Abstract
Herpesviruses have mastered host cell modulation and immune evasion to augment productive infection, life-long latency and reactivation1,2. A long appreciated, yet undefined relationship exists between the lytic–latent switch and viral non-coding RNAs3,4. Here we identify viral microRNA (miRNA)-mediated inhibition of host miRNA processing as a cellular mechanism that human herpesvirus 6A (HHV-6A) exploits to disrupt mitochondrial architecture, evade intrinsic host defences and drive the switch from latent to lytic virus infection. We demonstrate that virus-encoded miR-aU14 selectively inhibits the processing of multiple miR-30 family members by direct interaction with the respective primary (pri)-miRNA hairpin loops. Subsequent loss of miR-30 and activation of the miR-30–p53–DRP1 axis triggers a profound disruption of mitochondrial architecture. This impairs induction of type I interferons and is necessary for both productive infection and virus reactivation. Ectopic expression of miR-aU14 triggered virus reactivation from latency, identifying viral miR-aU14 as a readily druggable master regulator of the herpesvirus lytic–latent switch. Our results show that miRNA-mediated inhibition of miRNA processing represents a generalized cellular mechanism that can be exploited to selectively target individual members of miRNA families. We anticipate that targeting miR-aU14 will provide new therapeutic options for preventing herpesvirus reactivations in HHV-6-associated disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The sequencing datasets produced in this study have been deposited at the Gene Expression Omnibus with accession number GSE179867. BAC sequencing results have been deposited at the NCBI BioProject database with the BioProject ID PRJNA792929. Raw experimental data have been deposited at Mendeley (https://doi.org/10.17632/grnz4krxp2.3). Source data are provided with this paper.
References
Paludan, S. R., Bowie, A. G., Horan, K. A. & Fitzgerald, K. A. Recognition of herpesviruses by the innate immune system. Nat. Rev. Immunol. 11, 143–154 (2011).
York, I. A. Immune evasion strategies of the herpesviruses. Chem. Biol. 3, 331–335 (1996).
Cohen, J. I. Herpesvirus latency. J. Clin. Invest. 130, 3361–3369 (2020).
Pfeffer, S. et al. Identification of virus-encoded microRNAs. Science 304, 734–736 (2004).
Bushati, N. & Cohen, S. M. microRNA functions. Annu. Rev. Cell Dev. Biol. 23, 175–205 (2007).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Bartel, D. P. Metazoan MicroRNAs. Cell 173, 20–51 (2018).
Cai, X. et al. Kaposi’s sarcoma-associated herpesvirus expresses an array of viral microRNAs in latently infected cells. Proc. Natl Acad. Sci. USA 102, 5570–5575 (2005).
Kaufer, B. B. & Flamand, L. Chromosomally integrated HHV-6: impact on virus, cell and organismal biology. Curr. Opin. Virol. 9C, 111–118 (2014).
Agut, H., Bonnafous, P. & Gautheret-Dejean, A. Laboratory and clinical aspects of human herpesvirus 6 infections. Clin. Microbiol. Rev. 28, 313–335 (2015).
Mishra, P. & Chan, D. C. Metabolic regulation of mitochondrial dynamics. J. Cell Biol. 212, 379–387 (2016).
Chowdhury, S. R. et al. Chlamydia preserves the mitochondrial network necessary for replication via microRNA-dependent inhibition of fission. J. Cell Biol. 216, 1071–1089 (2017).
Smirnova, E., Griparic, L., Shurland, D. L. & van der Bliek, A. M. Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol. Biol. Cell 12, 2245–2256 (2001).
Kalia, R. et al. Structural basis of mitochondrial receptor binding and constriction by DRP1. Nature 558, 401–405 (2018).
Li, J. et al. miR-30 regulates mitochondrial fission through targeting p53 and the dynamin-related protein-1 pathway. PLoS Genet. 6, e1000795 (2010).
Prusty, B. K. et al. HHV-6 encoded small non-coding RNAs define an intermediate and early stage in viral reactivation. NPJ Genom. Med. 3, 25 (2018).
Nukui, M., Mori, Y. & Murphy, E. A. A human herpesvirus 6A-encoded microRNA: role in viral lytic replication. J. Virol. 89, 2615–2627 (2015).
