Abstract
Adhesion G-protein-coupled receptors (aGPCRs) are important for organogenesis, neurodevelopment, reproduction and other processes1,2,3,4,5,6. Many aGPCRs are activated by a conserved internal (tethered) agonist sequence known as the Stachel sequence7,8,9,10,11,12. Here, we report the cryogenic electron microscopy (cryo-EM) structures of two aGPCRs in complex with Gs: GPR133 and GPR114. The structures indicate that the Stachel sequences of both receptors assume an α-helical–bulge–β-sheet structure and insert into a binding site formed by the transmembrane domain (TMD). A hydrophobic interaction motif (HIM) within the Stachel sequence mediates most of the intramolecular interactions with the TMD. Combined with the cryo-EM structures, biochemical characterization of the HIM motif provides insight into the cross-reactivity and selectivity of the Stachel sequences. Two interconnected mechanisms, the sensing of Stachel sequences by the conserved ‘toggle switch’ W6.53 and the constitution of a hydrogen-bond network formed by Q7.49/Y7.49 and the P6.47/V6.47φφG6.50 motif (φ indicates a hydrophobic residue), are important in Stachel sequence-mediated receptor activation and Gs coupling. Notably, this network stabilizes kink formation in TM helices 6 and 7 (TM6 and TM7, respectively). A common Gs-binding interface is observed between the two aGPCRs, and GPR114 has an extended TM7 that forms unique interactions with Gs. Our structures reveal the detailed mechanisms of aGPCR activation by Stachel sequences and their Gs coupling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The cryo-EM density map for the GPR133-CTF–Gs complex has been deposited in the Electron Microscopy Data Bank under accession code EMD-31232. The coordinates for the model of GPR133-CTF–Gs have been deposited in the PDB under accession number 7EPT. The cryo-EM density map for the GPR144-CTF–Gs complex has been deposited in the Electron Microscopy Data Bank under accession code EMD-31254. The coordinates for the model of GPR144-CTF–Gs have been deposited in the PDB under accession number 7EQ1. All other data are available from the corresponding authors.
Change history
20 April 2022
In the version of this article initially published, there was an error in the Peer review statement, which has now been amended.
References
Bassilana, F., Nash, M. & Ludwig, M. G. Adhesion G protein-coupled receptors: opportunities for drug discovery. Nat. Rev. Drug Discov. 18, 869–884 (2019).
Bondarev, A. D. et al. Opportunities and challenges for drug discovery in modulating adhesion G protein-coupled receptor (GPCR) functions. Expert Opin. Drug Discov. 15, 1291–1307 (2020).
Hamann, J. et al. International Union of Basic and Clinical Pharmacology. XCIV. Adhesion G protein-coupled receptors. Pharmacol. Rev. 67, 338–367 (2015).
Hochreiter-Hufford, A. E. et al. Phosphatidylserine receptor BAI1 and apoptotic cells as new promoters of myoblast fusion. Nature 497, 263–267 (2013).
Eubelen, M. et al. A molecular mechanism for Wnt ligand-specific signaling. Science 361, eaat1178 (2018).
Zhang, D. L. et al. Gq activity- and β-arrestin-1 scaffolding-mediated ADGRG2/CFTR coupling are required for male fertility. eLife 7, e33432 (2018).
Liebscher, I. et al. A tethered agonist within the ectodomain activates the adhesion G protein-coupled receptors GPR126 and GPR133. Cell Rep. 9, 2018–2026 (2014).
Stoveken, H. M., Hajduczok, A. G., Xu, L. & Tall, G. G. Adhesion G protein-coupled receptors are activated by exposure of a cryptic tethered agonist. Proc. Natl Acad. Sci. USA 112, 6194–6199 (2015).
Petersen, S. C. et al. The adhesion GPCR GPR126 has distinct, domain-dependent functions in Schwann cell development mediated by interaction with laminin-211. Neuron 85, 755–769 (2015).
Wilde, C. et al. The constitutive activity of the adhesion GPCR GPR114/ADGRG5 is mediated by its tethered agonist. FASEB J. 30, 666–673 (2016).
Scholz, N. et al. Mechano-dependent signaling by Latrophilin/CIRL quenches cAMP in proprioceptive neurons. eLife 6, e28360 (2017).
Yeung, J. et al. GPR56/ADGRG1 is a platelet collagen-responsive GPCR and hemostatic sensor of shear force. Proc. Natl Acad. Sci. USA 117, 28275–28286 (2020).
Arac, D. et al. A novel evolutionarily conserved domain of cell-adhesion GPCRs mediates autoproteolysis. EMBO J. 31, 1364–1378 (2012).
Krasnoperov, V. et al. Post-translational proteolytic processing of the calcium-independent receptor of α-latrotoxin (CIRL), a natural chimera of the cell adhesion protein and the G protein-coupled receptor. Role of the G protein-coupled receptor proteolysis site (GPS) motif. J. Biol. Chem. 277, 46518–46526 (2002).
Lin, H. H. et al. Autocatalytic cleavage of the EMR2 receptor occurs at a conserved G protein-coupled receptor proteolytic site motif. J. Biol. Chem. 279, 31823–31832 (2004).
Langenhan, T., Aust, G. & Hamann, J. Sticky signaling-adhesion class G protein-coupled receptors take the stage. Sci. Signal. 6, re3 (2013).
Stoveken, H. M., Larsen, S. D., Smrcka, A. V. & Tall, G. G. Gedunin- and khivorin-derivatives are small-molecule partial agonists for adhesion G protein-coupled receptors GPR56/ADGRG1 and GPR114/ADGRG5. Mol. Pharmacol. 93, 477–488 (2018).
Bradley, E. C. et al. In vivo identification of small molecules mediating Gpr126/Adgrg6 signaling during Schwann cell development. Ann. NY Acad. Sci. 1456, 44–63 (2019).
Ping, Y. Q. et al. Structures of the glucocorticoid-bound adhesion receptor GPR97–Go complex. Nature 589, 620–626 (2021).
Diamantopoulou, E. et al. Identification of compounds that rescue otic and myelination defects in the zebrafish adgrg6 (gpr126) mutant. eLife 8, e44889 (2019).
Gupte, J. et al. Signaling property study of adhesion G-protein-coupled receptors. FEBS Lett. 586, 1214–1219 (2012).
Liebscher, I. & Schoneberg, T. Tethered agonism: a common activation mechanism of adhesion GPCRs. Handb. Exp. Pharmacol. 234, 111–125 (2016).
Beliu, G. et al. Tethered agonist exposure in intact adhesion/class B2 GPCRs through intrinsic structural flexibility of the GAIN domain. Mol. Cell 81, 905–921 (2021).
Wootten, D., Simms, J., Miller, L. J., Christopoulos, A. & Sexton, P. M. Polar transmembrane interactions drive formation of ligand-specific and signal pathway-biased family B G protein-coupled receptor conformations. Proc. Natl Acad. Sci. USA 110, 5211–5216 (2013).
Duan, J. et al. Structures of full-length glycoprotein hormone receptor signalling complexes. Nature 598, 688–692 (2021).
Demberg, L. M. et al. Activation of adhesion G protein-coupled receptors: agonist specificity of Stachel sequence-derived peptides. J. Biol. Chem. 292, 4383–4394 (2017).
Promel, S. et al. Characterization and functional study of a cluster of four highly conserved orphan adhesion-GPCR in mouse. Dev. Dynam. 241, 1591–1602 (2012).
