Abstract
The capacity of planktonic marine microorganisms to actively seek out and exploit microscale chemical hotspots has been widely theorized to affect ocean-basin scale biogeochemistry1,2,3, but has never been examined comprehensively in situ among natural microbial communities. Here, using a field-based microfluidic platform to quantify the behavioural responses of marine bacteria and archaea, we observed significant levels of chemotaxis towards microscale hotspots of phytoplankton-derived dissolved organic matter (DOM) at a coastal field site across multiple deployments, spanning several months. Microscale metagenomics revealed that a wide diversity of marine prokaryotes, spanning 27 bacterial and 2 archaeal phyla, displayed chemotaxis towards microscale patches of DOM derived from ten globally distributed phytoplankton species. The distinct DOM composition of each phytoplankton species attracted phylogenetically and functionally discrete populations of bacteria and archaea, with 54% of chemotactic prokaryotes displaying highly specific responses to the DOM derived from only one or two phytoplankton species. Prokaryotes exhibiting chemotaxis towards phytoplankton-derived compounds were significantly enriched in the capacity to transport and metabolize specific phytoplankton-derived chemicals, and displayed enrichment in functions conducive to symbiotic relationships, including genes involved in the production of siderophores, B vitamins and growth-promoting hormones. Our findings demonstrate that the swimming behaviour of natural prokaryotic assemblages is governed by specific chemical cues, which dictate important biogeochemical transformation processes and the establishment of ecological interactions that structure the base of the marine food web.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The raw metabolome data files were deposited in MetaboLights under accession number MTBLS1980. The raw metagenome fastq files were deposited in the Sequence Read Archive (SRA) under accession number PRJNA639602. The raw amplicon fastq files were deposited in the SRA under accession number PRJNA707306. The 16S rRNA gene sequences of the three isolates were deposited in GenBank under accession numbers: MT826233, MT826234 and MZ373175. Source data are provided with this paper.
Code availability
All custom analysis scripts are available on GitHub (https://github.com/JB-Raina-codes/ISCA-paper).
References
Azam, F. & Malfatti, F. Microbial structuring of marine ecosystems. Nat. Rev. Microbiol. 5, 782–791 (2007).
Blackburn, N., Fenchel, T. & Mitchell, J. Microscale nutrient patches in planktonic habitats shown by chemotactic bacteria. Science 282, 2254–2256 (1998).
Stocker, R. Marine microbes see a sea of gradients. Science 338, 628 (2012).
Levin, S. A. The problem of pattern and scale in ecology. Ecology 73, 1943–1967 (1992).
Azam, F. Microbial control of oceanic carbon flux: the plot thickens. Science 280, 694–696 (1998).
Strom, S. L. Microbial ecology of ocean biogeochemistry: a community perspective. Science 320, 1043–1045 (2008).
Sarmento, H. & Gasol, J. M. Use of phytoplankton-derived dissolved organic carbon by different types of bacterioplankton. Env. Microbiol. 14, 2348–2360 (2012).
Grossart, H.-P., Riemann, L. & Azam, F. Bacterial motility in the sea and its ecological implications. Aquat. Microb. Ecol. 25, 247–258 (2001).
Brumley, D. R. et al. Bacteria push the limits of chemotactic precision to navigate dynamic chemical gradients. Proc. Natl Acad. Sci. USA 116, 10792–10797 (2019).
Fenchel, T. Eppur si muove: many water column bacteria are motile. Aquat. Microb. Ecol. 24, 197–201 (2001).
Son, K., Menolascina, F. & Stocker, R. Speed-dependent chemotactic precision in marine bacteria. Proc. Natl Acad. Sci. USA 113, 8624–8629 (2016).
Fenchel, T. Microbial behavior in a heterogeneous world. Science 296, 1068–1071 (2002).
Kiørboe, T. & Jackson, G. A. Marine snow, organic solute plumes, and optimal chemosensory behavior of bacteria. Limnol. Oceanogr. 46, 1309–1318 (2001).
Lambert, B. S., Fernandez, V. I. & Stocker, R. Motility drives bacterial encounter with particles responsible for carbon export throughout the ocean. Limnol. Oceanogr. Lett. 4, 113–118 (2019).