Xu, X., Zhao, P. & Chen, S. J. Vfold: a web server for RNA structure and folding thermodynamics prediction. PLoS ONE 9, e107504 (2014).
Sharma, C. M. & Vogel, J. Differential RNA-seq: the approach behind and the biological insight gained. Curr. Opin. Microbiol. 19, 97–105 (2014).
Whisnant, A. W. et al. Integrative functional genomics decodes herpes simplex virus 1. Nat. Commun. 11, 2038 (2020).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Lee, Y. & Kim, V. N. In vitro and in vivo assays for the activity of Drosha complex. Methods Enzymol. 427, 89–106 (2007).
Tiku, V., Tan, M. W. & Dikic, I. Mitochondrial functions in infection and immunity. Trends Cell Biol. 30, 263–275 (2020).
West, A. P. et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature 520, 553–557 (2015).
Rasmussen, S. B. et al. Herpes simplex virus infection is sensed by both Toll-like receptors and retinoic acid-inducible gene- like receptors, which synergize to induce type I interferon production. J. Gen. Virol. 90, 74–78 (2009).
Samanta, M., Iwakiri, D. & Takada, K. Epstein–Barr virus-encoded small RNA induces IL-10 through RIG-I-mediated IRF-3 signaling. Oncogene 27, 4150–4160 (2008).
Castanier, C., Garcin, D., Vazquez, A. & Arnoult, D. Mitochondrial dynamics regulate the RIG-I-like receptor antiviral pathway. EMBO Rep. 11, 133–138 (2010).
Hornung, V. et al. 5′-triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006).
Liu, X. et al. MicroRNA-148/152 impair innate response and antigen presentation of TLR-triggered dendritic cells by targeting CaMKIIα. J. Immunol. 185, 7244–7251 (2010).
Newman, M. A., Thomson, J. M. & Hammond, S. M. Lin-28 interaction with the Let-7 precursor loop mediates regulated microRNA processing. RNA 14, 1539–1549 (2008).
Rybak, A. et al. A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nat. Cell Biol. 10, 987–993 (2008).
Viswanathan, S. R., Daley, G. Q. & Gregory, R. I. Selective blockade of microRNA processing by Lin28. Science 320, 97–100 (2008).
Treiber, T. et al. A compendium of RNA-binding proteins that regulate microRNA biogenesis. Mol. Cell 66, 270–284.e213 (2017).
Zisoulis, D. G., Kai, Z. S., Chang, R. K. & Pasquinelli, A. E. Autoregulation of microRNA biogenesis by let-7 and Argonaute. Nature 486, 541–544 (2012).
Tang, R. et al. Mouse miRNA-709 directly regulates miRNA-15a/16-1 biogenesis at the posttranscriptional level in the nucleus: evidence for a microRNA hierarchy system. Cell Res. 22, 504–515 (2012).
Wang, D. et al. Nuclear miR-122 directly regulates the biogenesis of cell survival oncomiR miR-21 at the posttranscriptional level. Nucleic Acids Res. 46, 2012–2029 (2018).
Patel, N., Tahara, S. M., Malik, P. & Kalra, V. K. Involvement of miR-30c and miR-301a in immediate induction of plasminogen activator inhibitor-1 by placental growth factor in human pulmonary endothelial cells. Biochem. J. 434, 473–482 (2011).
Mori, J. et al. Human herpesvirus-6 U14 induces cell-cycle arrest in G2/M phase by associating with a cellular protein, EDD. PLoS ONE 10, e0137420 (2015).
Takemoto, M. et al. Human herpesvirus 6 open reading frame U14 protein and cellular p53 interact with each other and are contained in the virion. J. Virol. 79, 13037–13046 (2005).
Sever, J. L. et al. Virus antibodies and multiple sclerosis. Arch. Neurol. 24, 489–494 (1971).
Schreiner, P. et al. Human herpesvirus-6 reactivation, mitochondrial fragmentation, and the coordination of antiviral and metabolic phenotypes in myalgic encephalomyelitis/chronic fatigue syndrome. Immunohorizons 4, 201–215 (2020).
Gulve, N., Frank, C., Klepsch, M. & Prusty, B. K. Chromosomal integration of HHV-6A during non-productive viral infection. Sci. Rep. 7, 512 (2017).
