Abstract
The B.1.1.529/Omicron variant of SARS-CoV-2 was only recently detected in southern Africa, but its subsequent spread has been extensive, both regionally and globally1. It is expected to become dominant in the coming weeks2, probably due to enhanced transmissibility. A striking feature of this variant is the large number of spike mutations3 that pose a threat to the efficacy of current COVID-19 vaccines and antibody therapies4. This concern is amplified by the findings of our study. Here we found that B.1.1.529 is markedly resistant to neutralization by serum not only from patients who recovered from COVID-19, but also from individuals who were vaccinated with one of the four widely used COVID-19 vaccines. Even serum from individuals who were vaccinated and received a booster dose of mRNA-based vaccines exhibited substantially diminished neutralizing activity against B.1.1.529. By evaluating a panel of monoclonal antibodies against all known epitope clusters on the spike protein, we noted that the activity of 17 out of the 19 antibodies tested were either abolished or impaired, including ones that are currently authorized or approved for use in patients. Moreover, we also identified four new spike mutations (S371L, N440K, G446S and Q493R) that confer greater antibody resistance on B.1.1.529. The Omicron variant presents a serious threat to many existing COVID-19 vaccines and therapies, compelling the development of new interventions that anticipate the evolutionary trajectory of SARS-CoV-2.
Similar content being viewed by others
Main
The COVID-19 pandemic continues as the causative agent SARS-CoV-2 continues to evolve. Many diverse viral variants have emerged (Fig. 1a), each characterized by mutations in the spike protein that raise concerns of both antibody evasion and enhanced transmission. The B.1.351/Beta variant was found to be highly refractory to antibody neutralization4 and therefore compromised the efficacy of vaccines5,6,7 and therapeutic antibodies. The B.1.1.7/Alpha variant became dominant globally in early 2021 due to having an edge in transmission8, only to be replaced by the B.1.617.2/Delta variant, which exhibited an even greater propensity to spread coupled with a moderate level of antibody resistance9. Next came the B.1.1.529/Omicron variant, which was first detected in southern Africa in November 2021 (refs. 3,10,11) (Fig. 1a). Omicron has since spread rapidly in the region, as well as to over 60 countries, gaining traction even in regions in which the Delta variant is prevalent. The short doubling time (2–3 days) of Omicron cases suggests that it could soon become dominant2. Moreover, its spike protein contains an alarming number of mutations (>30) (Fig. 1b and Extended Data Fig. 1), including at least 15 in the receptor-binding domain (RBD), the principal target of neutralizing antibodies. These extensive spike mutations raise the possibility that current vaccines and therapeutic antibodies would be greatly compromised. This concern is amplified by the findings that we now report.
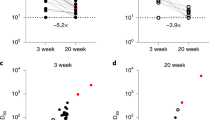
a, Unrooted phylogenetic tree of B.1.1.529 with other major SARS-CoV-2 variants. b, Key spike mutations found in the viruses isolated in the major lineage of B.1.1.529. Del, deletion; ins, insertion. c, Neutralization of D614G and B.1.1.529 pseudoviruses by convalescent patient sera. d, Neutralization of D614G and B.1.1.529 pseudoviruses by vaccinee sera. Within the four standard vaccination groups, individuals who were vaccinated without documented infection are denoted as circles and individuals who were both vaccinated and infected are denoted as triangles. Within the boosted group, Moderna vaccinees are denoted as squares and Pfizer vaccinees are denoted as diamonds. e, Neutralization of authentic D614G and B.1.1.529 viruses by vaccinee sera. Moderna vaccinees are denoted as squares and Pfizer vaccinees are denoted as diamonds. Data represent one out of two independent experiments. For all of the panels, the values above the symbols denote the geometric mean titre and the numbers in parentheses denote the number of samples above the limit of detection. P values were determined by using two-tailed Wilcoxon matched-pairs signed-rank tests.