Wadhams, G. H. & Armitage, J. P. Making sense of it all: bacterial chemotaxis. Nat. Rev. Mol. Cell. Biol. 5, 1024–1037 (2004).
Stocker, R., Seymour, J. R., Samadani, A., Hunt, D. E. & Polz, M. F. Rapid chemotactic response enables marine bacteria to exploit ephemeral microscale nutrient patches. Proc. Natl Acad. Sci. USA 105, 4209–4214 (2008).
Raina, J.-B., Fernandez, V., Lambert, B., Stocker, R. & Seymour, J. R. The role of microbial motility and chemotaxis in symbiosis. Nat. Rev. Microbiol. 17, 284–294 (2019).
Seymour, J. R., Amin, S. A., Raina, J.-B. & Stocker, R. Zooming in on the phycosphere: the ecological interface for phytoplankton–bacteria relationships. Nat. Microbiol. 2, 17065 (2017).
Bell, W. & Mitchell, R. Chemotactic and growth responses of marine bacteria to algal extracellular products. Biol. Bull. 143, 265–277 (1972).
Smriga, S., Fernandez, V. I., Mitchell, J. G. & Stocker, R. Chemotaxis toward phytoplankton drives organic matter partitioning among marine bacteria. Proc. Natl Acad. Sci. USA 113, 1576–1581 (2016).
Amin, S. A. et al. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature 522, 98–101 (2015).
Lambert, B. S. et al. A microfluidics-based in situ chemotaxis assay to study the behaviour of aquatic microbial communities. Nat. Microbiol. 2, 1344–1349 (2017).
Larsen, M. H., Blackburn, N., Larsen, J. L. & Olsen, J. E. Influences of temperature, salinity and starvation on the motility and chemotactic response of Vibrio anguillarum. Microbiology 150, 1283–1290 (2004).
Rinke, C. et al. Validation of picogram- and femtogram-input DNA libraries for microscale metagenomics. PeerJ 4, e2486 (2016).
Becker, J. et al. Closely related phytoplankton species produce similar suites of dissolved organic matter. Front. Microbiol. 5, 111 (2014).
Vraspir, J. M. & Butler, A. Chemistry of marine ligands and siderophores. Annu. Rev. Mar. Sci. 1, 43–63 (2009).
Tagliabue, A. et al. The integral role of iron in ocean biogeochemistry. Nature 543, 51–59 (2017).
Hopkinson, B. M. & Morel, F. M. M. The role of siderophores in iron acquisition by photosynthetic marine microorganisms. BioMetals 22, 659–669 (2009).
Amin, S. A. et al. Photolysis of iron–siderophore chelates promotes bacterial–algal mutualism. Proc. Natl Acad. Sci. USA 106, 17071–17076 (2009).
Croft, M. T., Lawrence, A. D., Raux-Deery, E., Warren, M. J. & Smith, A. G. Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature 438, 90–93 (2005).
Helliwell, K. E. The roles of B vitamins in phytoplankton nutrition: new perspectives and prospects. New Phytol. 216, 62–68 (2017).
Berg, G. Plant–microbe interactions promoting plant growth and health: perspectives for controlled use of microorganisms in agriculture. Appl. Microbiol. Biotechnol. 84, 11–18 (2009).
Christie, P. J., Whitaker, N. & González-Rivera, C. Mechanism and structure of the bacterial type IV secretion systems. Biochim. Biophys. Acta 1843, 1578–1591 (2014).
Preston, G. M. Metropolitan microbes: type III secretion in multihost symbionts. Cell Host Microbe 2, 291–294 (2007).
Deakin, W. J. & Broughton, W. J. Symbiotic use of pathogenic strategies: rhizobial protein secretion systems. Nat. Rev. Microbiol. 7, 312–320 (2009).
Luo, H. & Moran, M. A. Evolutionary ecology of the marine Roseobacter clade. Microbiol. Mol. Biol. Rev. 78, 573–587 (2014).
Rolland, J. L., Stien, D., Sanchez-Ferandin, S. & Lami, R. Quorum sensing and quorum quenching in the phycosphere of phytoplankton: a case of chemical interactions in ecology. J. Chem. Ecol. 42, 1201–1211 (2016).