Hilger, D. et al. Structural insights into differences in G protein activation by family A and family B GPCRs. Science 369, eaba3373 (2020).
Isberg, V. et al. Generic GPCR residue numbers—aligning topology maps while minding the gaps. Trends Pharmacol. Sci. 36, 22–31 (2015).
Zhou, F. et al. Molecular basis of ligand recognition and activation of human V2 vasopressin receptor. Cell Res. 31, 929–931 (2021).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Melero, R. et al. Continuous flexibility analysis of SARS-CoV-2 spike prefusion structures. IUCrJ https://doi.org/10.1107/S2052252520012725 (2020).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Yang, F. et al. Structural basis of GPBAR activation and bile acid recognition. Nature 587, 499–504 (2020).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Bianco, G., Forli, S., Goodsell, D. S. & Olson, A. J. Covalent docking using autodock: two-point attractor and flexible side chain methods. Protein Sci. 25, 295–301 (2016).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/openMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2016).
Lomize, M. A., Pogozheva, I. D., Joo, H., Mosberg, H. I. & Lomize, A. L. OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res. 40, D370–D376 (2012).
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graphics 14, 33–38 (1996).
Schoneberg, T., Liu, J. & Wess, J. Plasma membrane localization and functional rescue of truncated forms of a G protein-coupled receptor. J. Biol. Chem. 270, 18000–18006 (1995).
Strachan, R. T. et al. Divergent transducer-specific molecular efficacies generate biased agonism at a G protein-coupled receptor (GPCR). J. Biol. Chem. 289, 14211–14224 (2014).
Olsen, R. H. J. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
Yang, F. et al. Structure, function and pharmacology of human itch receptor complexes. Nature 600, 164–169 (2021).
Acknowledgements
We thank all staff members at the Cryo-Electron Microscopy Research Center, Shanghai Institute of Material Medica, for help with cryo-EM data collection for the GPR133-CTF–Gs complex and those at Shuimu BioSciences, Ltd, and the cryo-EM centre of the Southern University of Science and Technology for help with data collection for the GPR114-CTF–Gs–scFv16 complex. We also thank Y. Yu of the Translational Medicine Core Facility of Advanced Medical Research Institute, Shandong University, for technical support with the Envision multimode plate reader. The scientific calculations in this study were performed using the HPC Cloud Platform at Shandong University. This work was supported by the National Key Research and Development Program of China (2018YFC1003600 to J.-P.S., 2019YFA0904200 to J.-P.S. and P.X. and 2018YFA0507002 to H.E.X.), the National Science Fund for Distinguished Young Scholars Grant (81825022 to J.-P.S.), the National Science Fund for Excellent Young Scholars (82122070 to F.Y.), the Shandong Provincial Natural Science Fund for Excellent Young Scholars (ZR2021YQ18 to P.X.), the National Natural Science Foundation of China Grant (81773704 to J.-P.S., 31971195 to P.X., 31900936 to F.Y. and 31770796 to Y.J.), the Key Research Project of the Natural Science Foundation of Beijing, China (Z200019 to J.-P.S.), the Shanghai Municipal Science and Technology Major Project (2019SHZDZX02 to H.E.X.), the CAS Strategic Priority Research Program (XDB37030103 to H.E.X.), the Key Research and Development Program of Shandong Province (2021CXGC011105 to J.-P.S., GG201709260059 to P.X. and 2021ZLGX02 to J.-P.S. and L.D.), the Shandong Provincial Natural Science Foundation Grant (ZR2020ZD39 to J.-P.S. and ZR2016CQ07 to P.X.), the German Research Foundation CRC1052 and CRC1423 (project numbers 209933838 and 421152132 to I.L. and T.S.), the COST Association (COST Action CA18240 Adher ’n Rise to I.L. and T.S.) and Fundamental Research Funds for the Central Universities (2021JCG020 to J.-P.S. and P.X.).
Author information
Authors and Affiliations
Contributions
J.-P.S. and H.E.X. conceived, designed and supervised the overall project and supervised and guided all structural analyses. J.-P.S. initiated the study of structural understanding of the mechanism underlying the tethered activation and force induced activation of aGPCRs. J.-P.S., H.E.X., I.L., Y.-Q.P., P.X. and F.Y. participated in data analysis and interpretation. S.-C.G. and P.X. generated the GPR133-CTF insect cell expression constructs, established the purification procedure for the GPR133-CTF–Gs–Nb35 complex and prepared samples for cryo-EM. F.Y., X.Y. and X.W. generated the GPR114-CTF insect cell expression constructs, established the purification procedure for the GPR114-CTF–Gs–scFv16 complex and prepared samples for cryo-EM. W.Y. designed the modified Gαs construct. Y.-Q.P. and F. Zhao prepared the cryo-EM grids. Y.-Q.P. and P.X. collected the cryo-EM data and performed cryo-EM map calculation, model building and refinement with the assistance of Z.-L.Z. F. Zhou and D.-F.H. participated in the cryo-EM data analysis. C.Z. and Y.L. performed MD simulations. J.-P.S. designed, and I.L. and R.-J.Z. performed mechano-force assays with the assistance of F.D.H. I.L., T.S., F.-Z.L. and R.-J.Z. performed N-terminally elongated assays. R.-J.Z., X.Y., Y.-T.X. and D.-L.Z. generated all of the GPR133-CTF and GPR114-CTF constructs for the cell-based assays. I.L., L.D., S.Q.F. and T.S. participated in the design and explanation of the mechano-force assay results and the activation of GPR133 and GPR114 full-length Stachel sequences and induced cross-reactivity and cross-talk between the GAIN domain and TM7. I.L. and T.S. provided insightful ideas and experimental designs for many other experiments. R.-J.Z., F.Y., P.X., X.Y., Y.-T.X. and I.L. performed all cellular functional assays for GPR133-CTF and GPR114-CTF. Y.-Q.P., P.X., S.-C.G. and R.-J.Z. prepared the figures with the assistance of Y.J. and Y.L. J.-P.S. and H.E.X. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Antony Boucard, Aashish Manglik and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 GPR133 and GPR114 basal activity and complex purification.
