genome Initiative or GenBank–EMBL databases under the following accession numbers: SCREW1 (AT1G06135), SCREW2 (AT2G31345), SCREW3 (AT1G06137), SCREW4 (AT2G31335), NUT (AT5G25930), WRKY30 (AT5G24110), WRKY33 (AT2G38470), WRKY53 (AT4G23810), PR1 (AT2G14610), FRK1 (AT2G19190), MPK3 (AT3G45640), MPK4 (AT4G01370), MPK6 (AT2G43790), UBQ10 (AT4G05320), ACTIN2 (AT3G18780), BAK1 (AT4G33430), SERK1 (AT1G71830), SERK2 (AT1G34210), SERK4 (AT2G13790), FLS2 (AT5G46330), BIK1 (AT2G39660), MIK2 (AT4G08850), MIK2L (AT1G35710), RLK7 (AT1G09970), IKU2 (AT3G19700), HAE (AT4G28490), HSL2 (AT5G65710), LET1 (AT2G23200), LET2 (AT5G38990), NILR1 (AT1G74360), MIN7 (AT3G43300), ABI1 (AT4G26080), ABI2 (AT5G57050) and OST1 (AT4G33950).
References
Hetherington, A. M. & Woodward, F. I. The role of stomata in sensing and driving environmental change. Nature 424, 901–908 (2003).
Sussmilch, F. C., Schultz, J., Hedrich, R. & Roelfsema, M. R. G. Acquiring control: the evolution of stomatal signalling pathways. Trends Plant Sci. 24, 342–351 (2019).
Hsu, P. K., Dubeaux, G., Takahashi, Y. & Schroeder, J. I. Signaling mechanisms in abscisic acid-mediated stomatal closure. Plant J. 2, 307–321 (2021).
Melotto, M., Underwood, W. & He, S. Y. Role of stomata in plant innate immunity and foliar bacterial diseases. Annu. Rev. Phytopathol. 46, 101–122 (2008).
Guzel Deger, A. et al. Guard cell SLAC1-type anion channels mediate flagellin-induced stomatal closure. New Phytol. 208, 162–173 (2015).
Mustilli, A. C., Merlot, S., Vavasseur, A., Fenzi, F. & Giraudat, J. Arabidopsis OST1 protein kinase mediates the regulation of stomatal aperture by abscisic acid and acts upstream of reactive oxygen species production. Plant Cell 14, 3089–3099 (2002).
Takahashi, F., Hanada, K., Kondo, T. & Shinozaki, K. Hormone-like peptides and small coding genes in plant stress signaling and development. Curr. Opin. Plant Biol. 51, 88–95 (2019).
Olsson, V. et al. Look closely, the beautiful may be small: precursor-derived peptides in plants. Annu. Rev. Plant Biol. 70, 153–186 (2019).
Tang, D., Wang, G. & Zhou, J. M. Receptor kinases in plant–pathogen interactions: more than pattern recognition. Plant Cell 29, 618–637 (2017).
Dievart, A., Gottin, C., Perin, C., Ranwez, V. & Chantret, N. Origin and diversity of plant receptor-like kinases. Annu. Rev. Plant Biol. 71, 131–156 (2020).
Couto, D. & Zipfel, C. Regulation of pattern recognition receptor signalling in plants. Nat. Rev. Immunol. 16, 537–552 (2016).
Zhou, J. M. & Zhang, Y. Plant immunity: danger perception and signaling. Cell 181, 978–989 (2020).
Hou, S., Liu, D. & He, P. Phytocytokines function as immunological modulators of plant immunity. Stress Biol. 1, 8 (2021).
Tanaka, K. & Heil, M. Damage-associated molecular patterns (DAMPs) in plant innate immunity: applying the danger model and evolutionary perspectives. Annu. Rev. Phytopathol. 59, 53–75 (2021).
Hou, S. et al. The Arabidopsis MIK2 receptor elicits immunity by sensing a conserved signature from phytocytokines and microbes. Nat. Commun. 12, 5494 (2021).
Rhodes, J. et al. Perception of a divergent family of phytocytokines by the Arabidopsis receptor kinase MIK2. Nat. Commun. 12, 705 (2021).
Li, F. et al. Modulation of RNA polymerase II phosphorylation downstream of pathogen perception orchestrates plant immunity. Cell Host Microbe 16, 748–758 (2014).
Liang, X. & Zhou, J. M. Receptor-like cytoplasmic kinases: central players in plant receptor kinase-mediated signaling. Annu. Rev. Plant Biol. 69, 267–299 (2018).
Xu, G., Moeder, W., Yoshioka, K. & Shan, L. A tale of many families: calcium channels in plant immunity. Plant Cell 2022, koac033 (2022).
Ross, A. et al. The Arabidopsis PEPR pathway couples local and systemic plant immunity. EMBO J. 33, 62–75 (2014).
Santiago, J. et al. Mechanistic insight into a peptide hormone signaling complex mediating floral organ abscission. eLife 5, e15075 (2016).
Yamaguchi, Y., Huffaker, A., Bryan, A. C., Tax, F. E. & Ryan, C. A. PEPR2 is a second receptor for the Pep1 and Pep2 peptides and contributes to defense responses in Arabidopsis. Plant Cell 22, 508–522 (2010).
Hou, S. et al. The secreted peptide PIP1 amplifies immunity through receptor-like kinase 7. PLoS Pathog. 10, e1004331 (2014).
Furumizu, C. et al. The sequenced genomes of nonflowering land plants reveal the innovative evolutionary history of peptide signaling. Plant Cell 33, 2915–2934 (2021).
Chen, Y. et al. An aphid RNA transcript migrates systemically within plants and is a virulence factor. Proc. Natl Acad. Sci. USA 117, 12763–12771 (2020).
Ma, X., Xu, G., He, P. & Shan, L. SERKing coreceptors for receptors. Trends Plant Sci. 21, 1017–1033 (2016).
Lin, W., Ma, X., Shan, L. & He, P. Big roles of small kinases: the complex functions of receptor-like cytoplasmic kinases in plant immunity and development. J. Integr. Plant Biol. 55, 1188–1197 (2013).
Melotto, M., Underwood, W., Koczan, J., Nomura, K. & He, S. Y. Plant stomata function in innate immunity against bacterial invasion. Cell 126, 969–980 (2006).
Aung, K., Jiang, Y. & He, S. Y. The role of water in plant–microbe interactions. Plant J. 93, 771–780 (2018).
Beattie, G. A. Water relations in the interaction of foliar bacterial pathogens with plants. Annu. Rev. Phytopathol. 49, 533–555 (2011).
Freeman, B. C. & Beattie, G. A. Bacterial growth restriction during host resistance to Pseudomonas syringae is associated with leaf water loss and localized cessation of vascular activity in Arabidopsis thaliana. Mol. Plant Microbe Interact. 22, 857–867 (2009).