Fei, C. et al. Quorum sensing regulates ‘swim-or-stick’ lifestyle in the phycosphere. Environ. Microbiol. 22, 4761–4778 (2020).
Landa, M., Burns, A. S., Roth, S. J. & Moran, M. A. Bacterial transcriptome remodeling during sequential co-culture with a marine dinoflagellate and diatom. ISME J. 11, 2677 (2017).
Rinke, C. et al. A phylogenomic and ecological analysis of the globally abundant Marine Group II archaea (Ca. Poseidoniales ord. nov.). ISME J. 13, 663–675 (2019).
Fenchel, T. & Blackburn, N. Motile chemosensory behaviour of phagotrophic protists: mechanisms for and efficiency in congregating at food patches. Protist 150, 325–336 (1999).
Hughes, D. J. et al. Impact of nitrogen availability upon the electron requirement for carbon fixation in Australian coastal phytoplankton communities. 63, 1891–1910 (2018).
Sumner, L. W. et al. Proposed minimum reporting standards for chemical analysis. Metabolomics 3, 211–221 (2007).
Chong, J., Wishart, D. S. & Xia, J. Using MetaboAnalyst 4.0 for comprehensive and integrative metabolomics data analysis. Curr. Protoc. Bioinformatics 68, e86 (2019).
Xia, J. & Wishart, D. S. Web-based inference of biological patterns, functions and pathways from metabolomic data using MetaboAnalyst. Nat. Protoc. 6, 743–760 (2011).
Lambert, B. S. & Raina, J.-B. Fabrication and deployment of the in situ chemotaxis assay (ISCA). protocols.io https://doi.org/10.17504/protocols.io.kztcx6n (2019).
Ritchie, R. J. Consistent sets of spectrophotometric chlorophyll equations for acetone, methanol and ethanol solvents. Photosynth. Res. 89, 27–41 (2006).
Marie, D., Partensky, F., Jacquet, S. & Vaulot, D. Enumeration and cell cycle analysis of natural populations of marine picoplankton by flow cytometry using the nucleic acid stain SYBR Green I. Appl. Environ. Microbiol. 63, 186–193 (1997).
Bramucci, A. R. et al. Microvolume DNA extraction methods for microscale amplicon and metagenomic studies. ISME Commun. 1, 79 (2021).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://doi.org/10.48550/arXiv.1303.3997 (2013).
Boyd, J. A., Woodcroft, B. J. & Tyson, G. W. GraftM: a tool for scalable, phylogenetically informed classification of genes within metagenomes. Nucleic Acids Res. 46, e59 (2018).
Kuever, J., Rainey, F. A. & Widdel, F. In Bergey’s Manual of Systematics of Archaea and Bacteria https://doi.org/10.1002/9781118960608.obm00084 (2015).
Bianchi, D., Weber, T. S., Kiko, R. & Deutsch, C. Global niche of marine anaerobic metabolisms expanded by particle microenvironments. Nat. Geosci. 11, 263–268 (2018).
Liu, X. et al. Wide distribution of anaerobic ammonium-oxidizing bacteria in the water column of the South China Sea: implications for their survival strategies. Divers. Distrib. 27, 1893–19003 (2021).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Suzek, B. E., Huang, H., McGarvey, P., Mazumder, R. & Wu, C. H. UniRef: comprehensive and non-redundant UniProt reference clusters. Bioinformatics 23, 1282–1288 (2007).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59 (2014).
Paulson, J. N., Stine, O. C., Bravo, H. C. & Pop, M. Differential abundance analysis for microbial marker-gene surveys. Nat. Methods 10, 1200 (2013).
McMurdie, P. J. & Holmes, S. Waste not, want not: why rarefying microbiome data is inadmissible. PLOS Comput. Biol. 10, e1003531 (2014).
Berges, J. A., Franklin, D. J. & Harrison, P. J. Evolution of an artificial seawater medium: improvements in enriched seawater, artificial water over the last two decades. J. Phycol. 37, 1138–1145 (2001).
Lane, D. In Nucleic Acid Techniques in Bacterial Systematics (eds Stackebrandt, E. & Goodfellow, M.) 115–175 (1991).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nature Methods 13, 581–583 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Oksanen, J. et al. Package ‘Vegan’ Community Ecology Package Version 2 (2013).