a-b, Intracellular cAMP levels measured by GloSensor assay showing the basal activities of different constructs of GPR133 (a) and GPR114 (b) at similar cell surface expression levels (c) and (d), respectively. The pcDNA3.1 vector was used as a control. All data represent the mean ± SEM from three independent experiments (n=3) performed in triplicate. ***, P<0.001; **, P<0.01; *, P<0.05. Comparisons were performed between GPR133-FL (or GPR114-FL) and its derivative constructs at respective expression levels. The mutant receptor expression was normalized to comparable levels with WT receptor expression in our ELISA assay by adjusting the amount of plasmid transfected. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.9607, <0.0001, =0.0012, =0.0001 from left to right for GPR133-FL curve; P=0.9607, <0.0001, <0.0001, <0.0001 from left to right for GPR133-CTF-ΔGPS curve; P>0.9999, =0.0024, =0.0045, <0.0001 from left to right for GPR114-FL curve; P=0.1207, =0.0003, =0.0005, <0.0001 from left to right for GPR114-CTF-ΔGPS curve). c, ELISA data to determine the cell surface expression levels of the wild-type (WT) GPR133-FL, GPR133-CTF and GPR133-CTF-ΔGPS (GPR133-CTF with truncated Stachel sequence) in HEK293 cells. The x-axis shows the amount of transfected plasmid DNA. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of the GPR133-FL at the lowest amount of transfected DNA (indicated as 1). Data is shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistical significance of differences between WT and mutants was determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.8727, 0.0609, 0.6418, 0.6170, 0.6977, 0.8011, 0.6404, 0.5124 from left to right). d, ELISA data to determine the cell surface expression levels of the WT GPR114-FL, GPR114-CTF and GPR114-CTF-ΔGPS (GPR114-CTF with truncated Stachel sequence) in HEK293 cells. The x-axis shows the amount of transfected plasmid DNA. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of the GPR114-FL at the lowest amount of transfected DNA (indicated as 1). Data is shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistically significant differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P>0.9999, >0.9999, =0.1483, >0.9999, =0.7754, >0.9999, =0.3840, >0.9999 from left to right). e, Concentration-response curves of GPR133-CTF-ΔGPS (GPR133-CTF with truncated Stachel sequence) under the incubation with the Stachel peptide (p133). Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. f, Concentration-response curves of GPR114-CTF-ΔGPS (GPR114-CTF with truncated Stachel sequence) under the incubation with the Stachel peptide (p114). Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. g-j, To elucidate the molecular basis for the Stachel sequence-dependent activation of GPR133 and GPR114, we set out to determine the cryo-EM structures of GPR133-CTF-Gs and GPR114-CTF-Gs complexes. Baculoviruses encoding human Gαs, Gβ1, and Gγ2 were co-expressed with GPR133-CTF or GPR114-CTF in Spodoptera frugiperda (Sf9) insect cells to enable complex formation. The nanobody 35 (Nb35) was added for stabilization during further purification. The GPR114-CTF-Gs-trimer complex was unstable and often underwent dissociation. We therefore used a modified Gαs construct, where its N terminus was replaced with the corresponding Gαi residues, and the antibody fragment scFv16 to improve the percentage of intact particles in the cryo-EM study. (g-h), Molecular sieve peak diagram and Coomassie brilliant blue-stained gel image of the GPR114-CTF-Gs-trimer. (i), Molecular sieve peak diagram and Coomassie brilliant blue-stained gel image of the GPR133-CTF-Gs-Nb35 complex. (j), Molecular sieve peak diagram and Coomassie brilliant blue-stained gel image of the GPR114-CTF-Gs-scFv16 complex. Representative Figures from three independent experiments were shown. k-l, The final model of GPR133-CTF encompasses all residues from H543 to T827, with the exception of partial third intracellular loop (ICL3) residues (743–755). The GPR114-CTF model includes residues from T227 to A520, lacking the ICL3 residues between E443 and H454. (k), Orthogonal views of the model of GPR133-CTF-Gs-Nb35 complex (scale bar: 10 Å). GPR133-CTF, dark cyan; Gαs, gold; Gβ, light blue; Gγ, hot pink; Nb35, grey. (l), Orthogonal views of the model of GPR114-CTF-Gs-scFv16 complex (scale bar: 10 Å). GPR114-CTF, blue; Gαs, yellow; Gβ, light blue; Gγ, hot pink; scFv16, thistle.
Extended Data Fig. 2 Cryo-EM data processing, and overall resolution analysis of electron density of transmembrane helices, α5-helix of Gαs.
a-b, Cryo-EM micrographs of GPR133-CTF-Gs-Nb35 (a) and GPR114-CTF-Gs-scFv16 (b) complexes (scale bar: 30 nm) and 2D class averages (scale bar: 5 nm). 4,882 movies of GPR133-CTF-Gs complex and 2,777 movies of GPR114-CTF-Gs complex were collected using Titan Krios equipped with a K3 Summit direct electron detector. c-d, Flow chart of cryo-EM data processing of GPR133-CTF-Gs-Nb35 (c) and GPR114-CTF-Gs-scFv16 (d) complexes. EM maps with global resolutions of 3.1 Å and 3.3 Å were finally acquired for the GPR133-CTF-Gs-Nb35 and GPR114-CTF-Gs-scFv16 complexes from 575,193 and 299,210 particles, respectively. e-f, 3D density map of GPR133-CTF-Gs-Nb35 (e) and GPR114-CTF-Gs-scFv16 (f). Complexes colored according to local resolution (Å). Fourier shell correlation (FSC) curves of the final reconstruction and the refined model versus the final map. g-h, Cryo-EM density of transmembrane helices of GPR133-CTF, GPR114-CTF, α5-helix of Gαs, in the determined structures of GPR133-CTF-Gs-Nb35 (g) and GPR114-CTF-Gs-scFv16 (h). Well-defined EM densities corresponding to most regions of the TMD and the heterotrimeric Gs could be traced in both the GPR133-CTF-Gs-Nb35 and GPR114-CTF-Gs-scFv16 complexes.
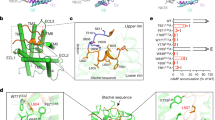
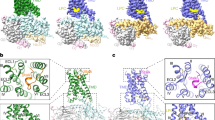
Extended Data Fig. 3 Overall characteristics of GPR133-CTF/Gs and GPR114-CTF/Gs TMD.
a-b, Structural comparison of TMD of Gs-bound GPR133-CTF (7EPT, teal) (a) and Gs-bound GPR114-CTF (7EQ1, marine) (b) with Gs-bound GLP1R (6B3J, green), Gs-bound β2AR (3SN6, light orange), active GABAB2 (7C7Q, olive) GB2 subunit, and Gi-bound Smoothened receptor (6OT0, gray) shown in orthogonal views. c, The detailed presentation of the Stachel sequences in GPR133-CTF (left panel). The Stachel sequence of the receptors turned approximately 80° toward the TM5, and then flipped back to TM1. The Stachel sequence of GPR133-CTF is highlighted (salmon). Cryo-EM density of the Stachel sequence in GPR133-CTF (right panel). d, The detailed presentation of the Stachel sequences in GPR114-CTF (left panel). The Stachel sequence of the receptors turned approximately 80° toward the TM5, and then flipped back to TM1. The Stachel sequence is highlighted (yellow) of GPR114-CTF. Cryo-EM density of the Stachel sequence in GPR114-CTF (right panel). e, The detailed presentation and the cryo-EM density of the Stachel sequences and ECL2 in GPR133-CTF (teal) or GPR114-CTF (marine) shown in the extracellular views. The ECL2 contacts with the Stachel sequences inserted into the TM bundle and makes extensive hydrophobic interactions in both receptors. The Stachel sequences of GPR133-CTF (salmon) and GPR114-CTF (yellow) are highlighted. f, Close-up view and the cryo-EM density of the conserved disulfide bonds between C6323.29 and C704ECL2 in GPR133 and C3143.29 and C404ECL2 in GPR114. g, The cytoplasmic view of the outward shifting of TM6 and TM7 in GPR133-CTF-Gs and GPR114-CTF-Gs complex. h, The unique extended TM7 of GPR114-CTF compared with the primary sequence equivalent H8 of GPR133-CTF. The extended TM7 of GPR114-CTF inserts into the crevice formed between Gαs and Gβ.