Xin, X. F. et al. Bacteria establish an aqueous living space in plants crucial for virulence. Nature 539, 524–529 (2016).
Axtell, C. A. & Beattie, G. A. Construction and characterization of a proU-gfp transcriptional fusion that measures water availability in a microbial habitat. Appl. Environ. Microbiol. 68, 4604–4612 (2002).
Tanaka, T., Tanaka, H., Machida, C., Watanabe, M. & Machida, Y. A new method for rapid visualization of defects in leaf cuticle reveals five intrinsic patterns of surface defects in Arabidopsis. Plant J. 37, 139–146 (2004).
Yoshida, T., Mogami, J. & Yamaguchi-Shinozaki, K. ABA-dependent and ABA-independent signaling in response to osmotic stress in plants. Curr. Opin. Plant Biol. 21, 133–139 (2014).
Zhu, J. K. Abiotic stress signaling and responses in plants. Cell 167, 313–324 (2016).
Munemasa, S. et al. Mechanisms of abscisic acid-mediated control of stomatal aperture. Curr. Opin. Plant Biol. 28, 154–162 (2015).
Vahisalu, T. et al. SLAC1 is required for plant guard cell S-type anion channel function in stomatal signalling. Nature 452, 487–U415 (2008).
Maierhofer, T. et al. Site- and kinase-specific phosphorylation-mediated activation of SLAC1, a guard cell anion channel stimulated by abscisic acid. Sci. Signal. 7, ra86 (2014).
Yoshida, R. et al. ABA-activated SnRK2 protein kinase is required for dehydration stress signaling in Arabidopsis. Plant Cell Physiol. 43, 1473–1483 (2002).
Ye, W. et al. Stomatal immunity against fungal invasion comprises not only chitin-induced stomatal closure but also chitosan-induced guard cell death. Proc. Natl Acad. Sci. USA 117, 20932–20942 (2020).
Lin, P. A. et al. Stomata-mediated interactions between plants, herbivores, and the environment. Trends Plant Sci. (2021).
Liu, X. S. et al. The LRR-RLK protein HSL3 regulates stomatal closure and the drought stress response by modulating hydrogen peroxide homeostasis. Front. Plant Sci. 11, 548034 (2020).
Rhodes, J. et al. Perception of a conserved family of plant signalling peptides by the receptor kinase HSL3. Preprint at bioRxiv https://doi.org/10.1101/2021.10.25.465685 (2022).
Wu, F. et al. Hydrogen peroxide sensor HPCA1 is an LRR receptor kinase in Arabidopsis. Nature 578, 577–581 (2020).
Mendy, B. et al. Arabidopsis leucine-rich repeat receptor–like kinase NILR1 is required for induction of innate immunity to parasitic nematodes. PLoS Pathog. 13, e1006284 (2017).
Huang, Y. et al. A trimeric CrRLK1L–LLG1 complex genetically modulates SUMM2-mediated autoimmunity. Nat. Commun. 11, 4859 (2020).
Liu, J. et al. The malectin-like receptor-like kinase LETUM1 modulates NLR protein SUMM2 activation via MEKK2 scaffolding. Nat. Plants 6, 1106–1115 (2020).
Meng, X. et al. Ligand-induced receptor-like kinase complex regulates floral organ abscission in Arabidopsis. Cell Rep. 14, 1330–1338 (2016).
Li, B. et al. Phosphorylation of trihelix transcriptional repressor ASR3 by MAP KINASE4 negatively regulates Arabidopsis immunity. Plant Cell 27, 839–856 (2015).
de Oliveira, M. V. V. et al. Specific control of Arabidopsis BAK1/SERK4-regulated cell death by protein glycosylation. Nat. Plants 2, 15218 (2016).
Li, L. et al. TMK4 receptor kinase negatively modulates ABA signaling by phosphorylating ABI2 and enhancing its activity. J. Integr. Plant Biol. 63, 1161–1178 (2021).
Wu, Y. et al. Genome-wide expression pattern analyses of the Arabidopsis leucine-rich repeat receptor-Like kinases. Mol. Plant 9, 289–300 (2016).
Li, F. et al. Modulation of RNA polymerase II phosphorylation downstream of pathogen perception orchestrates plant immunity. Cell Host Microbe 16, 748–758 (2014).
Yu, X. et al. The receptor kinases BAK1/SERK4 regulate Ca2+ channel-mediated cellular homeostasis for cell death containment. Curr. Biol. 29, 3778–3790 (2019).
He, P., Shan, L. & Sheen, J. in Plant–Pathogen Interactions (ed. Ronald, P. C.) 1–9 (Humana Press, 2007).
Maintz, J. et al. Comparative analysis of MAMP-induced calcium influx in Arabidopsis seedlings and protoplasts. Plant Cell Physiol. 55, 1813–1825 (2014).
Zhou, J. et al. Proteolytic processing of SERK3/BAK1 regulates plant immunity, development, and cell death. Plant Physiol. 180, 543–558 (2019).
Lei, J. et al. BOTRYTIS-INDUCED KINASE1 modulates Arabidopsis resistance to green peach aphids via PHYTOALEXIN DEFICIENT4. Plant Physiol. 165, 1657–1670 (2014).
Rodrigues, O. et al. Aquaporins facilitate hydrogen peroxide entry into guard cells to mediate ABA- and pathogen-triggered stomatal closure. Proc. Natl Acad. Sci. USA 114, 9200–9205 (2017).
Babilonia, K. et al. A nonproteinaceous Fusarium cell wall extract triggers receptor-like protein-dependent immune responses in Arabidopsis and cotton. New Phytol. 230, 275–289 (2021).
Eisele, J. F., Fassler, F., Burgel, P. F. & Chaban, C. A rapid and simple method for microscopy-based stomata analyses. PLoS One 11, e0164576 (2016).
Brandt, B. et al. Calcium specificity signaling mechanisms in abscisic acid signal transduction in Arabidopsis guard cells. eLife 4, e03599 (2015).
Munemasa, S. et al. Ethylene inhibits methyl jasmonate-induced stomatal closure by modulating guard cell slow-type anion channel activity via the OPEN STOMATA 1/SnRK2.6 kinase-independent pathway in Arabidopsis. Plant Cell Physiol. 60, 2263–2271 (2019).
Sun, Y. et al. Structural basis for flg22-induced activation of the Arabidopsis FLS2–BAK1 immune complex. Science 342, 624–628 (2013).