Durham, B. P. et al. Sulfonate-based networks between eukaryotic phytoplankton and heterotrophic bacteria in the surface ocean. Nat. Microbiol. 4, 1706–1715 (2019).
Durham, B. P. et al. Recognition cascade and metabolite transfer in a marine bacteria–phytoplankton model system. Environ. Microbiol. 19, 3500–3513 (2017).
Durham, B. P. et al. Cryptic carbon and sulfur cycling between surface ocean plankton. Proc. Natl Acad. Sci. USA 112, 453–457 (2015).
Landa, M. et al. Sulfur metabolites that facilitate oceanic phytoplankton–bacteria carbon flux. ISME J. 13, 2536–2550 (2019).
Acknowledgements
The authors thank G. Gorick for his work on Fig. 1a, L. Bennar, G. Kholi, M. Fabris, N. Le Reun, M. Giardina and D. Hughes for laboratory assistance and E. Botté for her support throughout this project. This work was supported by the Gordon and Betty Moore Foundation through a grant to J.R.S., G.W.T., P.H. and R.S. (GBMF3801) and two Investigator Awards to R.S. (GBMF3783 and GBMF9197; https://doi.org/10.37807/GBMF9197), and through Australian Research Council grants DP110103091 to J.R.S., G.W.T. and R.S and DP180100838 to J.R.S., R.S. and J.-B.R. B.S.L. was supported by the Simons Foundation (Award 594111). G.W.T. was supported by an Australian Research Council Future Fellowship (FT170100070), P.H. was supported by Australian Research Council Laureate Fellowship (FL150100038). C.R. was supported by an Australian Research Council Future Fellowship (FT170100213). J.-B.R. was supported by an Australian Research Council Fellowship (DE160100636).
Author information
Authors and Affiliations
Contributions
J.-B.R., B.S.L., C.R., R.S., P.H., G.W.T. and J.R.S. designed all the experiments. J.-B.R., C.R., A.B. and N.S. performed all the experiments. J.-B.R., A.L. and H.M. generated and analysed the metabolomic data. J.-B.R., C.R., F.R., B.S.L. and D.H.P. generated and analysed the metagenomic data. J.-B.R., M.O., B.S.L., A.B., V.I.F. and B.S. generated and analysed the amplicon data and performed the network and correlative analyses. J.-B.R., B.S.L., R.S. and J.R.S. wrote the manuscript. All authors edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Hans-Peter Grossart, Karla Heidelberg and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 ISCA deployments through a two years period at Clovelly Beach (33.91°S, 151.26°E).
Chemotactic index Ic, denoting the concentration of cells within ISCA wells, normalized by the mean concentration of cells within wells containing filtered seawater (FSW), after 60 min field deployment. Solid bars are significantly different from wells containing FSW (ANOVA (one-sided), p < 0.05, all p-values are reported in Supplementary Table 3). Each treatment was replicated across four different ISCAs (n = 4), except between April and August 2016 (n = 3). Data are presented as mean values ± SEM. FSW: filtered seawater, Syne: Synechococcus, Proch: Prochlorococcus, Duna: Dunaliella, Rhodo; Rhodomonas, Phaeo: Phaeocystis, Ehux: Emiliania, Prym: Prymnesium, Chae: Chaetoceros, Dityl: Ditylum, Phae: Phaeodactylum, Thala: Thalassiosira, Durus: Durusdinium, Alex: Alexandrium, Amphi: Amphidinium.
Extended Data Fig. 2 Environmental variables influencing the strength of chemotaxis.
(a) Average chemotactic index (Ic) elicited by the phytoplankton-derived DOM for each of the 12 ISCA deployments described in this study at Clovelly Beach (33.91°S, 151.26°E). Error bars: standard errors. (b) Correlogram of the metadata measured during each deployment (the size and colour of each bubble is proportional to the strength of the correlation). Only statistically significant correlations are not crossed (Pearson’s correlation (two-sided), p < 0.01). (c) Significant correlation between chemotactic index and temperature (Pearson’s correlation (two-sided), p < 0.01).
Extended Data Fig. 3 Differences in chemical composition between the phytoplankton-derived DOM.