Extended Data Fig. 4 Intramolecular interactions of the Stachel sequence.
a-b, The overall structures of GPR133-CTF (a) and GPR114-CTF (b) and the binding poses of their Stachel sequences. The Stachel sequences are shown in salmon in GPR133-CTF and yellow in GPR114-CTF. c-h, Detailed interactions of the HIM within the binding pocket of GPR114-CTF-Gs-scFv16 complex. i, Cryo-EM density of the HIM in Stachel sequence and its interacting residues in GPR133. j, Cryo-EM density of the HIM within the Stachel sequence and its interacting residues in GPR114. k, Residues in the Stachel sequence binding pocket of the GPR133-CTF-Gs-Nb35 and GPR114-CTF-Gs-scFv16 complexes in contact with their HIMs. The HIMs of GPR133-CTF and GPR114-CTF are in red and yellow, respectively. Corresponding residues are indicated by red circles. The different interaction residues in the two binding pockets are indicated by green (GPR133-CTF) and blue (GPR114-CTF) circles.
Extended Data Fig. 5 Detailed interactions of the Stachel sequence.
a-b, The detailed interactions of the last residue in the HIM with the hydrophobic pocket created by ECL2, TM5 and TM6 in GPR133 (a) and GPR114 (b). c, The Stachel sequence alignment of aGPCRs. The last residue in the HIM is highlighted in yellow. d, Close-up view of the interactions of the β strand in the Stachel sequence with ECL3 and its cryo-EM density in the GPR133-CTF-Gs complex. The upper β strand formed substantial interactions with ECL3, including the hydrophobic packing of V554 and L558 in the β-strand contacts with V780ECL3 and V785ECL3, respectively, as well as the hydrogen bonds anchored between Y789 and H562. Hydrogen bonds are highlighted by blue dashed lines. e, Close-up view of the interactions of the β strand in the Stachel sequence with ECL2 in GPR114-CTF-Gs complex. f, ELISA data to determine the cell surface expression levels of WT GPR133-CTF or mutants (Figure subtitles, MUT indicated mutant) of the Stachel sequence in HEK293 cells. The x-axis shows the amount of transfected plasmid DNA. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of the WT GPR133-CTF at the lowest amount of DNA (indicated as 1). The mutants show no significant difference compared to the WT. Data is given as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistical significance of differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.2673, 0.0519, 0.7478, 0.7386, 0.7767, 0.9636, 0.3029, 0.3934, 0.4981, 0.5609, 0.0691, 0.6844, 0.9734, 0.8165, 0.2466, 0.2920, 0.5868, 0.9218, 0.0621, 0.2494, 0.1486, 0.7737, 0.7689, 0.5235, 0.9452, 0.4328, 0.7852, 0.4602 from left to right). g, Expression-response curves of the receptor mutants at the Stachel sequence of GPR133-CTF in cAMP accumulation assay. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate for the WT and mutants. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.9860, =0.0069, =0.0003, <0.0001 from left to right for T545A curve; P=0.9031, =0.3850, =0.0021, <0.0001 from left to right for N546A curve; P=0.2874, <0.0001, =0.0001, <0.0001 from left to right for F547A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for I549A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for L550A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for M551A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for V553A curve). h, Effects of mutations in the Stachel sequences of GPR114-CTF on their basal activities. The bar graphs were generated based on the representative differences between WT GPR114-CTF and their mutants (MUT), at a relative expression level of 1.5 for GPR114-CTF shown in Extended Data Fig. 5i. Data are normalized and presented as the response percentage for WT GPR114-CTF, and shown as the mean ± SEM of three experiments (n=3) performed in triplicate. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.3065, 0.0003, 0.0008, 0.0041, 0.0001, 0.0013, 0.0012 from top to bottom). i, ELISA data to determine the cell surface expression levels of WT GPR114-CTF or mutants (Figure subtitles, MUT indicated mutant) of the Stachel sequence in HEK293 cells. The x-axis shows the amount of transfected plasmid DNA. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of the WT GPR114-CTF at the lowest amount of DNA (indicated as 1). The mutants show no significant difference compared to the WT. Data is given as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistical significance of differences between WT and mutants was determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.9176, 0.1835, >0.9999, >0.9999, >0.9999, 0.4778, >0.9999, 0.9951, 0.9027, 0.9285, 0.9712, 0.5330, 0.5783, 0.5036, 0.9166, 0.3091, 0.9249, 0.8114, 0.3435, 0.6027, 0.1607, 0.7014, 0.8360, 0.6672, 0.4233, 0.9954, 0.6394, 0.6221 from left to right). j, Expression-response curves of the receptor mutants at the Stachel sequence of GPR114-CTF in cAMP accumulation assay. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate for the WT and mutants. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P= 0.1012, =0.0280, =0.3065, =0.0042 from left to right for T227I curve; P= 0.0688, =0.0012, =0.0003, =0.0002 from left to right for Y228A curve; P= 0.0011, =0.0011, =0.0008, =0.0002 from left to right for F229A curve; P= 0.0026, =0.0027, =0.0041, =0.0002 from left to right for V231A curve; P= 0.0668, =0.0006, =0.0001, =0.0002 from left to right for L232A curve; P= 0.0114, =0.0050, =0.0013, =0.0004 from left to right for M233A curve; P= 0.0023, =0.0047, =0.0012, =0.0002 from left to right for L235A curve).
Extended Data Fig. 6 Related expression and signaling data of GPR133 and GPR114 receptor and peptide mutants.
a, ELISA data to determine the cell surface expression levels of WT GPR133-CTF or mutants of the binding pocket in HEK293 cells. The x-axis shows the amount of transfected DNA. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of the WT GPR133-CTF at the lowest amount of DNA (indicated as 1). The mutants show no significant difference from the WT. Data is given as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistical significance of differences between WT and mutants was determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.4727, 0.1565, 0.7767, 0.3254, 0.6891, 0.0871, 0.4670, 0.4542, 0.4230, 0.9260, 0.6733, 0.5506, 0.6659, 0.9314, 0.7897, 0.4868, 0.5883, 0.3622, 0.7033, 0.9226, 0.8527, 0.6059, 0.3041, 0.7325, 0.8596, 0.7842, 0.8259, 0.9677, 0.9067, 0.7272, 0.0752, 0.8870, 0.7973, 0.8011, 0.9420, 0.9441 from left to right). b, Expression-response curves of the mutants at the binding pocket of GPR133-CTF in cAMP accumulation assay. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate for the WT and mutants. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for W705A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for I713A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for W714A curve; P<0.0001, =0.1291, =0.0047, <0.0001 from left to right for F716A curve; P<0.0001, =0.0415, =0.0016, <0.0001 from left to right for V717A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for A720G curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for W773A curve; P<0.0001, =0.0048, =0.0066, <0.0001 from left to right for V777A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for F791A curve.). c, Effects of mutations in the binding pocket of GPR114-CTF on their basal activity. The bar graphs were generated based on the representative differences between WT GPR114-CTF and their mutants (MUT), at a relative expression level of 1.5 for GPR114-CTF shown in (d). Data are normalized and presented as the response percentage for WT GPR114-CTF and shown as the mean ± SEM of three experiments (n=3) performed in triplicate. All data were analysed by two-sided one-way ANOVA with Tukey’s test. ***, P<0.001; **, P<0.01 (P<0.0001, <0.0001, =0.0007, =0.0019, =0.0007, <0.0001, =0.0002, =0.0008, =0.0007 from top to bottom). d, ELISA data to determine the cell surface expression levels of WT GPR114-CTF or mutants of the binding pocket in HEK293 cells. The x-axis shows the amount of transfected DNA. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of the WT GPR114-CTF at the lowest amount of DNA (indicated as 1). The mutants show no significant difference from the WT. Data is given as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistical significance of differences between WT and mutants was determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.4235, 0.4226, 0.4226, 0.3091, 0.3223, 0.4095, 0.4778, 0.6505, 0.1903, 0.4319, 0.3254, 0.3702, 0.3194, 0.4279, 0.2488, 0.2451, 0.2869, 0.3321, 0.2632, 0.6253, 0.0625, 0.9249, 0.0864, 0.4283, 0.9007, 0.9249, 0.8107, 0.3834, 0.0864, 0.3197, 0.1158, 0.3410, 0.2095, 0.2236, 0.5310, 0.2913 from left to right). e, Expression-response curves of the mutants at the binding pocket of GPR114-CTF in cAMP accumulation assay. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate for the WT and mutants. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P= 0.0647, =0.0011, =0.0004, =0.0001 from left to right for C256A curve; P= 0.1161, <0.0001, =0.0003, =0.0001 from left to right for F294A curve; P= 0.1347, <0.0001, =0.0007, =0.0004 from left to right for F298A curve; P= 0.1890, =0.0451, =0.0019, =0.0005 from left to right for P302A curve; P= 0.0254, =0.0871, =0.0007, =0.0004 from left to right for L321A curve; P= 0.3739, <0.0001, =0.0003, =0.0001 from left to right for L325A curve; P= 0.6087, =0.0011, =0.0002, =0.0003 from left to right for M417A curve; P= 0.3465, <0.0001, =0.0005, =0.0002 from left to right for W470A curve; P= 0.0013, =0.0003, =0.0007, =0.0002 from left to right for F488A curve). f, Effects of mutations in the synthetic GPR133 Stachel peptide (p133) tested in cAMP accumulation assays with HEK293 cells expressing GPR133-CTF-ΔGPS. Data are normalized to the WT Stachel peptide and presented as the mean ± SEM of three experiments performed in triplicate. Statistical differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey’s test. ***, P<0.001; **, P<0.01 (P<0.0001, <0.0001). g, Concentration-response curves of GPR133-CTF-ΔGPS (GPR133-CTF with truncated Stachel sequence) in cAMP accumulation assay under the incubation with p133 and its mutants at Stachel sequence. Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. h, Effects of mutations in the synthetic GPR114 Stachel peptide (p114) tested in cAMP accumulation assays with HEK293 cells expressing GPR114-CTF-ΔGPS. Data shown in (i) are normalized to the WT Stachel peptide and presented as the mean ± SEM of three experiments performed in triplicate. Statistically significant differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey’s test. ***, P<0.001 (P=0.0001). i, Concentration-response curves of GPR114-CTF-ΔGPS (GPR114-CTF with truncated Stachel sequence) in cAMP accumulation assay under the incubation with p114 and its Stachel sequence mutants. Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate.
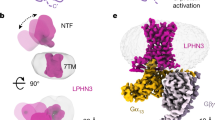
Extended Data Fig. 7 Stachel sequence-mediated activation of Gs-coupled aGPCRs.
a, Effects of increasing amounts of transfected receptor plasmids on the cleavage-deficient GPR133-FL-AA activation in responses to mechanical force stimulations. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate for the different transfection quantity of WT. All data were analysed by two-sided one-way ANOVA with Tukey’s test. ***, P<0.001; **, P<0.01; ns, no significant difference (P=0.5974, =0.6685, =0.0047, =0.0004, <0.0001 from left to right). b, COS-7 cells were transiently transfected with 500 ng of empty vector and mGPR133-FL as well as the cleavage-deficient mutant mGPR133-FL-H540R. Accumulation of cAMP was measured after shaking the cells at 100 rpm. Empty vector served as negative control (cAMP level: 3.01 ± 1.81 nM/well) (effect of construct p=0.0008, effect of shake p=0.0475, interaction construct × shake p=0.126; two-way ANOVA). Data are given as means ± SEM of three independent experiments each performed in triplicate. Statistics were performed by applying a two-way ANOVA followed by Sidak’s post hoc analysis; *, P<0.05; **, P<0.01; ***, P<0.001. All significance given above individual points in the graphs show the result of the post hoc analysis (P=0.0398, 0.0375). c, COS-7 cells were transiently transfected with 500 ng of empty vector and hGPR133-FL or the cleavage-deficient mutant hGPR133-FL-H540R. Accumulation of cAMP was measured after shaking the cells at 100 rpm. Empty vector served as negative control (cAMP level: 1.30 ± 0.37 nM/well) (effect of construct p=0.0054, effect of shake p=0.0122, interaction construct × shake p=0.0528; two-way ANOVA). Data are given as means ± SEM of three independent experiments each performed in triplicate. Statistics were performed by applying a two-way ANOVA followed by Sidak’s post hoc analysis; *, P<0.05; **, P<0.01; ***, P<0.001. All significance given above individual points in the graphs show the result of the post hoc analysis (P=0.0216, 0.0198). d, COS-7 cells were transfected with 500 ng/well of tagged constructs. Specific optical density (OD) readings (OD value of double HA/Flag-tagged aGPCR constructs minus OD value of vector-transfected cells) are given as percentage of the human P2Y12 receptor, which served as positive control. The non-specific OD value (pcDps=empty vector) was 0.05 ± 0.02 (set 0%) and the OD value of P2Y12 was 1.30 ± 0.16 (set 100%). Data are given as means ± SD of three independent experiments each performed in triplicates (P=0.9932, 0.9874, 0.5049, 0.9487 from left to right). e, ELISA of the binding pocket mutants to determine the cell surface expression levels of the GPR133-FL-AA and their corresponding mutants in HEK293 cells. The relative expression level of mutants was compared to GPR133-FL-AA (indicated as 1). The mutants show no significant difference from the WT. They are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistically significant differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.9998, 0.9946, 0.9999, 0.9997, 0.9997, 0.9996, 0.9994). f, Sequence alignment of the HIM among several aGPCR members. The lower case added in front of receptor represents different species. h, human; m, mouse; b, bovine. g, Detailed and key interactions between the Stachel sequence and the GAIN domain of GPR133-FL structure which was download from AlphaFold Protein Structure Database. h, Expression-response curves for mutants of the GAIN domain of GPR133-FL-AA in cAMP accumulation assay (Figure subtitles, MUT indicated mutant). Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate for the WT and mutants. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.6735, =0.0548, <0.0001, =0.0111 from left to right for T459A curve; P=0.7744. <0.0001, <0.0001, =0.0442 from left to right for L487S curve; P=0.7744, <0.0001, <0.0001, =0.0001 from left to right for V508A-Y509A curve; P=0.2291, <0.0001, <0.0001, =0.0201 from left to right for W522A curve). i, ELISA data to determine the cell surface expression levels of the WT GPR133-FL-AA or mutants of the GAIN domain in HEK293 cells. The x-axis shows the amount of plasmid. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of GPR133-FL-AA at the lowest transfected amount of DNA (indicated as 1). The mutants show no significant difference from the WT. Data are given as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistical significance of differences between WT and mutants was determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.3118, 0.3376, 0.9500, 0.6782, >0.9999, 0.7715, 0.5450, 0.4617, 0.9028, 0.4219, 0.9528, 0.9028, 0.9900, 0.8012, >0.9999, 0.9973 from left to right). j, Concentration-response curves of GPR114-CTF-ΔGPS (abbreviated as GPR114-C-Δ) and GPR133-CTF-ΔGPS (abbreviated as GPR133-C-Δ) in response to the stimulation with the GPR114 Stachel peptide (p114). Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. k, The histogram shows the relative cAMP levels in HEK293 cells overexpressing FL GPR114 constructs after stimulation with different elongated peptides. The degree of GPR114 activation is represented by cAMP accumulation (x-fold over empty vector). Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistically significant differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey test. ***, P<0.001; **, P<0.01 (P=0.9391, 0.8837, 0.9860, 0.8923, 0.9049, 0.0298 from top to bottom).