Prusty, B. K., Krohne, G. & Rudel, T. Reactivation of chromosomally integrated human herpesvirus-6 by telomeric circle formation. PLoS Genet. 9, e1004033 (2013).
Naldini, L. et al. In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector. Science 272, 263–267 (1996).
Wiznerowicz, M. & Trono, D. Conditional suppression of cellular genes: lentivirus vector-mediated drug-inducible RNA interference. J. Virol. 77, 8957–8961 (2003).
Tang, H. et al. Human herpesvirus 6 encoded glycoprotein Q1 gene is essential for virus growth. Virology 407, 360–367 (2010).
Tischer, B. K., von Einem, J., Kaufer, B. & Osterrieder, N. Two-step red-mediated recombination for versatile high-efficiency markerless DNA manipulation in Escherichia coli. Biotechniques 40, 191–197 (2006).
Wallaschek, N. et al. The telomeric repeats of human herpesvirus 6A (HHV-6A) are required for efficient virus integration. PLoS Pathog. 12, e1005666 (2016).
Tischer, B. K. & Kaufer, B. B. Viral bacterial artificial chromosomes: generation, mutagenesis, and removal of mini-F sequences. J. Biomed. Biotechnol. 2012, 472537 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Ingerman, E. et al. Dnm1 forms spirals that are structurally tailored to fit mitochondria. J. Cell Biol. 170, 1021–1027 (2005).
Mears, J. A. et al. Conformational changes in Dnm1 support a contractile mechanism for mitochondrial fission. Nat. Struct. Mol. Biol. 18, 20–26 (2011).
Rosenbloom, A. B. et al. Optimized two-color super resolution imaging of Drp1 during mitochondrial fission with a slow-switching Dronpa variant. Proc. Natl Acad. Sci. USA 111, 13093–13098 (2014).
Ji, W. K., Hatch, A. L., Merrill, R. A., Strack, S. & Higgs, H. N. Actin filaments target the oligomeric maturation of the dynamin GTPase Drp1 to mitochondrial fission sites. eLife 4, e11553 (2015).
Prusty, B. K., Chowdhury, S. R., Gulve, N. & Rudel, T. Peptidase inhibitor 15 (PI15) regulates chlamydial CPAF activity. Front. Cell. Infect. Microbiol. 8, 183 (2018).
Gulve, N. et al. Anti-herpesviral effects of a novel broad range anti-microbial quaternary ammonium silane, K21. Antiviral Res. 131, 166–173 (2016).
Balachandran, N., Amelse, R. E., Zhou, W. W. & Chang, C. K. Identification of proteins specific for human herpesvirus 6-infected human T cells. J. Virol. 63, 2835–2840 (1989).
Kim, Y. K., Yeo, J., Kim, B., Ha, M. & Kim, V. N. Short structured RNAs with low GC content are selectively lost during extraction from a small number of cells. Mol. Cell 46, 893–895 (2012).
Brown, B., Venneri, M., Zingale, A., Sergi, L. & Naldini, L. Endogenous microRNA regulation suppresses transgene expression in hematopoietic lineages and enables stable gene transfer. Nat. Med. 12, 585–591 (2006).
Jurges, C. S., Dolken, L. & Erhard, F. Integrarative transcription start site identification with iTiSS. Bioinformatics https://doi.org/10.1093/bioinformatics/btab170 (2021).
Hauptmann, J. et al. Biochemical isolation of Argonaute protein complexes by Ago-APP. Proc. Natl Acad. Sci. USA 112, 11841–11845 (2015).