Serum neutralization of B.1.1.529
We first examined the neutralizing activity of serum collected in spring 2020 from patients with COVID-19, who were presumably infected with the wild-type (WT) SARS-CoV-2 (9–120 days after symptoms) (Methods and Extended Data Table 1). Samples from ten individuals were tested for neutralization against both D614G (WT) and B.1.1.529 pseudoviruses. Although robust titres were observed against D614G, a significant drop (>32-fold) in 50% infectious dose (ID50) titres was observed against B.1.1.529, with only two samples showing titres above the limit of detection (LOD) (Fig. 1c and Extended Data Fig. 2a). We next assessed the neutralizing activity of sera from individuals who received one of the four widely used COVID-19 vaccines: BNT162b2 (Pfizer, 15–213 days after vaccination), mRNA-1273 (Moderna, 6–177 days after vaccination), Ad26.COV2.S (Johnson & Johnson, 50–186 days after vaccination) and ChAdOx1 nCoV-19 (AstraZeneca, 91–159 days after vaccination) (Methods and Extended Data Table 2). In all cases, a substantial loss in neutralizing potency was observed against B.1.1.529 (Fig. 1d and Extended Data Fig. 2b–f). For the two mRNA-based vaccines, BNT162b2 and mRNA-1273, a >21-fold and >8.6-fold decrease in ID50 was observed, respectively. Note that, for these two groups, we specifically chose samples with high titres so that the fold change in titre could be better quantified; the difference in the number of samples with titres above the LOD (6 out of 13 for BNT162b2 versus 11 out of 12 for mRNA-1273) may therefore be favourably biased. Within the Ad26.COV2.S and ChAdOx1 nCOV-19 groups, all of the samples were below the LOD against B.1.1.529, except for two Ad26.COV2.S samples from patients who had a previous history of SARS-CoV-2 infection (Fig. 1d). Collectively, these results suggest that individuals who were previously infected or fully vaccinated remain at risk for B.1.1.529 infection.
Booster shots are now routinely administered in many countries 6 months after full vaccination. We therefore also examined the serum neutralizing activity of individuals who had received three homologous mRNA vaccinations (13 with BNT162b2 and 2 with mRNA-1273, 14–90 days after vaccination). Every sample showed lower activity in neutralizing B.1.1.529, with a mean decrease of 6.5-fold compared with the WT (Fig. 1d). Although all of the samples had titres above the LOD, the substantial loss in activity may still pose a risk for B.1.1.529 infection despite the booster vaccination.
We next confirmed the above findings by testing a subset of the BNT162b2 and mRNA-1273 vaccinee serum samples using authentic SARS-CoV-2 isolates: WT and B.1.1.529. Again, a substantial decrease in the neutralization of B.1.1.529 was observed, with mean decreases of >6.0-fold and >4.1-fold for the fully vaccinated group and the boosted group, respectively (Fig. 1e).
Antibody neutralization of B.1.1.529
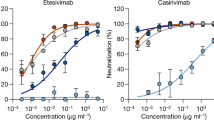
To understand the types of antibodies in the serum that lost neutralizing activity against B.1.1.529, we assessed the neutralization profile of 19 well-characterized monoclonal antibodies against the spike protein, including 17 directed to the RBD and 2 directed to the N-terminal domain (NTD). We included the following monoclonal antibodies that have been authorized or approved for clinical use, either individually or in combination: REGN10987 (imdevimab)12, REGN10933 (casirivimab)12, COV2-2196 (tixagevimab)13, COV2-2130 (cilgavimab)13, LY-CoV555 (bamlanivimab)14, CB6 (etesevimab)15, Brii-196 (amubarvimab)16, Brii-198 (romlusevimab)16 and S309 (sotrovimab)17. We also included other monoclonal antibodies of interest: 910-30 (ref. 18), ADG-2 (ref. 19), DH1047 (ref. 20), S2X259 (ref. 21) and our antibodies 1-20, 2-15, 2-7, 4-18, 5-7 and 10-40 (refs. 22,23,24). The footprints of monoclonal antibodies with structures available were drawn in relation to the mutations found in the B.1.1.529 RBD (Fig. 2a) and NTD (Fig. 2b). The risk to each of the four classes25 of RBD monoclonal antibodies, as well as to the NTD monoclonal antibodies, was immediately apparent. Indeed, neutralization studies on B.1.1.529 pseudovirus showed that 17 out of the 19 monoclonal antibodies tested lost neutralizing activity completely or partially (Fig. 2c and Extended Data Fig. 3). The potency of class 1 and class 2 RBD monoclonal antibodies all decreased by more than 100-fold, as did the more potent monoclonal antibodies in RBD class 3 (REGN10987, COV2-2130 and 2-7). The activities of S309 and Brii-198 were spared. All of the monoclonal antibodies in RBD class 4 lost neutralization potency against B.1.1.529 by at least tenfold, as did monoclonal antibodies directed to the antigenic supersite26 (4-18) or the alternative site23 (5-7) on the NTD. Strikingly, all four combination monoclonal antibody drugs in clinical use lost substantial activity against B.1.1.529, probably abolishing or impairing their efficacy in patients.