Extended Data Fig. 8 The selectivity of Stachel sequences and structural motifs in the active state of GPR133-CTF and GPR114-CTF.
a-b, RMSD of the N-terminally elongated Stachel peptides (upper panel) and RMSD of binding pocket residues which directly interact with the N-terminally elongated Stachel peptide (lower panel) of GPR133 (a), GPR114 (b) during 200 ns MD simulation. The shaded area represents the original value of RMSD, which was reduced the transparency, and the bold line is the result of averaging every 200 points of the original value of RMSD. c, GPR114-CTF N-terminal Stachel sequence elongation did not reduce its activation because H225 and A2421.33 in GPR114 do not have such negative effect compared to GPR133-CTF N-terminal elongation. The model of Stachel sequence-elongated GPR114-CTF was generated by computational simulation. d-e, Expression-response curves (d) and bar graphs (e) for GPR114-CTF and GPR114-CTF-HL mutants (GPR114-CTF N-terminal elongation with Hss-−2 and Lss-−1) in cAMP accumulation assay. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate. ns, no significance. Comparison between GPR114-CTF and GPR114-CTF-HL at respective transfection amount of plasmid. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.6843, 0.6943, 0.4025, 0.5021 from left to right for GPR114-CTF-HL curve). f, ELISA data to determine the cell surface expression levels of GPR114-CTF and GPR114-CTF-HL in HEK293 cells. Relative expression levels compared to WT GPR114-CTF (set as 1) are shown. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate. ns, no significant difference. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.2408). g-h, Structural representation of mutation effects on the extended N terminus of the GPR133 Stachel sequence. (g), The mutation of H543D enabled favorable charge attraction with R5601.33. (h), The mutation of L544N enabled potential hydrogen bond formation between N544 and S5671.40. The H-bond is described as a dashed line. i, Bar graph of mutations in the GPR133-CTF-HL and its mutants (H543D, L544N) and their impact on basal activity. The y-axis represents the corresponding RLU value when the relative expression amount is 6 (as shown in the x-axis of Fig. 3e). Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistically significant differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey test. ***, P<0.001; *, P<0.05 (P=0.0003, 0.0360 from left to right). j, ELISA data to determine the cell surface expression levels of GPR133-CTF-HL, GPR133-CTF-HL(H543D) and GPR133-CTF-HL(L544N) in HEK293 cells. Relative expression levels compared to GPR133-CTF-HL (set as 1) are shown. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate. ns, no significant difference. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.9025, 0.1695 from left to right). k-p, We chose two other experimentally verified uncleavable aGPCRs to investigate for potential Stachel elongation. Compared to GPR111 full length (GPR111-FL), GPR111-CTF and GPR111-CTF-LF had stronger effects on inhibiting intracellular cAMP levels in response to Fsk stimulation, while GPR111-CTF-ΔGPS shows no activity (k, l). Even though all constructs show similar expression levels (m). As the mouse orthologue of GPR115 has been shown to induce IP formation in the cell, we chose Gq dissociation BRET assay to investigate the activity of its CTF mutant. In contrast to mouse GPR115, the constitutive Gq activity of human GPR115-CTF as well as the elongated human GPR115-CTF-VV is stronger than its full length (GPR115-FL) or the Stachel sequence deletion mutant (GPR115-CTF-ΔGPS) (n-o) at comparable expression levels (p). These biochemical characterizations indicated that the Stachel sequences of GPR111 and GPR115 contribute to the constitutive activities of their CTFs. The results also showed that the Stachel of GPR111 and GPR115 can be N-terminally extended without significant alteration of the activities. According to our computational simulations, the positively charged residues H543 and R5601.33 in GPR133-CTF can be replaced by the corresponding hydrophobic residues L428 and I4451.38 in GPR111, or V383 and V4001.38 in GPR115, thus allowing favorable N-terminal extension at these positions (shown in Supplementary Fig. 5). (k-l), Expression-response curves and bar graphs for GPR111-CTF, GPR111-CTF-LF (GPR111-CTF N-terminal elongation with Lss-−2 and Fss-−1), GPR111-FL, and GPR111-CTF-ΔGPS in cAMP accumulation assay. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate. ***, P<0.001; **, P<0.01; *, P<0.05; ns, no significance. Comparison between GPR111-CTF, GPR111-CTF-LF, GPR111-CTF-ΔGPS, and GPR111-FL at respective transfection amount. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.9186, 0.3976, 0.2471, 0.9507 from left to right for GPR111-CTF-LF curve; P=0.9590, 0.2779, 0.0085, 0.0004 from left to right for GPR111-FL curve; P=0.8990, 0.2465, 0.0004, <0.0001 from left to right for GPR111-CTF-ΔGPS curve). (m), ELISA data to determine the cell surface expression levels of GPR111-CTF, GPR111-CTF-LF, GPR111-FL, and GPR111-CTF-ΔGPS in HEK293 cells. Relative expression levels compared to WT GPR111-CTF (set as 1) are shown. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate. ns, no significant difference. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.9660, 0.8201, 0.2839 from left to right). (n-o), Expression-response curves and bar graphs for GPR115-CTF, GPR115-CTF-VV (GPR115-CTF N-terminal elongation with Vss-−2 and Vss-−1), GPR115-FL, and GPR115-CTF-ΔGPS in BRET assay. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate. ***, P<0.001; **, P<0.01; ns, no significance. Comparison between GPR115-CTF, GPR115-CTF-VV, GPR115-FL, and GPR115-CTF-ΔGPS at respective transfection amount. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P>0.9999, =0.3456, 0.6854, 0.7109 from left to right for GPR115-CTF-LF curve; P>0.9999, =0.0215, 0.0002, 0.0010 from left to right for GPR115-FL curve; P>0.9999, =0.0012, <0.0001, <0.0001 from left to right for GPR115-CTF-ΔGPS curve). (p), ELISA data to determine the cell surface expression levels of the GPR115-CTF, GPR115-CTF-VV, GPR115-FL, and GPR115-CTF-ΔGPS in HEK293 cells. Relative expression levels compared to WT GPR115-CTF (set as 1) are shown. Values are the mean ± SEM of three independent experiments (n=3) performed in triplicate. ns, no significant difference. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.8281, 0.7432, 0.3261 from left to right).