Acknowledgements
We thank the Core Unit for Confocal Microscopy and Flow Cytometry-based Cell Sorting of the IZKF Würzburg for supporting the study; the Core Unit Systemmedizin of the JMU Würzburg for dRNA-seq analysis; S. R. Chowdhury for his help with initial SIM microscopy and mitochondrial data analysis; Y. Mori; E. Murphy for HHV-6A BAC constructs and reconstituted virus preparations; and J. Vogel for constructive suggestions during manuscript preparations. We acknowledge the Helmholtz Institute for RNA-based Infection Research (HIRI) for supporting this work with a seed grant to BKP through funds from the Bavarian Ministry of Economic Affairs and Media, Energy and Technology (Grant allocation nos 0703/68674/5/2017 and 0703/89374/3/2017). This work was supported by the European Research Council (ERC-2016-CoG 721016–HERPES to L.D. and ERC-2015-StG 677673–INTEGHER to B.B.K.). Additional support came from the Deutsche Forschungsgemeinschaft (FOR 2830–Do 1275/7-1 and FOR 5200–Do 1275/12-1 to L.D.). We also thank HHV-6 Foundation, USA for the Dharam Ablashi pilot research grant to B.K.P., Solve ME/CFS Initiative, USA for the Ramsay Research Award and Amar Foundation USA career development grant to B.K.P.
Author information
Authors and Affiliations
Contributions
B.K.P. and L.D. conceived the idea, developed and supervised the project, and wrote the manuscript. B.K.P. carried out majority of experiments supported by T.H. and A.B.P. T.H. and J.T. carried out interferon assays. A.B.P. designed expression constructs and carried out miRNA quantification assays. F.K. developed mutant virus carrying cells. A.E. carried out miRNA-mimic Northerrn blots. A.G. carried out immunoblotting experiments. A.W.W. contributed to miRNA interaction studies. M.L. carried out FACS analysis. C.J. and S.H. carried out the RNA-seq experiment and data analysis. B.B.K. designed and developed the HHV-6A mutant BAC. T.K. and M.S. carried out SIM microscopy and analysed the SIM data. F.E. analysed the sequencing data. G.M., U.F. and T.R. contributed essential reagents and analysed the data. All authors contributed to manuscript writing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Felicia Goodrum and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 HHV-6A reactivation induces mitochondrial fission via the p53/DRP1 axis.
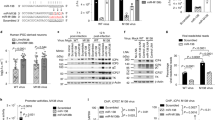
a. Mitochondria in lytic HHV-6A infected primary HUVEC cells expressing soluble GFP within mitochondrial lumen (mitoGFP). Expression of HHV-6 p41 protein was tested for virus infection. Representative images from 3 independent experiments. n = 3. b. HHV-6A reactivation in U2OS cells expressing mitoGFP and carrying latent HHV-6A genome was stimulated for 24 or 48 h using trichostatin A (TSA) (T) or DMSO as solvent control (C). Cells were either fixed after one (C1 or T1) or two (C2 or T2) days of treatment and studied using confocal microscopy. Representative confocal images of cells two days post treatment are shown. Mitochondrial number and average surface area as quantified from confocal images are presented as box and whiskers plot. Immunostaining against HHV-6 p41 protein was carried out to identify virus reactivation. n = 9 images per conditions. **P = 0.005, ****P < 0.001. c. Virus reactivation in U2OS cells with or without latent HHV-6A was induced for 48 h using TSA (T) or solvent control (C). Changes in DRP1 and p53 protein expression were quantified using densitometric analysis of immunoblots. GAPDH normalized DRP1 protein levels are presented as bar diagram. n = 3 per condition. *P = 0.04. d. Structured illumination microscopy (SIM) was used to quantify DRP1 rings around mitochondria. Cells from the above experiment were processed for SIM after DRP1 immunostaining. Enlarged DRP1 rings are shown within insets. DRP1 rings per cell together with Pearson’s overlap co-efficient were quantified from the obtained images. *P = 0.01, **P = 0.005. Box and whisker plots (b, d) show minimum to maximum values with all independent replicates, centre denotes median, and the bounds denote the 25th to 75th percentiles. Data are mean ± SD. Two-tailed non-parametric t-test (b-d). The uncropped blots are provided in Supplementary Fig. 1.