(a) Heatmap of the 111 compounds identified between the different phytoplankton species. Data were log-transformed and mean centred. An interactive version of this figure is available (Fig. S2). (b) Principal component analysis (PCA) of chemical composition of the phytoplankton-derived DOM: displaying the top three components (explaining 64.7% of the variance).
Extended Data Fig. 4 Relative abundance of the prokaryotic families present in the bulk seawater, the FSW controls, and in the different phytoplankton-derived DOM.
Only taxa representing more than 2% of the communities are displayed in colours, those representing less than 2% and grouped as “Other”. ND: taxonomy not determined at the family level.
Extended Data Fig. 5 Prokaryotic taxa significantly enriched the phytoplankton-derived DOM treatments.
(a) Number of prokaryotic taxa enriched in each phytoplankton-DOM treatment (compared to filtered seawater controls). The full list of taxa significantly enriched in phytoplankton-derived DOM treatments can be found in Supplementary Table 6. (b) Network analysis showing the differentiation between “generalist” and “specialist” families at the taxonomic level. This network has the same topology than the Figure 2b. Chemotactic prokaryotic taxa (small circles; nodes) are linked to the treatments they responded to (large circles) by lines with colours corresponding to each treatment. Each node is colour coded based on its taxonomy. (c) Number of prokaryote taxa significantly enriched in one or more phytoplankton-derived DOM treatments (compared to filtered seawater controls). Another graphical representation of this data can be found in Figure 2b.
Extended Data Fig. 6 Genes involved in motility, chemotaxis and surface-attachment were significantly enriched in the ISCA treatments compared to the bulk seawater.
Data were log-transformed and mean-centred (n = 4) for each ISCA treatment. Asterisks highlight significant enrichment compared to the bulk seawater (F-tests (one-sided), p < 0.05).
Extended Data Fig. 7 Genes involved in the uptake and degradation of phytoplankton-derived molecules (selected from the literature)39,67,68,69,70, as well as in the transport of a range of labile substrates, were significantly enriched in the prokaryotic communities responding to phytoplankton-derived DOM.
Data were log-transformed and mean-centred (n = 4) for each ISCA treatment. Asterisks highlight significant enrichment compared to the FSW treatment (F-tests (one-sided), p < 0.05, all p-values are reported in Supplementary Table 8). DMSP: dimethylsulfoniopropionate; DHPS: 2,3-dihydroxypropane-1-sulfonate; GBT: Glycine betaine.
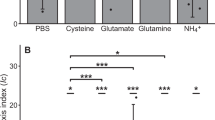
Extended Data Fig. 8 Assay testing the ability of the bacterial isolates to catabolize the validated chemoattractants (Figure 4b).
Each chemoattractant was inoculated at a concentration of 1 mM (n = 4) in an artificial seawater medium supplemented with 0.2% of casamino acids. After 48 h, the optical density (OD600) of each culture was compared to controls only containing casamino acids. DGDG: Digalactosyldiacylglycerol; 3-aminopip: 3-aminopiperidin-2-one. Solid bars are significantly different from wells containing FSW (ANOVA (one-sided), p < 0.05, all p-values are reported in Supplementary Table 9). Data are presented as mean values ± SEM.
Extended Data Fig. 9 Control for bacterial growth during the ISCA deployment time.
(a) Comparison of prokaryotic cell counts before, and then 1 h and 5 h after post incubation with phytoplankton-derived DOM (1 mg mL−1). The number of prokaryotic cells were not statistically different between pre-incubation and one hour of incubation (ANOVA (one-sided), n = 3, p = 0.8026). Data are presented as mean values ± SEM. (b) Principal component analysis (PCA) of bacterial community composition resulting from incubations (explaining 80.4% of the variance), revealing the overlap between bacterial community compositions pre-incubation and those after 1 h of incubation. An analysis of similarities confirmed that community compositions were not significantly different pre-incubation and after one hour of incubation (ANOSIM; 99,999 permutations; n = 33; R = 0.108; p = 0.2), but significant differences were observed after five hours (R = 0.602; p = 0.001).
Extended Data Fig. 10 Control for shifts in bacterial composition during the ISCA deployment time.
Relative abundance of the bacterial communities (at the ASV level) before, 1 h and 5 h of incubation with phytoplankton-derived DOM (1 mg mL−1). The legend only shows the 30 most abundant ASVs.