Extended Data Fig. 9 Molecular mechanisms of the active state of GPR133-CTF and GPR114-CTF.
a, The toggle switch W6.53 position in GPR114-CTF, which directly interacts with key residues of the Stachel sequence. b, Detailed interactions of the UQC motif tethering TM3-TM6-TM7 in Gs-coupled GPR114-CTF. c, The outward shifts of TM6 and TM7 in GPR114-CTF. d, Detailed interactions of the P/V6.47φφG6.50 motif in TM6 with TM5 and TM7 in GPR114-CTF. Hydrogen bonds are highlighted by blue dashed lines. e, The effects of mutations in the P/V6.47φφG6.50 motif on the basal activities of GPR114-CTF. The bar graphs were generated based on the representative differences between WT GPR114-CTF and its mutants (MUT) at a relative expression level of 1.5 for GPR114-CTF, shown in Extended Data Fig. 9g. Data are normalized and presented as the response percentage for WT GPR114-CTF, and are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. ***, P < 0.001; **, P<0.01. All data were analyzed by two-sided one-way ANOVA with Tukey’s test (P<0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001, <0.0001 from left to right). f-g, ELISA data of the P6.47/V6.47φφG6.50 motif and H(N)L(M)Y motif to determine the cell surface expression levels of WT GPR133-CTF (f) and GPR114-CTF (g) or mutants (MUT) in HEK293 cells. The x-axis shows the amount of plasmid. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of the WT GPR133-CTF at the lowest transfected DNA amount (indicated as 1). The mutants show no significant difference from the WT. Data is given as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistically significant differences between WT and mutations were determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.1493, 0.3963, 0.1717, 0.3963, 0.1717, 0.1494, 0.1493, 0.2596, 0.0686, 0.0816, 0.1837, 0.8972, 0.1515, 0.7702, 0.1693, 0.7280, 0.9719, 0.9935, 0.4168, 0.2119, 0.2927, 0.5876, 0.7719, 0.1109, 0.6018, 0.1098, 0.5852, 0.5433, 0.8103, 0.3385, 0.2500, 0.2901, 0.4567, 0.4565, 0.0531, 0.4221, 0.0554, 0.4088, 0.7554, 0.4565, 0.2060, 0.2099, 0.2426, 0.1916 from left to right for figure f; P=0.9239, 0.9066, 0.9400, 0.4204, 0.4226, 0.4226, 0.4226, 0.7931, 0.4372, >0.9999, 0.7235, 0.1936, 0.7829, 0.6508, 0.8823, 0.9493, 0.6694, 0.9958, 0.3211, >0.9999, 0.8288, 0.7490, 0.7456, 0.4023, 0.9157, 0.9048, 0.6303, 0.9166, 0.0690, >0.9999, 0.9147, 0.6926, 0.6479, 0.8901, 0.4141, 0.5741, 0.9948, 0.7041, 0.4770, >0.9999 from left to right for figure g). h-i, Expression-response curves of the mutants in the P6.47/V6.47φφG6.50 motif and H(N)L(M)Y motif of GPR133-CTF (h) and GPR114-CTF (i) in cAMP accumulation assay. Values are the mean ± SEM of three independent experiments (n=3) for the WT and mutants. All data were analysed by two-sided one-way ANOVA with Tukey’s test (h: P=0.0003, <0.0001, <0.0001, <0.0001 from left to right for H656A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for L657A curve; P=0.3262, <0.0001, <0.0001, <0.0001 from left to right for Y658A curve; P=0.0002, <0.0001, <0.0001, <0.0001 from left to right for N727A curve; P=0.0002, <0.0001, <0.0001, <0.0001 from left to right for P767A curve; P=0.0002, <0.0001, <0.0001, <0.0001 from left to right for I768A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for L769A curve; P<0.0001, <0.0001, <0.0001, <0.0001 from left to right for Q798A curve; P=0.0002, <0.0001, <0.0001, <0.0001 from left to right for F805A curve; P=0.0005, <0.0001, <0.0001, <0.0001 from left to right for H806A curve; i: P= 0.0132, <0.0001, =0.0013, =0.0004 from left to right for N338A curve; P= 0.0100, <0.0001, =0.0002, =0.0002 from left to right for L339A curve; P= 0.1161, <0.0001, =0.0002, =0.0001 from left to right for Y340A curve; P= 0.1161, <0.0001, =0.0001, <0.0001 from left to right for N427A curve; P= 0.0474, <0.0001, =0.0001, =0.0001 from left to right for L431A curve; P> 0.9999, <0.0001, =0.0002, =0.0001 from left to right for V464A curve; P= 0.1890, =0.0029, =0.0002, =0.0002 from left to right for L466A curve; P= 0.1161, <0.0001, =0.0002, =0.0004 from left to right for Y495A curve; P= 0.3739, =0.0002, =0.0002, =0.0001 from left to right for F498A curve; P= 0.1161, <0.0001, =0.0002, =0.0001 from left to right for L499A curve).
Extended Data Fig. 10 The active state and G-protein coupling of GPR133-CTF and GPR114-CTF.
a, The cytoplasmic view of the GPR97 TMD compared with GPR133-CTF and GPR114-CTF. The distance of the cytosol-oriented part of TM3 and TM6 of GPR133-CTF and GPR114-CTF is larger than in GPR97. b, The interactions of the HLY motif with the α5 helix of Gαs in GPR133-CTF-Gs, GPR114-CTF-Gs and GPR97-Go complex. Complex superimposed based on TM3. Hydrogen bonds are highlighted by blue dashed lines. c. Effects of mutations in the H(N)LY motif of GPR133-CTF (upper panel) and GPR114-CTF (lower panel) on their basal activity. Mutations of any of these residues in both GPR133-CTF and GPR114-CTF significantly decreased their basal activity suggesting the functional importance of these positions in these aGPCRs. Represented differences between WT and its mutants expressed at relative expression level 3 for GPR133-CTF and level 1 for GPR114-CTF. Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistical significance of differences between WT and mutations was determined by two-sided one-way ANOVA with Tukey test. ***, P<0.001; **, P<0.01 (P<0.0001, <0.0001, <0.0001, from top to bottom for upper panel; P=0.0013, 0.0002, 0.0002 from top to bottom for lower panel). d, Effects of p133 (left panel) or p114 (right panel)-induced cAMP accumulation in HEK293 cells expressing the mutants at the H(N)LY motif of GPR133-CTF-ΔGPS (GPR133-CTF with truncated Stachel sequence, left panel) and GPR114-CTF-ΔGPS (GPR114-CTF with truncated Stachel sequence, right panel). Mutations of any of these residues in both GPR133-CTF and GPR114-CTF significantly decreased the Stachel peptide-induced activation. Data was normalized according to WT p133 (left panel) or p114 (right panel) and presented as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistical significance of differences between receptor WT and mutations was determined by two-sided one-way ANOVA with Tukey test. ***, P<0.001 (P<0.0001, <0.0001, =0.0003 from top to bottom for left panel; P<0.0001, <0.0001 from top to bottom for right panel). e, Concentration-response curves of WT GPR133-CTF-ΔGPS (GPR133-CTF with truncated Stachel sequence) and GPR114-CTF-ΔGPS (GPR114-CTF with truncated Stachel sequence) or mutants at H(N)L(M)Y motif under the incubation with p133 (left panel) or p114 (right panel). Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. f, Comparison of TM6 and TM7 in Go-coupled GPR97 (7D77), Gs-coupled GPR133-CTF, and Gs-coupled GPR114-CTF. The separation distances of TM6 and TM7 in GPR133-CTF and GPR114-CTF were smaller than in GPR97. The rotational shift of the α5 helix is due to the smaller separation between TM6 and TM7 in the two Gs-coupled aGPCR structures compared to the GPR97-Go complex. The insertion of the palmitoyl moiety of the Go protein into the TMD of GPR97 increased the separation between TM6 and TM7. g, Residues in GPR133-CTF, GPR114-CTF and GPR97 TMD (PDB:7D77) that contact their corresponding G proteins.