Extended Data Fig. 2 HHV-6A lytic infection and reactivation impairs miR-30 processing.
a. HSB-2 cells were infected for 48 h with HHV-6A for lytic virus infection. Total RNA from mock- and HHV-6A-infected cells were analyzed by Northern blotting. n = 3. b. Experiment similar to ‘a’ from three independent replicate experiments are shown. RNA decade marker (DM) and in vitro transcribed (IVT) RNA (300 nt) were run as size marker. c. HHV-6A reactivation in U2OS cells carrying latent HHV-6A was induced for 48 h using TSA. Cells without HHV-6A served as mock. The pri-miR-30c to mature miR-30c ratios, as determined by densitometric analysis, are shown as bar diagrams. n = 3. *P = 0.04. d. Virus reactivation of U2OS cells carrying latent HHV-6A was induced for 48 h using TSA. RNA from three independent replicates are shown. Cells without HHV-6A served as mock. Blots for human U6, miR-aU14 and sncRNA-U77 are shown only once as the same blot was stripped and used for reprobing against multiple miR-30 family members. All the images shown are from a single gel. HHV-6 miR-aU14 and sncRNA-U77 were probed to confirm virus reactivation. A probe against human U6 was used as loading control. n = 3. e. MA plot showing normalized log2 fold changes in miR-30 family members from HSB-2 cells. Upper panel shows read counts of major 5’ miRNAs. The lower panel shows read counts of minor miR-30 members 3′ miRNAs. n = 3. A prominent band of 80-90 nt was detected on miR-30d blots using LNA probes and are marked as ? as no known miR-30d products are known to have such sizes. Data are mean ± SD. two-tailed non-parametric t-test (c). The uncropped blots are provided in Supplementary Fig. 1.
Extended Data Fig. 3 HHV-6A infection and impairment of miR-30 biogenesis.
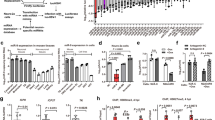
a. HHV-6A lytic infection impairs miR-30 biogenesis. HSB-2 cells were infected for 48 h with HHV-6A for lytic virus infection. Total RNA from mock- and HHV-6A-infected cells were analyzed for pri-miRNA/pre-miRNA and mature miRNA expression by qRT-PCR using TaqMan primers. n = 5 independent experiments. *P < 0.05, ***P < 0.004. b. HHV-6A reactivation impairs miR-30 biogenesis. HHV-6A reactivation in U2OS cells carrying latent HHV-6A was induced for 48 h using TSA. Solvent control (DMSO) treated cells served as mock. n = 3 independent experiments. *P < 0.04, **P < 0.08, ***P < 0.0001. pri-/pre-miR values represents the sum of primary (pri-) and precursor (pre-) miRNAs as TaqMan qPCR primers can’t differentiate between the two miRNA intermediates. c and d. Human NFYC mRNA levels were studied in the same samples as in Fig. a and b respectively. Cq values for mature miRNA amplification were normalized with U6 snRNA. Cq values for pri-/pre-miRNAs were normalized with beta actin. Furthermore, fold change values of virus infected samples were normalized against the same from mock infected samples and TSA treated samples were normalized against mock (solvent control) treated samples. NFYC Cq values were normalized aginst beta actin. Data are mean ± SD. Two-tailed non-parametric t-test (a-d). e. Schematic of potential binding site of miR-aU14 in the hairpin loop of different miR-30 family members. Sequence complementarity between miR-aU14 and pre-miR-30a, pre-miR-30d. Potential RNA-RNA interaction regions are highlighted in grey. Mature miR-30 sequences are indicated in red. miR-aU14 is shown in blue. f, g. Two different possible conformations (f and g) for the interaction between miR-aU14 and pre-miR-30c were predicted by VfoldCPX 2D RNA analysis. Free binding energies for respective conformations as predicted by the software are indicated.
Extended Data Fig. 4 Characterization of HHV-6A encoded miR-aU14.
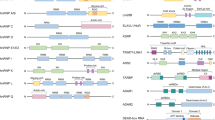
a. Identification of AGO-bound viral miRNAs during lytic HHV-6A infection. HSB-2 cells were infected for 72 h with wild-type HHV-6A followed by human AGO immunoprecipitation and small RNA sequencing. n = 2 independent experiments. b. dRNA-seq normalized read counts from HSB-2 cells infected with HHV-6A (n = 2) are shown for both the sense (upper panel) and antisense (lower panel) strand. Location of miR-aU14 and U14 ORF as well as the identified transcription start sites (TSS) of the U14 mRNA (highlighted as ‘A’ in blue) and of pri-miR-aU14 (highlighted as ‘B’ in red) are indicated against the GenBank accession number X83413.2. TSS identified pri-miR-aU14 to initiate 158 nt upstream of miR-aU14 within the first third of the U14 ORF. The TSS that we identified for the U14 mRNA indicates that translation of the U14 ORF only initiates at a second AUG (indicated) that is located 15 nt downstream of the reference AUG. c. Predicted folding of pre-miR-aU14. The mature miR-aU14 sequence is highlighted in red. d. Putative miR-aU14 sequences from two different strains of HHV-6A (HHV-6A U1102, GenBank X83413.2; HHV-6A GS, GenBank KC465951.1) and one strain of HHV-6B (HHV-6B Z29, GenBank AF157706.1) were aligned. Sequence homology of mature miR-aU14 region is highlighted with red rectangles.