Acknowledgements
We thank the Arabidopsis Biological Resource Center (ABRC) and the Nottingham Arabidopsis Stock Centre (NASC) for providing the Arabidopsis T-DNA insertion lines; G. A Beattie for sharing water potential reporters, bacterial strains and assay protocols; R. Panstruga for the p35S::mCherry-AEQ construct; F. Yu for providing abi1-2/abi2-2 mutants; J. Chai and Z. Han for providing BAK1ECD proteins; and J. Li for providing pNUT::GUS seeds. We also thank J. Schroeder and P.-K. Hsu for help with the stomatal conductance experiment and for critical reading of the manuscript, and members of the laboratories of L.S. and P.H. for discussions and comments on the experiments. The work was supported by the National Science Foundation (NSF) (IOS-1951094) and the National Institutes of Health (NIH) (R01GM092893) to P.H.; the NIH (R35GM144275), the NSF (IOS-2049642) and the Robert A. Welch Foundation (A-2122) to L.S.; the National Natural Science Foundation of China (NSFC) (31500971), the Youth Innovation Technology Project of Higher School in Shandong Province (2020KJF013) and the Natural Science Foundation of Shandong Province (ZR2020MC022) to S.H.; the Science and Technology Development Program of Shandong Province (2012GSF11712) to H.W.; and the Natural Science Foundation of Shandong Province (ZR201807100168) to W.Z.
Author information
Authors and Affiliations
Contributions
S.H., Z.L., L.S. and P.H. conceived the project, designed experiments and analysed data. Z.L. and S.H. performed most of the experiments. O.R. performed stomatal movement assays. P.W. constructed the Pst DC3000 reporter strains and contributed to water potential assays. D.L. contributed to observations of the subcellular localization of SCREW and NUT, and to the GUS staining assays. S.M. performed the patch-clamp experiment. J. Lei in the laboratory of K.Z.-S. performed aphid bioassays. K.N. in the laboratory of S.Y.H. contributed to min7 disease assays. X.W. in the laboratory of W.Z. built p35S::SCREW1/2 transgenic lines. J. Liu contributed to the NUT or FLS2 and SERK interaction assays. F.A.O.-M. contributed to SCREW and NUT binding and endocytosis assays. C.Y. contributed to genotyping and Arabidopsis transformation. S.Y.H., K.Z.-S., W.Z. and H.W. analysed data and provided critical feedback; L.S., P.H., S.H. and Z.L. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Jian-Min Zhou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification of SCREWs in Arabidopsis.
a, Upregulation of Arabidopsis peptide genes by flg22 in an FLS2-dependent manner. The gene expression data were extracted from an RNA-Seq analysis17 and subjected to data adjustment by log2 transformation using TBtools for the heat map. b, Phylogenetic analysis of Arabidopsis SCREW homologs and schematic diagrams of different domains. The phylogenetic tree was constructed with MEGAX using neighbour-joining methods. The bootstrap values from 1,000 replications are indicated on the branches. c, SCREW orthologs are present in dicots and monocots. Protein sequences were blast-searched against the NCBI database, and wheat, rice, and maize genomes using Arabidopsis SCREW1 as a query. The phylogenetic tree was constructed as indicated in (b) and displayed with iTOL v5 online software (https://itol.embl.de/). The bootstrap values from 1,000 replications are shown on the branches. d, SCREWs are upregulated after elf18 treatments. Ten-day-old plate-grown seedlings were treated with 200 nM elf18. The expression of SCREWs normalized to UBQ10 was analysed by RT–qPCR. Means (n = 4, biologically independent samples) of fold induction shown as log2 values were used to construct heat map using TBtools. e, SCREWs are upregulated after Pep1 treatments. Ten-day-old plate-grown seedlings were treated with 200 nM Pep1. The expression of SCREWs was detected and shown as in (d) (n = 4, biologically independent samples). f, SCREWs are upregulated after Pst DC3000 infections. Ten-day-old plate-grown seedlings were treated with Pst DC3000 at OD600 nm = 0.01, and gene expression was analysed as in (d) (n = 3, biologically independent samples).
Extended Data Fig. 2 SCREWs activate PTI responses.
a, Recombinant His-SCREW1 induces MAPK activation. Ten-day-old plate-grown seedlings were treated with protein elution buffer (Ctrl) or 1 μM His-SCREW1. MAPK activation was analysed by immunoblotting with anti-pERK1/2 antibodies (top), and protein loading is shown by Ponceau S staining (Ponc.) for RBC (bottom). b, SCREW1 upregulates the expression of WRKY30, WRKY33, and WRKY53. Ten-day-old plate-grown seedlings were treated with 1 µM GST or GST-SCREW1 for 1 h, and gene expression was analysed by RT–qPCR. Means (n = 3, biologically independent samples) of fold induction shown as log2 values were used to construct heat map using TBtools. c, SCREWs induce FRK1 promoter activities. Protoplasts were co-transfected with pFRK1::LUC and pUBQ10::GUS followed by treatment with ddH2O (Ctrl), 100 nM flg22, 1 μM GST, GST-SCREW1, GST-SCREW2, or GST-SCREW3 for 4 h. The FRK1 promoter activity was presented as the ratio of luciferase to GUS values. Data were analysed by one-way ANOVA followed by Tukey’s test, and are shown as mean ± s.d. (n = 12, biologically independent samples). P values are provided in the graph and Supplementary Table 3. d, Schematic diagrams of full-length (FL) and truncated SCREW1 variants. Domains of the signal peptide, variable region, and C-terminal conserved region are shown with aa positions labelled. e, SCREW1-induced MAPK activation requires its conserved C-terminus. Ten-day-old seedlings were treated with 1 µM GST or different GST-SCREW1 truncation proteins for 15 min, and the MAPK activation was detected as in (a). f, Synthesized peptides corresponding to the conserved C-terminal domain in SCREW1 induce MAPK activation. Ten-day-old plate-grown seedlings were incubated with 100 nM flg22, 100 nM Pep1, or 100 nM different SCREW1 variants, and the MAPK activation was detected as in (a). g, SCREW1 induces MAPK activation at a subnanomolar scale. Ten-day-old plate-grown seedlings were treated with synthesized peptides corresponding to SCREW139–69 with indicated concentrations for 15 min, and MAPK activation was detected as in (a). h, SCREW1 induces a comparable MAPK activation with flg22. Ten-day-old plate-grown WT seedlings were treated with 500 nM SCREW1 or 100 nM flg22 for the indicated time. i, SCREW1 induces a weak ROS burst. Leaf discs from four-week-old soil-grown WT plants were treated with or without 100 nM flg22, 100 nM or 1 μM SCREW1, and ROS production was measured as relative light units (RLU) by a luminometer over 60 min. Data are shown as mean ± s.e.m. (n = 12, biologically independent samples). j, SCREW1 induces a moderate cytoplasmic Ca2+ increase relative to flg22. Protoplasts were transfected with p35S::mCherry-AEQ and incubated for 6 h, followed by treatment with 1 μM SCREW1 or 200 nM flg22. Cytoplasmic Ca2+ concentration was detected over 15 min. Data are shown as mean ± s.d. (n = 3, biologically independent samples). k, SCREW1 does not induce BIK1 phosphorylation. Protoplasts expressing BIK1-HA were treated with 100 nM flg22, 100 nM, or 1 μM SCREW1 for 10 min. BIK1-HA proteins were detected by immunoblotting using anti-HA antibodies (top) with protein loading shown by Coomassie brilliant blue (CBB) staining for RBC (bottom). Experiments were repeated at least three times with similar results.