a, Footprints of RBD-directed antibodies. Mutations within B.1.1.529 are highlighted in cyan. Approved or authorized antibodies are shown in bold. The receptor-binding motif (RBM) residues are highlighted in yellow. b, Footprints of NTD-directed antibodies. Mutations within B.1.1.529 are highlighted in cyan. The NTD supersite residues are highlighted in light pink. c, Neutralization of D614G and B.1.1.529 pseudoviruses by RBD-directed and NTD-directed monoclonal antibodies (mAbs). d, Neutralization of D614G and B.1.1.529+R346K pseudoviruses by RBD-directed and NTD-directed monoclonal antibodies. Data represent one out of two independent experiments.
Approximately 10% of the B.1.1.529 viruses in the Global Initiative on Sharing All Influenza Data (GISAID)1 also contain an additional RBD mutation, R346K, which is the defining mutation for the B.1.621/Mu variant27. We therefore constructed another pseudovirus (B.1.1.529+R346K) containing this mutation for additional testing using the same panel of monoclonal antibodies (Fig. 2d). The overall findings resembled those already shown in Fig. 2c, with the exception that the neutralizing activity of Brii-198 was abolished. In fact, nearly the entire panel of antibodies was essentially rendered inactive against this minor form of the Omicron variant.
The fold changes in IC50 of the monoclonal antibodies against B.1.1.529 and B.1.1.529+R346K relative to D614G are summarized in the first two rows of Fig. 3a. The considerable loss of activity observed for all classes of monoclonal antibodies against B.1.1.529 suggests that perhaps the same is occurring in the serum of convalescent patients and vaccinated individuals.
a, Neutralization of pseudoviruses containing single mutations found within B.1.1.529 by a panel of 19 monoclonal antibodies. The fold change relative to neutralization of D614G is denoted, with resistance coloured red and sensitization coloured green. b, Modelling of critical mutations in B.1.1.529 that affect antibody neutralization.
Mutations conferring antibody resistance
To understand the specific B.1.1.529 mutations that confer antibody resistance, we next individually tested the same panel of 19 monoclonal antibodies against pseudoviruses for each of the 34 mutations (excluding D614G) found in B.1.1.529 or B.1.1.529+R346K. Our findings not only confirmed the role of known mutations at spike residues 142–145, 417, 484 and 501 in conferring resistance to NTD or RBD (class 1 or class 2) antibodies4 but also revealed several mutations that were not previously known to have functional importance to neutralization (Fig. 3a and Extended Data Fig. 4). Q493R, which was previously shown to affect binding of CB6 and LY-CoV555 (ref. 28) as well as polyclonal sera29, mediated resistance to CB6 (class 1) as well as to LY-CoV555 and 2-15 (class 2), findings that could be explained by the abolishment of hydrogen bonds due to the long side chain of arginine and induced steric clashes with CDRH3 in these antibodies (Fig. 3b (left)). Both N440K and G446S mediated resistance to REGN10987 and 2-7 (class 3), observations that could also be explained by steric hindrance (Fig. 3b (middle)). The most striking and perhaps unexpected finding was that S371L broadly affected neutralization by monoclonal antibodies in all four RBD classes (Fig. 3a and Extended Data Fig. 4). Although the precise mechanism of this resistance is unknown, in silico modelling suggested two possibilities (Fig. 3b (right)). First, in the RBD-down state, mutating Ser to Leu results in an interference with the N343 glycan, thereby possibly alteringits conformation and affecting class 3 antibodies that typically bind to this region. Second, in the RBD-up state, S371L may alter the local conformation of the loop consisting of Ser371-Ser373-Ser375, thereby affecting the binding of class 4 antibodies that generally target a portion of this loop24. It is not clear how class 1 and class 2 RBD monoclonal antibodies are affected by this mutation.