Extended Data Fig. 5 Viral inhibition of miR-30c processing triggers mitochondrial fragmentation.
a. Doxycycline-(Dox)-inducible wild-type and mutant miR-aU14 shRNAs were expressed in HeLa cells under control of a dox-inducible Pol III promoter. Mature miR-aU14 sequence are indicated in red. Point mutations are highlighted in blue within a black rectangle. b. HeLa cells were stably transduced to express dox-inducible miR-aU14. miR-30c and miR-aU14 expression was probed by Northern blotting. Probe against human U6 served as loading control. Ratio of pri-miR-30c/miR-30c, as determined by densitometric quantification, are shown. n = 3. *P = 0.04 c. Polyclonal HeLa cells stably transduced to express either wild-type (HeLa-Wt1, 2 and 3) or mutant (HeLa-Mut1 and 2) miR-aU14 were induced. miR-30c and miR-aU14 expression was probed by Northern blotting. d. miR-aU14 mimic induces DRP1 protein expression. Total protein lysates from cells transfected with either a control mimic, miR-aU14 mimic (Wt mimic), or the mutant miR-aU14 mimic (Mut mimic) were analysed by immunoblotting for DRP1 and p53 protein levels. *P = 0.04. e. Total protein lysates from polyclonal HeLa cells with dox-inducible wild-type (HeLa-Wt1) or mutant miR-aU14 (HeLa-Mut1) were subjected to immunoblot analysis. DRP1 protein levels were quantified by densitometric analysis and were normalized against GAPDH. *P = 0.01. f. Mitochondrial architecture was studied in U2OS cells transfected either with miR-aU14 mimic (Wt mimic) or the mutant miR-aU14 mimic (Mut mimic). In upper panel, fixed cells were used for confocal microscopy. n = 4. In lower panel, SIM images co-stained with DRP1 are shown. Average mitochondrial area and mitochondrial numbers per cell graphs were generated from confocal images. DRP1 aggregates per cell and associated Pearson’s overlap co-efficient were generated from SIM images. *P < 0.04. Magnified areas of mitochondria are shown within insets. Box and whisker plots (f) show minimum to maximum values with all independent replicates, centre denotes median, and the bounds denote the 25th to 75th percentiles. Data are mean ± SD. Two-tailed non-parametric t-test (b, d, e, f). The uncropped blots are provided in Supplementary Fig. 1.
Extended Data Fig. 6 Characterization of miR-aU14 mutant HHV-6A virus.
a. The miR-aU14 sequence was altered in the HHV-6A BAC without disturbing the amino acid (aa) sequences of the U14 ORF on the opposing strand. Sequence information at the site of mutation is shown. b. HHV-6A reactivation of U2OS cells carrying latent wild-type (HHV-6A-Wt) or miR-aU14 mutant (HHV-6A-Mut) HHV-6A genomes was stimulated for 48 h using TSA. U2OS cells without latent HHV-6A served as mock. Total protein lysates were subjected to immunoblot analysis. GAPDH was used as loading control. Normalized fold change in DRP1 protein levels are presented as a bar diagram. n = 2. c. Mitochondrial architecture in wild-type or mutant HHV-6A reactivating cells (green arrowheads) carrying stable RFP within mitochondria (mitoRFP). The BAC-derived HHV-6A backbone carries a GFP expression cassette facilitating expression of GFP upon virus reactivation. DAPI served as counterstaining in confocal imaging. n = 3 independent experiments with at least 2 images per condition. Data are mean ± SD. two-tailed non-parametric t-test (b). The uncropped blots are provided in Supplementary Fig. 1.