Extended Data Fig. 3 Two conserved cysteine residues are required for SCREW activities, and SCREW1 is not mobile.
a, Predicted structures of SCREW1 and SCREW2 C-terminal 23 aa. Structures were predicted using AlphaFold Protein Structure Database (https://www.alphafold.ebi.ac.uk/). Disulfide bonds are shown by yellow sticks. b, Two cysteine residues are required for SCREW2 activation of MAPKs. Ten-day-old plate-grown WT seedlings were treated with or without 100 nM SCREW2, SCREW2(CC), SCREW2(CC/SS) and SCREW2(ΔC8). MAPK activation was analysed by immunoblots with anti-pERK1/2 antibodies (top), and the protein loading is shown by CBB staining for RBC (bottom). c, Two cysteine residues are required for SCREW2-induced seedling growth inhibition. Three-day-old plate-grown WT seedlings were transferred into liquid ½MS medium without (Ctrl) or with 1 μM peptides. Images were taken (left), and fresh weights of seedlings (right) were measured seven days later. Data are shown as box plots with the interquartile range as the upper and lower confines, minima and maxima as whiskers, and the median as a solid line (n = 12, biologically independent samples). d, The biotin-SCREW1 peptide is not mobile. The third pair of true leaves of four-week-old plants were infiltrated with biotin-SCREW1, and both third (local) and fourth (systemic) pairs of true leaves were collected for the detection of biotin-SCREW1 by immunoblotting with HRP-labelled Streptavidin (top). The protein loading control is shown by CBB staining for RBC (bottom). e, Local application of SCREW1 does not induce PR1 expression in distal leaves. The third pair of true leaves from four-week-old plants were infiltrated with H2O, 500 nM Pep1, or 500 nM SCREW1, and both third (local) and fourth (systemic) pairs of true leaves were collected 24 h later for RT–qPCR using ACTIN2 as internal controls. Data of induction fold compared to H2O treatment are shown as mean ± s.d. (n = 3, biologically independent samples). f, Local application of SCREW1 does not induce PR1 accumulation in distal leaves. The experiment was performed as in e, and PR1 proteins were detected by immunoblotting with anti-PR1 antibodies (top). The protein loading control is shown by CBB staining for RBC (bottom). g, Local application of SCREW1 does not induce disease resistance in distal leaves. The third pair of leaves were pre-infiltrated with 500 nM SCREW1 followed by Pst DC3000 inoculation 24 h later on both third and fourth pairs of leaves. Bacterial growth was detected at three days post-inoculation (dpi). Data are shown as the means ± s.d. (n = 8, biologically independent samples). h, The biotin-SCREW1 and SCREW1-HA peptides have similar activities with SCREW1 for MAPK activation. Ten-day-old plate-grown WT seedlings were treated with or without 100 nM SCREW1, biotin-SCREW1, and SCREW1-HA. MAPK activation was analysed by immunoblots using anti-pERK1/2 antibodies (top) with the protein loading shown by CBB staining for RBC (bottom). i, The biotin-SCREW1 and SCREW1-HA peptides have similar activities with SCREW1 for seedling growth inhibition. The experiment was performed as in (c). Data are shown as box plots with the interquartile range as the upper and lower confines, minima and maxima as whiskers, and the median as a solid line (n = 12, biologically independent samples). Experiments were repeated three times with similar results. Data were analysed by one-way (c, i), or two-way (e, g) ANOVA followed by Tukey’s test. Exact P values are provided in the graphs and Supplementary Table 3.
Extended Data Fig. 4 SCREW1 and SCREW2 are involved in plant immunity.
a, Inducible overexpression of SCREW1 leads to leaf chlorosis. Four-week-old transgenic plants carrying pEst::SCREW1-HA in WT (L12 & L19) were sprayed with 0.05% DMSO (Ctrl) or 50 μM β-oestradiol (Est), and then imaged five days later (Scale bar, 1 cm) with trypan blue staining (Scale bar, 0.5 cm). SCREW1-HA proteins were detected by immunoblots with anti-HA antibodies (bottom). b, SCREW1 overexpression elevates PR1 expression. Four-week-old soil-grown pEst::SCREW1-HA transgenic plants were sprayed with 50 μM β-oestradiol. The PR1 expression normalized to ACTIN2 was analysed by RT–qPCR. Data are shown as mean ± s.d. (n = 3, biologically independent samples). c, Overexpression of SCREW1 or SCREW2 leads to plant growth retardation and leaf curling. Transgenic lines carrying p35S::SCREW1 or p35S::SCREW2 were grown on soil for five weeks before photography. Scale bar, 1 cm. The expression of SCREWs normalized to ACTIN2 in ten-day-old plate-grown seedlings was analysed with RT–qPCR. Data are shown as mean ± s.d. (n = 3, biologically independent samples). d, Overexpression of SCREW1 or SCREW2 upregulates PR1 expression. Relative PR1 expression levels normalized to UBQ10 in four-week-old soil-grown plants were analysed with RT–qPCR. Data are shown as mean ± s.d. (n = 3, biologically independent samples). e, CRISPR/Cas9-mediated gene editing of SCREW1 and SCREW2. Mutations of SCREW1 and SCREW2 in screw1/2-1 and screw1/2-2 were detected by DNA sequencing and shown as chromatographs. Two homozygous lines, screw1/2-1 and screw1/2-2, carry the same nucleotide insertion in SCREW1 for both lines, and a nucleotide insertion and a sixteen-nucleotide deletion in SCREW2 for screw1/2-1 and screw1/2-2, respectively. f, The screw1/2 mutants are morphologically indistinguishable from WT plants. Plants were grown on soil for four weeks and photographed. Scale bar, 1 cm. g, The screw1/2 mutants are more susceptible to Pst DC3000. Leaves of four-week-old WT and screw1/2 mutant lines (1 & 2) were hand-inoculated with Pst DC3000 at OD600 nm = 5 × 10−4. Bacterial numbers were measured at 0 and 3 dpi and shown as mean ± s.d. (n = 8, biologically independent samples). h, The screw1/2 mutants show enhanced susceptibility to Pst DC3000 hrcC −. Leaves of four-week-old soil-grown WT and screw1/2 mutant lines (1 & 2) were hand-inoculated with Pst DC3000 hrcC − at OD600 nm = 0.005, and the bacterial number was measured at 3 dpi. Data are shown as mean ± s.d. (n = 8, biologically independent samples). Experiments were repeated at least three times with similar results. Data were analysed by one-way (b-d, h) or two-way (g) ANOVA followed by Tukey’s test. Exact P values are provided in the graphs and Supplementary Table 3.