Supplementary information
Supplementary Information
This file contains Supplementary Fig. 1; legend for Supplementary Fig. 2; Supplementary Tables 2, 10, 11; legends for Supplementary Tables 1, 3–9, 12; and Supplementary References.
Supplementary Figure 2
See Supplementary Information file for Supplementary Fig. 2 legend.
Supplementary Table 1
Occurrence of the eukaryotic phytoplankton genera used for our in situ chemotaxis assay (raw counts) at three coastal sites near Sydney, Australia, over a two years period (Cobblers Beach, Salmon Haul, Taren Point). This table is too large to be displayed in this document and is available as a separate file.
Supplementary Table 3
Sums of squares (SS), mean squares (MS) and significance levels for the ANOVAs of the chemotactic responses reported in Fig. 1b and Extended Data Fig. 1, with simple main effect test (diff: differences in mean; lower and upper: confidence intervals). All data were log transformed to meet the test assumptions. This table is too large to be displayed in this document and is available as a separate file.
Supplementary Table 4
Summary of PERMANOVA comparing the chemical composition of the ten phytoplankton-derived DOM treatments. Bray–Curtis similarity, 999 unrestricted permutations, main effect and pairwise tests. Abbreviation: Alex: Alexandrium; Amphi: Amphidinium; Chae: Chaetoceros; Dityl: Ditylum; Duna: Dunaliella; Ehux: Emiliania; Phae: Phaeodactylum; Prym: Prymnesium; Syne: Synechococcus; Thala: Thalassiosira. This table is too large to be displayed in this document and is available as a separate file.
Supplementary Table 5
Summary of PERMANOVA comparing the taxonomic composition of the prokaryotic communities between the different treatments. Bray–Curtis similarity, 999 unrestricted permutations, main effect and pairwise tests. This table is too large to be displayed in this document and is available as a separate file.
Supplementary Table 6
Summary of the taxa-specific enrichments between filtered seawater (control) and phytoplankton-derived DOM treatments using the R package metagenomeSeq1. This table is too large to be displayed in this document and is available as a separate file.
Supplementary Table 7
Summary of PERMANOVA comparing the functional capabilities of the prokaryotic communities between the different treatments. Bray–Curtis similarity, 999 unrestricted permutations, main effect and pairwise tests. This table is too large to be displayed in this document and is available as a separate file.
Supplementary Table 8
Summary of the functional gene enrichment within phytoplankton-derived DOM treatments relative to the filtered seawater control using the R package metagenomeSeq1. This table is too large to be displayed in this document and is available as a separate file.
Supplementary Table 9
Sums of squares (SS), mean squares (MS) and significance levels for the ANOVAs of the chemotactic responses and growth reported in Fig. 4b, with simple main effect test (diff: differences in mean; lower and upper: confidence intervals). This table is too large to be displayed in this document and is available as a separate file.
Supplementary Table 12
Quality check results of the ISCA metagenomes and all additional controls (mock communities: “mock_ACE”; DNA extraction controls: “Neg_Feb”; Library prep controls: “Neg_ACE”; and undeployed ISCA treatments: “Control_undeployed”). This table is too large to be displayed in this document and is available as a separate file.
Rights and permissions
About this article
Cite this article
Raina, JB., Lambert, B.S., Parks, D.H. et al. Chemotaxis shapes the microscale organization of the ocean’s microbiome. Nature 605, 132–138 (2022). https://doi.org/10.1038/s41586-022-04614-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04614-3
This article is cited by
-
Algal blooms in the ocean: hot spots for chemically mediated microbial interactions
Nature Reviews Microbiology (2024)
-
Bacteria contribute exopolysaccharides to an algal-bacterial joint extracellular matrix
npj Biofilms and Microbiomes (2024)
-
Seasonal patterns in microbial carbon and iron transporter expression in the Southern Ocean
Microbiome (2023)
-
Strong chemotaxis by marine bacteria towards polysaccharides is enhanced by the abundant organosulfur compound DMSP
Nature Communications (2023)
-
Chemotaxis increases metabolic exchanges between marine picophytoplankton and heterotrophic bacteria
Nature Microbiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.