Evolution of SARS-CoV-2 to antibodies
To gain insights into the antibody resistance of B.1.1.529 relative to previous SARS-CoV-2 variants, we evaluated the neutralizing activity of the same panel of neutralizing monoclonal antibodies against pseudoviruses for B.1.1.7 (ref. 8), B.1.526 (ref. 30), B.1.429 (ref. 31), B.1.617.2 (ref. 9), P.1 (ref. 32) and B.1.351 (ref. 33). It is evident from these results (Fig. 4 and Extended Data Fig. 5) that previous variants developed resistance to only NTD antibodies and class 1 and class 2 RBD antibodies. Here B.1.1.529, with or without R346K, has made a big mutational leap by becoming not only nearly completely resistant to class 1 and class 2 RBD antibodies, but also substantially resistant to both class 3 and class 4 RBD antibodies. B.1.1.529 is now the most complete ‘escapee’ from neutralization by currently available antibodies.
Discussion
The Omicron variant caused fear almost as soon as it was detected to be spreading in South Africa. The suggestion that this new variant would transmit more readily has come true in the ensuing weeks2. The extensive mutations found in its spike protein raised concerns that the efficacy of current COVID-19 vaccines and antibody therapies might be compromised. Indeed, in this study, sera from convalescent patients (Fig. 1c) and vaccinees (Fig. 1d, e) showed markedly reduced neutralizing activity against B.1.1.529. Other studies have found similar losses34,35,36,37,38. These findings are consistent with emerging clinical data on the Omicron variant demonstrating higher rates of reinfection11 and vaccine breakthroughs. In fact, recent reports showed that the efficacy of two doses of BNT162b2 vaccine has dropped from over 90% against the original SARS-CoV-2 strain to approximately 40% and 33% against B.1.1.529 in the United Kingdom39 and South Africa40, respectively. Even a third booster shot may not adequately protect against Omicron infection39,41, although the protection against disease still makes it advisable to administer booster vaccinations. Vaccines that elicited lower neutralizing titres35,42 are expected to fare worse against B.1.1.529.
The nature of the loss in serum neutralizing activity against B.1.1.529 could be discerned from our findings on a panel of monoclonal antibodies directed to the viral spike. The neutralizing activities of all four major classes of RBD monoclonal antibodies and two distinct classes of NTD monoclonal antibodies are either abolished or impaired (Fig. 2c, d). In addition to previously identified mutations that confer antibody resistance4, we have uncovered four previously undescribed spike mutations with functional consequences. Q493R confers resistance to some class 1 and class 2 RBD monoclonal antibodies; N440K and G446S confer resistance to some class 3 RBD monoclonal antibodies; and S371L confers global resistance to many RBD monoclonal antibodies through mechanisms that are not yet apparent. While performing these monoclonal antibody studies, we also observed that nearly all of the currently authorized or approved monoclonal antibody drugs are rendered weak or inactive by B.1.1.529 (Figs. 2c and 3a). In fact, the Omicron variant that contains R346K renders almost all current antibody therapy for COVID-19 ineffective (Figs. 2d and 3a).