Extended Data Fig. 5 Identification of the SCREW receptor NUT.
a, Scheme to identify SCREW receptor candidates. Among 269 LRR-RKs in Arabidopsis, 26 members are upregulated by flg22 treatment, among which 11 members contain at least 18 LRRs in the extracellular domain. The cognate ligands of five of them remain unknown at the time of the study. b, NUT belongs to the XI subfamily of LRR-RK and is phylogenetically close to HAE and HSLs. Phylogenetic analysis of 52 LRR-RKs from the subfamily VII (orange curved line), X (purple curved line), XI (green curved line), and XII (grey curved line) is shown. Purple and red bars indicate the induction folds after elf18 and flg22 treatments, respectively. Olive green bars with numbers indicate the number of LRRs. Blue squares indicate cognate ligands of LRR-RKs. AT5G25930 (NUT) is highlighted in bold red font. The protein sequences were retrieved from NCBI (https://www.ncbi.nlm.nih.gov/) for MEGAX phylogenetic analysis using the neighbour-joining method with 1,000 bootstrap replicates. The phylogenetic tree was displayed by iTOL (https://itol.embl.de/). The expression data were from GENEVESTIGATOR V3. c, SCREWs and NUT are conserved in dicots and monocots. Protein sequences were blast-searched in NCBI using Arabidopsis SCREW1, NUT, or FLS2 as queries, and the phylogenetic analysis was performed as in (b). Red, purple, and olive curved lines indicate monocots, dicots, and other plant classes, respectively; Orange and teal bars indicate the percentage of homology of FLS2 and NUT in different plant species, respectively. Blue dots, olive stars, and red squares indicate FLS2, NUT, and SCREW homologs, respectively. Peach, lime green, grey, and brown fans denote different plant families. d, Diagram of AT5G25930 (NUT) with annotated T-DNA insertion sites in nut-1 (WiscDslox450B04) and nut-2 (SALK_207895). Solid bars indicate exons, lines for introns, and open boxes for UTRs. Arrows indicate primers used for genotyping. e, Genotyping of nut-1 and nut-2. The T-DNA insertions in the NUT coding region were PCR-amplified using genomic DNAs of WT, nut-1, or nut-2 as templates and primers shown in (d). f, RT–qPCR analysis of NUT transcripts. NUT expression levels in ten-day-old plate-grown seedlings were analysed using RT–qPCR with UBQ10 as an internal control. Data are shown as mean ± s.d. (n = 3, biologically independent samples) with one-way ANOVA followed by Tukey’s test. Exact P values are provided in the graph and Supplementary Table 3. Experiments were repeated three times with similar results (e, f).
Extended Data Fig. 6 NUT is required for SCREW-triggered responses and resistance to B. cinerea and green peach aphids.
a, NUT-Flag restores SCREW1-induced MAPK activation in nut protoplasts. Protoplasts were transfected with an empty vector (Ctrl) or NUT-Flag followed by treatment with 100 nM SCREW1 for 15 min. MAPK activation and NUT-Flag proteins were detected with anti-pERK1/2 (top) and anti-Flag antibodies (middle), respectively. Protein loading is shown by Ponceau S staining (Ponc.) for RBC (bottom). b, Inducible overexpressing SCREW1-triggered leaf chlorosis is blocked in nut-2. Five-week-old pEst::SCREW1-HA transgenic plants in WT and nut-2 were sprayed with 50 μM β-oestradiol and photographed five days later. Scale bar, 1cm. c, The nut mutants are more susceptible to Pst DC3000 than WT. Leaves of four-week-old plants were hand-inoculated with Pst DC3000 at OD600 nm = 5 × 10−4. The bacterial numbers were measured at 0 and 3 dpi. Data are shown as mean ± s.d. (n = 6, biologically independent samples). d, The nut and screw1/2 mutants are more susceptible to spray-inoculated Pst DC3000. Four-week-old plants were sprayed with Pst DC3000 at OD600 nm = 0.2. The disease symptoms and bacterial numbers were determined at 72 hpi. Data are shown as mean ± s.d. (n = 8, biologically independent samples). e, NUT is upregulated by MAMPs and pathogens. Data were retrieved from the Arabidopsis eFP browser (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi) and shown as histograms. Grey squares indicate no data available from eFP browser. Flg22, NLP, HrpZ, and lipopolysaccharide (LPS) are MAMPs. Pst DC3000, Pst DC3000 hrcC-, Pst DC3000 avrRpm1 and P. syringae pv. phaseolicola (Psp) are bacterial pathogens. Phytophthora infestans is an oomycete, and B. cinerea is a fungal pathogen. f, Flg22 upregulates NUT promoter activities. Two-week-old plate-grown pNUT::GUS/WT transgenic seedlings were treated without (Ctrl) or with 100 nM flg22 for 2 h, followed by GUS staining and microscopic imaging under a stereomicroscope. Scale bar, 2 mm. g, The nut and screw1/2 mutants do not affect plant resistance to Pst DC3000 avrRpm1 (left) and Pst DC3000 avrRpt2 (right). Bacteria were infiltrated into four-week-old plant leaves at OD600 nm = 0.001, and bacterial populations were determined at 2 dpi. Data are shown as mean ± s.d. (n = 6, biologically independent samples). h, The nut and screw1/2 mutants do not affect plant HR to Pst DC3000 avrRpm1 (left) and Pst DC3000 avrRpt2 (right). Bacteria were infiltrated into four-week-old plant leaves at OD600 nm = 0.08, and wilting leaves were counted at the indicated time points. At least 20 leaves were inoculated for each genotype and inoculum. The cell death rate was presented as the ratio of wilting leaves to total inoculated leaves. Data are shown as mean ± s.d. (n = 3, biologically independent samples). i, The nut and screw1/2 mutants are more susceptible to B. cinerea. Detached leaves of four-week-old plants were drop-inoculated with B. cinerea at 107 spores/mL. Disease phenotype was recorded at 48 hpi. Data are shown as mean ± s.d. (left to right: n = 30, 29, 27, 25, 28, biologically independent samples). j, The p35S::SCREW1 and p35S::SCREW2 plants show enhanced resistance to aphids. Six-age-synchronized second instar nymphs of Myzus persicae were inoculated onto leaves of four-week-old plants. The number of neonates per plant (n) was counted at 10 dpi. Data are shown as mean ± s.d. (left to right: n = 19, 12, 12, 12, 8, biologically independent samples). k, The nut mutants are more susceptible to aphid infections than WT plants. The experiments and statistics were performed as in j (left to right: n = 10, 10, 10, 12, 12, biologically independent samples). l, The expression of SCREWs and NUT is induced by aphid infections. Leaves of two-week-old plants were inoculated with or without aphid nymphs for 24, 48, and 72 h. The expression of SCREWs and NUT normalized to UBQ10 was analysed by RT–qPCR. Means (n = 3, biologically independent samples) of fold induction compared to non-treatment are shown as log2 transformation to construct heat map using TBtools. m, The NUT promoter activity is induced by aphid infections. Two-week-old soil-grown transgenic plants carrying pNUT::GUS were inoculated with aphid nymphs and subjected to GUS staining at 1 and 3 dpi. The pictures were taken under a stereomicroscope. Scale bar, 2 mm. Experiments except e were repeated three times with similar results. Data were analysed by one-way (d, i) or two-way (c, g) ANOVA followed by Tukey’s test, or one-way ANOVA followed by Dunnett’s test (j, k). Exact P values are provided in the graphs and Supplementary Table 3.