Extended Data Fig. 11 Molecular mechanisms of Gs coupling to GPR133-CTF and GPR114-CTF.
a-c, Cryo-EM density of the conserved interaction residues that mediate G-protein coupling in the GPR133-CTF-Gs and GPR114-CTF-Gs complex. (a), receptor residues interacting with Y391 and Q390 of the α5 helix of Gαs; (b), H387 and L388 in the Gs α5 helix form hydrophobic packings with residues of TM3 and TM5, including M660/L3423.57, V661/L3433.58, and I738/L4385.61 in GPR133-CTF and GPR114-CTF; (c), VICL2F/YICL2 motif in ICL2 of GPR133-CTF and GPR114-CTF located in a large hydrophobic pocket created by H41G.S1.02, V217G.S3.01, F219G.S3.03, F376G.H5.08, R380G.H5.12, I383G.H5.15, and Q384G.H5.16 from the α5 helix, β3 strand and αN-β1 junctions of Gs. d, Effects of mutations in the observed Gs interfaces of GPR133-CTF on its basal activities. Relative differences between WT and its mutants (MUT) at a relative expression level of 3 for GPR133-CTF (refer to ELISA data on receptor’s cell surface expression level in Extended Data Fig. 11e) are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistically significant differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey’s test. ***, P<0.001; **, P<0.01; *, P<0.05 (P=0.0001, =0.0010, <0.0001, <0.0001, =0.0004, =0.0002, <0.0001, <0.0001, <0.0001 from top to bottom). e, ELISA data to determine the cell surface expression levels of WT and G-protein interface mutants (MUT) of GPR133-CTF in HEK293 cells. The x-axis shows the amount of transfected plasmid DNA. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of the WT GPR133-CTF at the lowest amount of transfected DNA (indicated as 1). Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistically significant differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.1744, 0.3963, 0.1717, 0.1494, 0.0609, 0.0816, 0.2596, 0.1837, 0.0686, 0.6987, 0.1515, 0.7702, 0.9719, 0.6222, 0.2927, 0.4168, 0.5876, 0.2119, 0.5749, 0.1109, 0.6018, 0.5433, 0.8777, 0.2901, 0.3385, 0.4567, 0.2500, 0.3117, 0.0531, 0.4221, 0.7554, 0.2124, 0.2426, 0.2060, 0.1916, 0.2099 from left to right). f, Expression-response curves of WT and receptor mutants on the interface with the Gs protein of GPR133-CTF in cAMP accumulation assay. Values are the mean ± SEM of three independent experiments (n=3) for WT and mutants. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P=0.9615, <0.0001, =0.0001, <0.0001 from left to right for R601A curve; P=0.9615, <0.0001, =0.0010, =0.1053 from left to right for V661A curve; P=0.9615, <0.0001, <0.0001, <0.0001 from left to right for V664A curve; P=0.8195, <0.0001, <0.0001, <0.0001 from left to right for F665A curve; P=0.9615, <0.0001, =0.0004, =0.0266 from left to right for I738A curve; P=0.9615, <0.0001, =0.0002, <0.0001 from left to right for I741A curve; P=0.9494, <0.0001, <0.0001, <0.0001 from left to right for V764A curve; P=0.9889, <0.0001, <0.0001, <0.0001 from left to right for N810A curve; P=0.9930, <0.0001, <0.0001, <0.0001 from left to right for V823A curve). g, Effects of mutations in the observed Gs interfaces of GPR114-CTF on its basal activities. Relative differences between WT and its mutants (MUT) at a relative expression level of 1.5 for GPR114-CTF (refer to ELISA data on receptor’s cell surface expression level in Extended Data Fig. 11h) are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistically significant differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey’s test. ***, P<0.001;**, P<0.01; *, P<0.05 (P = 0.0105, 0.0145, 0.0003, 0.0003, 0.0003, 0.0003, 0.0002, 0.0117, 0.0093 from top to bottom). h, ELISA data to determine the cell surface expression levels of WT and G-protein interface mutants of GPR114-CTF in HEK293 cells. In the figure, MUT refers to mutant. The x-axis shows the amount of transfected plasmid DNA. The y-axis shows the ratio of corresponding receptor expression levels compared with the expression level of the WT GPR114-CTF at the lowest amount of transfected DNA (indicated as 1). Values are shown as the mean ± SEM of three experiments (n=3) performed in triplicate. Statistically significant differences between WT and mutants were determined by two-sided one-way ANOVA with Tukey test. ns, no significant difference (P=0.9257, 0.9629, 0.9890, 0.9885, 0.9470, 0.9873, 0.8718, 0.7410, >0.9999, 0.9572, 0.9630, 0.9421, 0.2164, 0.9287, 0.9122, 0.6140, 0.6078, 0.9342, 0.9776, 0.1833, 0.2764, 0.4905, 0.8861, 0.8497, 0.6788, 0.4138, 0.2324, 0.4640, 0.5049, 0.8859, 0.4774, 0.5553, 0.9974, 0.0803, 0.3843, 0.4444 from left to right). i, Expression-response curves of WT and receptor mutants on the interface with the Gs protein of GPR114-CTF in cAMP accumulation assay. Values are the mean ± SEM of three independent experiments (n=3) for WT and mutants. All data were analysed by two-sided one-way ANOVA with Tukey’s test (P= 0.0031, =0.7780, =0.0749, =0.0023 from left to right for L280A curve; P= 0.0020, =0.0003, =0.0145, =0.0140 from left to right for L342A curve; P= 0.0111, <0.0001, =0.0003, =0.0002 from left to right for L343A curve; P= 0.2378, <0.0001, =0.0003, =0.0002 from left to right for V346A curve; P= 0.0668, <0.0001, =0.0003, =0.0002 from left to right for T437A curve; P= 0.0647, <0.0001, =0.0003, =0.0006 from left to right for L441A curve; P= 0.0474, <0.0001, =0.0002, =0.0001 from left to right for L465A curve; P= 0.0178, =0.0004, =0.0927, =0.0018 from left to right for W502A curve; P= 0.0011, =0.0230, =0.0093, =0.0221 from left to right for Q506A curve). j, The detailed interactions of ICL2 with the α5 helix in GPR114-CTF-Gs-scFv16 complex, including hydrophobic interactions and hydrogen bonds formed between Y350ICL2 and K34G.HN.51, Q35G.HN.52 and R38G.hns1.02. Hydrogen bonds are highlighted by blue dashed lines. k, The detailed interactions of ICL1 with Gβ in GPR133-CTF-Gs-Nb35 complex. V594, L595 and R598 in GPR133 directly interact with R52, D312, D333 and F335 in the Gβ subunit. l, Structural superposition of GPR133-CTF-Gs and the GPR114-CTF-Gs complex. The extended TM7 in GPR114-CTF and helix 8 in GPR133-CTF are highlighted.
Supplementary information
Supplementary Information
This file contains Supplementary Figs. 1–8 and Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Ping, YQ., Xiao, P., Yang, F. et al. Structural basis for the tethered peptide activation of adhesion GPCRs. Nature 604, 763–770 (2022). https://doi.org/10.1038/s41586-022-04619-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04619-y
This article is cited by
-
CD97 inhibits osteoclast differentiation via Rap1a/ERK pathway under compression
International Journal of Oral Science (2024)
-
A method for structure determination of GPCRs in various states
Nature Chemical Biology (2024)
-
Constitutive activation mechanism of a class C GPCR
Nature Structural & Molecular Biology (2024)
-
Structure, function and drug discovery of GPCR signaling
Molecular Biomedicine (2023)
-
The activation mechanism and antibody binding mode for orphan GPR20
Cell Discovery (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.