Extended Data Fig. 7 miR-aU14 mimic interferes with in vitro processing of pri-miR-30 and pre-miR-30.
a. In vitro pri-miR-30c processing is inhibited by wild-type miR-aU14 mimic. HEK 293 cells were transfected with Flag-DGCR8 or a mock construct. Total cell lysate from both mock and Flag-DGCR8 expressing cells were used to process in vitro transcribed radiolabelled pri-miR-30c. Immunoprecipitated Flag-DGCR8 protein complex or mock IPs lysates were used in parallel. Either wild-type miR-aU14 mimic or a mutant mimic (Mut2 mimic) was used in the reaction to see their effect on miRNA biogenesis. b. Similar assays were carried out using in vitro transcribed pri-miR-30a. Immunoprecipitated human AGO protein complex was added to the reaction separately to check if it can help miR-aU14 to bind to pri-miR-30a and interfere with DGCR8 binding. c. miR-aU14 was able to interfere with pre-miR-30c processing by AGO:DICER complex. d. miR-aU14 was able to interfere with pre-miR-30a processing by AGO:DICER complex. e. Flag-DGCR8 immunoprecipitation was verified by immunoblotting. f. AGO immunoprecipitation was verified by immunoblotting. DM, Decade RNA marker. n = 2 (a-f). Same IP lysates were used for both replicates. The uncropped blots are provided in Supplementary Fig. 1.
Extended Data Fig. 8 miR-aU14 regulates virus lytic/latent switch.
a. U2OS cells carrying wild-type HHV-6A miR-aU14 (HHV-6A-Wt) were transfected with either the miR-aU14 mimic or a control mimic. Cells were induced with TSA and the extent of virus reactivation was analyzed by Northern blotting for viral miR-U2, miR-U86 and sncRNA-U77. miR-30c levels were checked for validating effects of miR-aU14 mimic on miR-30c processing. Human U6 normalized miR-U86 levels from six independent experiments are presented in a bar diagram. n = 6 independent experiments. *P < 0.02. b. U2OS cells carrying latent wild-type HHV-6A were transfected with miR-aU14 mimic or the two mutant miRNA mimics. Cells were induced with TSA and the extent of virus reactivation was analyzed by Northern blotting for viral miR-U86, miR-U2. n = 2. A probe against human U6 was used as loading control. Data are mean ± SD. Two-tailed non-parametric t-test (a). The uncropped blots are provided in Supplementary Fig. 1.
Extended Data Fig. 9 miR-30c inhibition induces p53/DRP1 axis and allows better HHV-6A growth.
a. miR-30c sponge efficiently downregulates miR-30c. U2OS cells were exposed to miR-30c inhibitor or miR-30c sponge lentivirus. 3 days later miR-30c levels were analyzed by Northern blotting. n = 3. b. Both miR-30c sponge and 30c inhibitor induces p53/DRP1 axis. U2OS cells were exposed to miR-30c sponge lentivirus or miR-30c inhibitor in presence of mock or p53 siRNA. 3 days later, p53 and DRP1 levels were studied by immunoblotting. GAPDH was used as a loading control. DRP1 levels are shown as a scatter plot. n = 3. *P = 0.04. c. Mitochondrial architecture was compared in U2OS cells in response to miR-aU14 Wt mimic or miR-30c sponge or miR-30c inhibitor. Average mitochondrial area are presented as a scatter plot. Magnified images of mitochondria are shown within insets. *P = 0.01. d. miR-30c sponge decreases Interferon beta (IFNβ) response. U2OS cells transduced with miR-30c sponge were transfected with hpRNA and the induced IFNβ response was analyzed after normalization against IFNβ mRNA in cells without hpRNA transfection. *P = 0.03. e. Effect of miR-30c sponge and HHV-6A infection on miR-30c biogenesis. HEK 293 cells were transduced with miR-30c sponge and were infected with cell free HHV-6A at MOI of 5 for 48 h. Total RNA was used for Northern blotting. U6 was used as loading control. n = 2. f. Effect of miR-30c mimic on HHV-6A lytic infection. HSB-2 cells were transfected with either a control or the miR-aU14 mimic (Wt mimic) together with miR-30c mimic. HHV-6A mCherry reporter virus was used to measure cell to cell spread of virus infection by flow cytometry. n = 2. *P = 0.03. g. miR-30c sponge alone reactivates mutant HHV-6A. U2OS cells carrying no virus (mock) or miR-aU14 mutant HHV-6A (HHV-6A-Mut) are transduced with miR-30c sponge lentivirus with or without TSA. Virus reactivation was assessed by measuring virus sncRNA-U77. n = 1. Data are mean ± SD. Two-tailed non-parametric t-test (b, c, d, f). The uncropped blots are provided in Supplementary Fig. 1. Gating Strategy for Extended Data Fig. 9f is provided in Supplementary Fig. 2.