Extended Data Fig. 7 SPR assays of SCREW and SCREW derivatives binding to NUTECD, and bik1 does not affect SCREW1-induced inhibition of seedling growth.
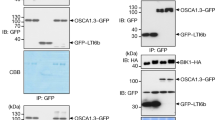
a, NUT-GFP is localized on the plasma membrane. Transgenic seedlings carrying p35S::GFP or p35S::NUT-GFP were grown on ½MS plates for seven days, and true leaves were imaged using a confocal laser scanning microscopy. Scale bar, 25 μm. b, NUT-GFP and SCREW2-GFP are localized on the plasma membrane in N. benthamiana. N. benthamiana leaves were infiltrated with A. tumefaciens GV3101 carrying p35S::NUT-GFP, p35S::SCREW2-GFP or p35S::GFP and imaged under a confocal microscope at 3 dpi. Scale bar, 25 μm. c, Sequence alignment of parts of cytosolic kinase domains of different RKs, including NUT, FLS2, BAK1, EFR, and BRI1. The sequences were aligned by MEGAX and visualized with ESPript 3.0 (http://espript.ibcp.fr/). The lysine (K) required for ATP-binding (blue asterisk) and the RD/non-RD motif (blue rectangle) were marked. The positions of amino acids in NUT were labelled on the top. d-g, SPR assays of SCREW1 (d), SCREW2 (e), SCREW1-HA (f), SCREW2CC/SS (g) binding to NUTECD under pH 5.7. NUTECD was immobilized on a sensor chip, and synthesized SCREW peptides were used as flow-through analytes. The top panel shows the SPR sensorgram profile of SCREW peptides at gradient concentrations flowing through the NUTECD-immobilized chip. The bottom panel shows the steady-state affinity (binding at equilibrium) with different Kd. h, The C-terminus of SCREW2 is essential for its binding to NUTECD. The SPR assays were performed as in Fig. 3f, g. i, BAK1 promotes the SCREW2-NUT binding affinity. (Left) SPR detection of BAK1ECD binding to NUTECD in the absence of SCREW peptides. BAK1ECD was used as flow-through analytes on a sensor chip immobilized with NUTECD. At the concentration tested, no binding between NUTECD and BAK1ECD was detected. (Right) SPR detection of SCREW2 binding to NUTECD in the presence of BAK1ECD. NUTECD was immobilized on a sensor chip, and SCREW2 peptides at gradient concentrations together with 0.1 µM BAK1ECD proteins were used as flow-through analytes. The Kd of SCREW2 binding to NUTECD in the presence of BAK1 is 0.38 ± 0.074 μM. The SPR assays were performed at pH 5.7. j, NUT does not associate with BIK1. Protoplasts were co-transfected with BIK1-HA and NUT-Flag, PEPR1-Flag, or a control vector (Ctrl). Total proteins were immunoprecipitated with anti-Flag agarose beads and detected with anti-HA or anti-Flag antibodies (top two). Proteins before immunoprecipitation (IP) are shown as input controls (bottom two). k, SCREW1 inhibits seedling growth in the bik1 mutant. Three-day-old plate-grown seedlings were transferred into liquid ½MS medium with or without 1 μM SCREW1 and grown for seven days. Fresh weights of seedlings are shown as as mean ± s.d. (n = 8, biologically independent samples). Data were analysed by two-sided Student’s t-test and two-way ANOVA followed by Tukey’s test. P values are provided in the graph and Supplementary Table 3. Experiments were repeated twice (a, b, d–i) or three times (j, k) with similar results.
Extended Data Fig. 8 SCREW–NUT does not affect flg22-triggered early signalling events but partially requires MIN7.
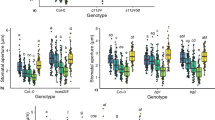
a, SCREW1 suppresses flg22- or ABA-induced stomatal closure. The stomatal apertures were measured after treatment without or with 1 μM SCREW1, 1 μM flg22, 10 μM ABA, or a combination of SCREW1 with flg22 or ABA for 2 h under the light. Data are shown as box plots with the interquartile range as the upper and lower confines, minima and maxima as whiskers, and the median as a solid line. Different letters denote a statistically significant difference (P < 0.05). n = number of stomata in the graph. b, SCREW1 does not affect the flg22-induced FLS2-BAK1 association. Four-day-old plate-grown WT seedlings were transferred into liquid ½MS for six days, followed by 100 nM flg22, 100 nM SCREW, or a combination of 100 nM flg22 and 100 nM SCREW for 10 min. Proteins were subjected for IP assays using anti-FLS2 antibodies and followed by immunoblotting using anti-BAK1 or anti-FLS2 antibodies. Top two panels show the IPed products, and bottom two panels show protein inputs. c, The nut and screw mutants do not affect the flg22-induced FLS2-BAK1 association. Four-day-old plate-grown seedlings were transferred into liquid ½MS for six days, followed by 100 nM flg22 for 10 min. Co-IP analysis was similar to the above in b. d, SCREW1 and NUT do not affect flg22-induced MAPK activation. Ten-day-old seedlings were treated with 100 nM flg22, 100 nM SCREW1, or a combination of 100 nM flg22 and 100 nM SCREW1. MAPK activation was analysed by immunoblotting with anti-pERK1/2 antibodies (top), and protein loading is shown by Ponceau S staining (Ponc.) for RBC (bottom). e, SCREW1-triggered resistance to Pst DC3000 is partially abolished in min7. Four-week-old plants were pre-infiltrated with 0.5 µM SCREW1, or H2O as a control. After 24 h, Pst DC3000 was infiltrated at a concentration of OD600 nm = 0.02 or 0.002, and bacterial counting was performed at 1 dpi for OD600 nm = 0.02, and 2 dpi for OD600 nm = 0.002. Data are shown as means ± s.d. (n = 3, biologically independent samples) (left). The picture was taken at 3 dpi with OD600 nm = 0.002 (right). f, MIN7 is not required for SCREW1 activation of MAPKs. Ten-day-old plate-grown seedlings were treated with 100 nM SCREW1. MAPK activation was analysed by immunoblots with anti-pERK1/2 antibodies (top), and the protein loading is shown by CBB staining for RBC (bottom). g, MIN7 is not required for SCREW1 suppression of seedling growth. Three-day-old plate-grown seedlings were transferred into liquid ½MS medium with or without 1 μM SCREW1. Seedlings were imaged (left) and weighed (right) seven days post-treatment. Scale bar, 1 cm. Fresh weights of seedlings are shown as box plots with the interquartile range as the upper and lower confines, minima and maxima as whiskers, and the median as a solid line (n = 12, biologically independent samples). h, The susceptibility of nut and screw1/2 mutants to Pst DC3000 is comparable to WT plants under high humidity with transient apoplast water supplementation. Leaves of four-week-old plants were inoculated with Pst DC3000 at OD600 nm = 1 × 10−4 and kept under 85–98% humidity for three days before bacterial counting. Transient water supplementation (+H2O) was performed by keeping plants under high humidity after syringe-infiltration without air-drying inoculated leaves as described previously32. Data are shown as means ± s.d. (n = 8, biologically independent samples). i, SCREW1-triggered plant resistance to Pst DC3000 is compromised under high humidity with transient apoplast water supplementation. Leaves of four-week-old WT plants were pre-infiltrated with H2O (Ctrl) or 1 µM SCREW1 for 24 h followed by Pst DC3000 inoculation. Plants were kept under 50% or 85–98% humidity for three days before bacterial counting. Data are shown as means ± s.d. (n = 8, biologically independent samples). Experiments were repeated three times with similar results. Data were analysed by two-sided Student’s t-test (e, g), one-way (a) or two-way (h, i) ANOVA followed by Tukey’s test. Exact P values are provided in the graphs and Supplementary Table 3.