The scientific community has chased after SARS-CoV-2 variants for a year. As more and more of them appeared, our interventions directed to the spike became increasingly ineffective. The Omicron variant has now put an exclamation mark on this point. It is not too farfetched to think that this SARS-CoV-2 is now only a mutation or two away from being pan-resistant to current antibodies, either monoclonal or polyclonal. We must devise strategies that anticipate the evolutional direction of the virus and develop agents that target better-conserved viral elements.
Methods
Data reporting
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Serum samples
Convalescent plasma samples were obtained from patients with documented SARS-CoV-2 infection. These samples were collected at the beginning of the pandemic in early 2020 at Columbia University Irving Medical Center and are therefore assumed to be infection by the WT strain of SARS-CoV-2 (ref. 4). Sera from individuals who received two or three doses of mRNA-1273 or BNT162b2 vaccine were collected at Columbia University Irving Medical Center at least two weeks after the final dose. Sera from individuals who received one dose of Ad26.COV2.S or two doses of ChAdOx1 nCov-19 were obtained from BEI Resources. Some individuals were also infected by SARS-CoV-2 in addition to the vaccinations they received. Note that, whenever possible, we specifically chose samples with high titres against the WT strain of SARS-CoV-2 such that the loss in activity against B.1.1.529 could be better quantified and the titres observed here should therefore be considered in that context. All collections were conducted under protocols reviewed and approved by the Institutional Review Board of Columbia University. All of the participants provided written informed consent. Additional information for the convalescent samples and vaccinee samples is provided in Extended Data Tables 1 and 2, respectively.
Monoclonal antibodies
Antibodies were expressed as previously described22 by synthesis of heavy chain variable (VH) and light chain variable (VL) genes (GenScript), transfection of Expi293 cells (Thermo Fisher Scientific) and affinity purification from the supernatant by rProtein A Sepharose (GE). REGN10987, REGN10933, COV2-2196 and COV2-2130 were provided by Regeneron Pharmaceuticals; Brii-196 and Brii-198 were provided by Brii Biosciences; CB6 was provided by B. Zhang and P. Kwong (NIH); and 910-30 was provided by B. DeKosky (MIT).
Cell lines
Expi293 cells were obtained from Thermo Fisher Scientific (A14527); Vero E6 cells were obtained from the ATCC (CRL-1586); HEK293T cells were obtained from the ATCC (CRL-3216); and Vero-E6-TMPRSS2 cells were obtained from JCRB (JCRB1819). Cells were purchased from authenticated vendors and morphology was confirmed visually before use. All cell lines tested mycoplasma negative.
Variant SARS-CoV-2 spike plasmid construction
An in-house high-throughput template-guide gene synthesis approach was used to generate spike genes with single or full mutations of B.1.1.529. In brief, 5′-phosphorylated oligos with designed mutations were annealed to the reverse strand of the WT spike gene construct and extended by DNA polymerase. Extension products (forward-stranded fragments) were then ligated together by Taq DNA ligase and subsequently amplified by PCR to generate variants of interest. To verify the sequences of variants, next-generation sequencing libraries were prepared according to a low-volume Nextera sequencing protocol43 and sequenced on the Illumina Miseq platform (single-end mode with 50 bp R1). Raw reads were processed using Cutadapt v.2.144 with the default settings to remove adapters, and were then aligned to reference variants sequences using Bowtie2 v.2.3.445 with the default settings. The resulting reads alignments were then visualized in the Integrative Genomics Viewer46 and manually inspected to verify the fidelity of variants. The sequences of the oligos used in variants generation are provided in Extended Data Table 3.
Pseudovirus production
Pseudoviruses were produced in the vesicular stomatitis virus (VSV) background, in which the native glycoprotein was replaced by that of SARS-CoV-2 and its variants, as previously described24. In brief, HEK293T cells were transfected with a spike expression construct with polyethylenimine (PEI) (1 mg ml−1) and cultured overnight at 37 °C under 5% CO2, and then infected with VSV-G pseudotyped ΔG-luciferase (G*ΔG-luciferase, Kerafast) 1 day after transfection. After 2 h of infection, cells were washed three times, changed to fresh medium and then cultured for approximately another 24 h before the supernatants were collected, centrifuged and aliquoted to use in assays.