Extended Data Fig. 10 Human miRNA processing can be selectively inhibited by synthetic small RNAs.
a. Schematic of putative binding of two different synthetic miRNAs to human pre-let-7f1. b. Targeting the hairpin loop of pre-let-7f1 with synthetic miRNAs interferes with miRNA processing. Two different miRNA mimics designed against the hairpin loop of pre-let-7f1 or control mimic were transfected into U2OS cells. let-7d miRNA levels were quantified by Northern blotting. A probe against human U6 was used as loading control. Pre-let-7f1 / let-7f1 ratio, as determined by densitometric analysis, are shown in the bar diagram. n = 4 independent replicates. Data are mean ± SD. *P = 0.03, two-tailed non-parametric t-test (b). c. Sequence details of miR-155 mimic and the mutant miR-155. Altered sequences are highlighted within a yellow box and are marked in red font. d. In vitro processing of pri-148b is inhibited by wild-type miR-155. HEK 293 cells were transfected with Flag-DGCR8 or a mock construct. Total cell lysate from both mock and Flag-DGCR8 expressing cells were used to process in vitro transcribed radiolabelled pri-miR-148b. Immunoprecipitated Flag-DGCR8 protein complex or mock IPs lysates were used in parallel. Either wild-type miR-155 mimic (155 mimic) or a mutant mimic (155 mut mimic) was used in the reaction to see it’s effect on miRNA biogenesis. n = 2 independent experiments. DM, Decade RNA marker. Predicted pre-miR-148b size is 99 nt, which was barely detectable in our assay. Interestingly, in this assay, the miR-155 mimic resulted in the appearance of an alternative processing intermediate of about 100 nt, which matched the 99 nt predicted miR-148b pre-miRNA included in miR-Base. While this 99 nt processing intermediate does not match to the actual cleavage site of mature miR-148b, this finding provides additional evidence that miR-155 inhibits pri-miR-155 processing resulting in alternative pri-miR-148b processing. The uncropped blots are provided in Supplementary Fig. 1.
Extended Data Fig. 11 Graphical abstract of regulation of miRNA-mediated miRNA processing.
Schematic of regulation of miRNA processing by other miRNAs.
Supplementary information
Supplementary Figure 1
This file contains raw, uncropped data for gels. Areas outlined by a dotted box were used to assemble figure panels.
Supplementary Figure 2
Gating strategy for flow cytometry analysis to obtain live cells (fixed) (SSC vs FSC) indicated by the sub-population representing ‘cells’, which were further screened for mCherry expression by analysing SSC vs FL-3 (mCherry) where the third quadrant represents the mCherry positive population.
Supplementary Tables
This file contains Supplementary Tables 1–3. Supplementary Table 1: List of plasmids used in the study; Supplementary Table 2: List of primers, oligonucelotides and synthetic RNA oligonucelotides used in the study; Supplementary Table 3: List of antibodies used in the study.
Source data
Rights and permissions
About this article
Cite this article
Hennig, T., Prusty, A.B., Kaufer, B.B. et al. Selective inhibition of miRNA processing by a herpesvirus-encoded miRNA. Nature 605, 539–544 (2022). https://doi.org/10.1038/s41586-022-04667-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04667-4
This article is cited by
-
Integrated analysis of microRNA and messenger RNA expression profiles reveals functional microRNA in infectious bovine rhinotracheitis virus-induced mitochondrial damage in Madin-Darby bovine kidney cells
BMC Genomics (2024)
-
Post-COVID-Syndrom mit Fatigue und Belastungsintoleranz: Myalgische Enzephalomyelitis bzw. Chronisches Fatigue-Syndrom
Die Innere Medizin (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.