Extended Data Fig. 9 SCREW–NUT regulates leaf water loss and ABA responses.
a, Transgenic plants carrying p35S::SCREW1 or p35S::SCREW2 exhibit curled leaves and increased sensitivity to dehydration stress. Leaves of five-week-old soil-grown plants were detached and imaged at 0 and 6 h after detachment. Scale bar, 1 cm. b, Increased water-loss rate in transgenic plants carrying p35S::SCREW1 or p35S::SCREW2. The rates of cumulative water loss from rosette leaves of five-week-old plants were measured at 6 h post-detachment. Data are shown as means ± s.d. (n = 6, biologically independent samples). c, Reduced water-loss rate in nut and screw1/2. The rate of cumulative water loss from detached leaves of four-week-old plants was measured at the indicated time points after detachment. Data are shown as means ± s.d. (n = 6, biologically independent samples). d, Enhanced resistance to mannitol treatment in nut mutants. Seedlings were grown on 1/2MS plates with 0, 50, 100, 150, or 200 mM mannitol for 15 days. Scale bar, 2 mm. e, Cuticle permeability of nut and screw1/2 seedlings is similar to that of WT. Three-week-old plate-grown plants were soaked with 0.05% toluidine blue for 15 min and washed with ddH2O before imaging. Scale bar, 1 cm. At least six seedlings for each genotype were analysed for the presence of the blue-coloured patches, which indicate an increased permeability of the stain into the leaf through the cuticle. No apparent differences were observed between WT and mutants. f, Leaf cuticle permeability of nut and screw1/2 is similar to that of WT. Leaves of four-week-old soil-grown plants were drop-stained with 0.05% toluidine blue on the adaxial surface for the indicated time. The red circles and rectangles indicate the sites of inoculation. Inserts show zoomed-in areas. No blue-coloured patches were observed, indicating the intact cuticle for each genotype. Scale bar, 20 µm. Scale bar, 1 cm. Ten leaves for each genotype were analysed. g, Leaf cuticle layers of nut and screw1/2 are similar to those of WT. Three-week-old plate-grown plant leaves were examined by transmission electron microscopy from the adaxial side. Red arrows indicate cuticles observed as a thin (~80–100 nm) electron-dense layer on the surface of the cell wall. Scale bar, 1 µM. Four leaves of each genotype were analysed. No apparent differences in thickness were detected among different genotypes. h, The nut and screw1/2 mutants are more sensitive to ABA treatment than WT plants. Seedlings were grown on 1/2MS plates without (Ctrl) or with 1 μM ABA for seven days (left). Cotyledon greening rates are shown as means ± s.d. (right, n = 4, biologically independent repeats). i, SCREW1 and SCREW2 suppress ABA-induced expression of RAB18 and RD29A in plants. Ten-day-old plate-grown WT seedlings were treated with H2O, 10 µM ABA, or combinations of 10 μM ABA and 1 μM SCREW1 or SCREW2 for 3 h. Transcript levels of RAB18 and RD29A normalized to UBQ10 were determined via RT–qPCR. Data are shown as mean ± s.d. (n = 4, biologically independent samples). j, NUT is upregulated by ABA, mannitol, and drought treatments. The expression data were extracted from Genevestigator V3 and shown as histograms. Grey squares indicate no data available. k, SCREWs and NUT are up-regulated after ABA treatments. Ten-day-old plate-grown WT seedlings were treated with 100 μM ABA for 0, 3, and 6 h. Transcript levels of SCREWs and NUT normalized to UBQ10 were determined via RT–qPCR. Data are shown as mean ± s.d. (n = 3, biologically independent samples). Experiments were repeated three times with similar results. Data were analysed by one-way (b) or two-way (c, h, i, k) ANOVA followed by Tukey’s test. Exact P values are provided in the graphs and Supplementary Table 3.
Extended Data Fig. 10 SCREW1 induces NUT-dependent ABI phosphorylation.
a, SCREW1 induces ABI2 phosphorylation. Protoplasts were transfected with ABI2-HA followed by treatment with 1 µM SCREW1 for the indicated time. Proteins were separated with Mn2+-Phos-tag (top two) or regular SDS-PAGE (middle two) and detected with anti-HA antibodies. The protein loading is shown by CBB staining for RBC. Signal intensities of the top two bands corresponding to the phosphorylated ABI2 (pABI2) normalized to the input ABI2 in the regular SDS-PAGE from six independent immunoblots were quantified by ImageJ (bottom). The phosphorylation of ABI2 without SCREW1 treatment was set as 1. Data are shown as mean ± s.d. (n = 6, biologically independent repeats). b, SCREW1 does not induce OST1 phosphorylation. Protoplasts were transfected with OST1-Flag, followed by treatment with 1 µM SCREW1 for the indicated time. Proteins were separated with Mn2+-Phos-tag (top two) or regular SDS-PAGE (middle two) and detected with anti-Flag antibodies. The protein loading is shown by CBB staining for RBC. Signal intensities of bands from three independent immunoblots were analysed by ImageJ (bottom). The relative phosphorylation of OST1 represents the ratio of phosphorylated to unphosphorylated OST1. Data are shown as mean ± s.d. (n = 3, biologically independent repeats). c, SCREW1 induces NUT-dependent ABI1 phosphorylation. Protoplasts were co-transfected with ABI1-HA and NUT-Flag or a control vector followed by treatment with 1 µM SCREW1 for the indicated time. Proteins were separated by SDS-PAGE with (top two) or without Mn2+-Phos-tag (middle four) and detected with anti-HA or anti-pERK1/2 antibodies. The protein loading is shown by CBB staining for RBC. Signal intensities of bands corresponding to phosphorylated ABI1 (pABI1) normalized to the input ABI1 in the regular SDS-PAGE from four independent immunoblots were quantified by ImageJ (bottom). The phosphorylation of ABI1 without SCREW1 treatment was set as 1. Data are shown as mean ± s.d. (n = 4, biologically independent repeats). Experiments were repeated three times with similar results. Data were analysed by one-way (a, b) or two-way (c) ANOVA followed by Tukey’s test. Exact P values are provided in graphs and Supplementary Table 3.