Pseudovirus neutralization assay
All viruses were first titrated to normalize the viral input between assays. Heat-inactivated sera or antibodies were first serially diluted in 96 well-plates in triplicate, starting at 1:100 dilution for sera and 10 µg ml−1 for antibodies. Viruses were then added and the virus–sample mixture was incubated at 37 °C for 1 h. Vero-E6 cells (ATCC) were then added at a density of 3 × 104 cells per well and the plates were incubated at 37 °C for approximately 10 h. Luciferase activity was quantified using the Luciferase Assay System (Promega) according to the manufacturer’s instructions using SoftMax Pro v.7.0.2 (Molecular Devices). Neutralization curves and IC50 values were derived by fitting a nonlinear five-parameter dose–response curve to the data in GraphPad Prism v.9.2.
Authentic virus isolation and propagation
Authentic B.1.1.529 was isolated from a specimen from the respiratory tract of a patient with COVID-19 in Hong Kong by K.-Y. Yuen and colleagues at the Department of Microbiology, The University of Hong Kong. Isolation of WT SARS-CoV-2 was previously described47. Viruses were propagated in Vero-E6-TMPRSS2 cells and the sequence was confirmed by next-generation sequencing before use.
Authentic virus neutralization assay
To measure neutralization of authentic SARS-CoV-2 viruses, Vero-E6-TMPRSS2 cells were first seeded in 96-well plates in cell culture medium (Dulbecco’s Modified Eagle Medium (DMEM) + 10% fetal bovine serum (FBS) + 1% penicillin–streptomycin) overnight at 37 °C under 5% CO2 to establish a monolayer. The next day, sera or antibodies were serially diluted in 96-well plates in triplicate in DMEM + 2% FBS and then incubated with 0.01 multiplicity of infection of WT SARS-CoV-2 or B.1.1.529 at 37 °C for 1 h. Sera were diluted from 1:100 dilution and antibodies were diluted from 10 µg ml−1. Next, the mixture was overlaid onto cells and further incubated at 37 °C under 5% CO2 for approximately 72 h. Cytopathic effects were then scored by plaque assay in a blinded manner. Neutralization curves and IC50 values were derived by fitting a nonlinear five-parameter dose–response curve to the data in GraphPad Prism v.9.2.
Antibody footprint analysis and RBD mutagenesis analysis
The SARS-CoV-2 spike structure used for displaying epitope footprints and mutations within emerging strains was downloaded from the Protein Data Bank (PDB: 6ZGE). The structures of antibody-spike complexes were also obtained from PDB (7L5B (2-15), 6XDG (REGN10933 and REGN10987), 7L2E (4-18), 7RW2 (5–7), 7C01 (CB6), 7KMG (LY-COV555), 7CDI (Brii-196), 7KS9 (910-30), 7LD1 (DH1047), 7RAL (S2X259), 7LSS (2–7) and 6WPT (S309)). Interface residues were identified using PISA48 using the default parameters. The footprint for each antibody was defined by the boundaries of all epitope residues. The border for each footprint was then optimized by ImageMagick v.7.0.10-31 (https://imagemagick.org). PyMOL v.2.3.2 was used to perform mutagenesis and to generate structural plots (Schrödinger).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Materials used in this study will be made available under an appropriate Materials Transfer Agreement. All the data are provided in the paper. The structures used for analysis in this study are available from PDB under accession numbers 6ZGE, 7L5B, 6XDG, 7L2E, 7RW2, 7C01, 7KMG, 7CDI, 7KS9, 7LD1, 7RAL, 7LSS and 6WPT.
References
Shu, Y. & McCauley, J. GISAID: global initiative on sharing all influenza data—from vision to reality. Euro Surveill. 22, 30494 (2017).