Extended Data Fig. 11 SCREW1 and ABI suppress ABA-induced OST1 phosphorylation.
a, ABI1 suppresses ABA-induced OST1 phosphorylation. Protoplasts were co-transfected with OST1-Flag and ABI1-HA with the indicated ratio of DNAs, followed by treatment with 1 µM SCREW1 for 5 min before adding 1 µM ABA for an additional 5 min. Proteins were separated with Mn2+-Phos-tag (top three) or regular SDS-PAGE (middle two) and detected with anti-HA or anti-Flag antibodies. The protein loading is shown by CBB staining for RBC. Signal intensities of bands corresponding to phosphorylated ABI1 (pABI1) and OST1 (pOST1) from four independent immunoblots were quantified by ImageJ (bottom two). The relative phosphorylation of OST1 represents the ratio of phosphorylated to unphosphorylated OST1. The phosphorylation of ABI1 without treatment was set as 1. Data are shown as mean ± s.d. (n = 4, biologically independent repeats). b, ABI2 suppresses ABA-induced OST1 phosphorylation. Protoplasts were co-transfected with ABI2-HA and OST1-Flag followed by treatment with 1 µM SCREW1 before adding 1 µM ABA. Proteins were separated with Mn2+-Phos-tag (top two) or regular SDS-PAGE (bottom three) and detected with anti-HA or anti-Flag antibodies. The protein loading is shown by CBB staining for RBC. Signal intensities of bands from four independent immunoblots were analysed by ImageJ (bottom). The relative phosphorylation of OST1 represents the ratio of phosphorylated to unphosphorylated OST1. Data are shown as mean ± s.d. (n = 4, biologically independent repeats). c, SCREW1 suppresses ABA-induced OST1 phosphorylation in a dosage-dependent manner. Protoplasts were transfected with OST1-Flag followed by treatments with 0, 10, 100 or 1,000 nM SCREW1 for 5 min before adding 1 µM ABA for another 5 min. Proteins were separated with Mn2+-Phos-tag (top two) and regular SDS-PAGE (middle two) and detected with anti-Flag antibodies. The protein loading is shown by CBB staining for RBC. Signal intensities of bands from four independent immunoblots were analysed by ImageJ. The relative phosphorylation of OST1 represents the ratio of phosphorylated to unphosphorylated OST1. Data are shown as mean ± s.d. (n = 4, biologically independent repeats). d, NUT interacts with ABI1 in protoplasts. Protoplasts were co-transfected with ABI1-HA and NUT-Flag or a control vector. Proteins were immunoprecipitated with anti-Flag agarose beads and detected with anti-HA or anti-Flag antibodies (top two). Proteins before IP are shown as input controls (bottom two). e, GST-NUTCD interacts with ABI1 in vitro. GST or GST-NUTCD proteins were immobilized on glutathione sepharose beads and incubated with His-ABI1-HA followed by immunoblotting with anti-HA antibodies (top). The protein loading is shown by CBB (bottom). Experiments were repeated three times with similar results. Data were analysed by one-way (b, c) or two-way (a) ANOVA followed by Tukey’s test. Exact P values are provided in the graphs and Supplementary Table 3.
Extended Data Fig. 12 SCREW1 suppresses flg22-induced stomatal closure through the ABI–OST1 module.
a, ABI1 and ABI2 are required for SCREW1 suppression of flg22-induced stomatal closure. The stomatal apertures from epidermal peels of WT and abi1-2/abi2-2 were measured after treatment without or with 1 μM SCREW1, 1 µM flg22, or a combination of SCREW1 and flg22 for 2 h. Data are shown as the box plots with the interquartile range as the upper and lower confines, minima and maxima as whiskers, and the median as a solid line (n = 202, the number of stomata). The different letters denote a statistically significant difference (P < 0.05, two-way ANOVA followed by the Tukey’s test). The effect of SCREW1 on flg22-induced stomatal closure in WT and abi1-2/abi2-2 was compared by a two-sided Student’s t-test. The experiment was repeated three times with similar results. Exact P values are provided in the graph and Supplementary Table 3. b, A model of SCREW–NUT in protecting plants against infections via promoting stomatal reopening and reducing apoplastic water levels. MAMP perception by PRRs induces stomatal closure to limit pathogen entry. Inevitably, stomatal closure increases the apoplastic water levels and creates aqueous habitats favourable for pathogen multiplication. To counteract, MAMP–PRR signalling induces the expression of SCREWs and NUT. Upon SCREW perception, NUT complexes with BAK1 and promotes stomatal reopening via regulating the ABI–OST1–SLAC1 phosphorylation module, thereby increasing water loss and reducing apoplastic water levels to prevent the pathogen colonization. To invade hosts, pathogens deliver effectors or toxins, some of which can open stomata. The blue lines indicate SCREW–NUT induction and function in plant immunity revealed in this study.
Supplementary information
Supplementary Table 1
Primers used in the study.
Supplementary Table 2
Synthetic Peptides used in this study.
Supplementary Table 3
P values.
Source data
Rights and permissions
About this article
Cite this article
Liu, Z., Hou, S., Rodrigues, O. et al. Phytocytokine signalling reopens stomata in plant immunity and water loss. Nature 605, 332–339 (2022). https://doi.org/10.1038/s41586-022-04684-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04684-3
This article is cited by
-
Enhancing drought, heat shock, and combined stress tolerance in Myrobalan 29C rootstocks with foliar application of potassium nitrate
BMC Plant Biology (2024)
-
Extracellular niche establishment by plant pathogens
Nature Reviews Microbiology (2024)
-
SCAB1 coordinates sequential Ca2+ and ABA signals during osmotic stress induced stomatal closure in Arabidopsis
Science China Life Sciences (2024)
-
Small secreted peptides (SSPs) in tomato and their potential roles in drought stress response
Molecular Horticulture (2023)
-
How plants manage pathogen infection
EMBO Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.