Grabowski, F., Kochańczyk, M. & Lipniacki, T. Omicron strain spreads with the doubling time of 3.2—3.6 days in South Africa province of Gauteng that achieved herd immunity to Delta variant. Preprint at https://doi.org/10.1101/2021.12.08.21267494 (2021).
Scott, L. et al. Track Omicron’s spread with molecular data. Science 374, 1454–1455 (2021).
Wang, P. et al. Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7. Nature 593, 130–135 (2021).
Abu-Raddad, L. J., Chemaitelly, H. & Butt, A. A. Effectiveness of the BNT162b2 COVID-19 vaccine against the B.1.1.7 and B.1.351 variants. N. Engl. J. Med. 385, 187–189 (2021).
Madhi, S. A. et al. Efficacy of the ChAdOx1 nCoV-19 COVID-19 vaccine against the B.1.351 variant. N. Engl. J. Med. 384, 1885–1898 (2021).
Sadoff, J. et al. Safety and efficacy of single-dose Ad26.COV2.S vaccine against COVID-19. N. Engl. J. Med. 384, 2187–2201 (2021).
Davies, N. G. et al. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science 372, eabg3055 (2021).
Mlcochova, P. et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 599, 114–119 (2021).
Hadfield, J. et al. Nextstrain: real-time tracking of pathogen evolution. Bioinformatics 34, 4121–4123 (2018).
Pulliam, J. R. C. et al. Increased risk of SARS-CoV-2 reinfection associated with emergence of the Omicron variant in South Africa. Preprint at https://doi.org/10.1101/2021.11.11.21266068 (2021).
Hansen, J. et al. Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail. Science 369, 1010–1014 (2020).
Zost, S. J. et al. Potently neutralizing and protective human antibodies against SARS-CoV-2. Nature 584, 443–449 (2020).
Jones, B. E. et al. The neutralizing antibody, LY-CoV555, protects against SARS-CoV-2 infection in nonhuman primates. Sci. Transl. Med. 13, eabf1906 (2021).
Shi, R. et al. A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2. Nature 584, 120–124 (2020).
Ju, B. et al. Human neutralizing antibodies elicited by SARS-CoV-2 infection. Nature 584, 115–119 (2020).
Pinto, D. et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature 583, 290–295 (2020).
Banach, B. B. et al. Paired heavy- and light-chain signatures contribute to potent SARS-CoV-2 neutralization in public antibody responses. Cell Rep 37, 109771 (2021).
Rappazzo, C. G. et al. Broad and potent activity against SARS-like viruses by an engineered human monoclonal antibody. Science 371, 823–829 (2021).
Li, D. et al. In vitro and in vivo functions of SARS-CoV-2 infection-enhancing and neutralizing antibodies. Cell 184, 4203–4219 (2021).
Tortorici, M. A. et al. Broad sarbecovirus neutralization by a human monoclonal antibody. Nature 597, 103–108 (2021).
Liu, L. et al. Potent neutralizing antibodies against multiple epitopes on SARS-CoV-2 spike. Nature 584, 450–456 (2020).
Cerutti, G. et al. Neutralizing antibody 5-7 defines a distinct site of vulnerability in SARS-CoV-2 spike N-terminal domain. Cell Rep. 37, 109928 (2021).
Liu, L. et al. Isolation and comparative analysis of antibodies that broadly neutralize sarbecoviruses. Preprint at https://doi.org/10.1101/2021.12.11.472236 (2021).
Barnes, C. O. et al. SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature 588, 682–687 (2020).
Cerutti, G. et al. Potent SARS-CoV-2 neutralizing antibodies directed against spike N-terminal domain target a single supersite. Cell Host Microbe 29, 819–833 (2021).
Uriu, K. et al. Neutralization of the SARS-CoV-2 Mu variant by convalescent and vaccine serum. N. Engl. J. Med. 385, 2397–2399 (2021).
Starr, T. N., Greaney, A. J., Dingens, A. S. & Bloom, J. D. Complete map of SARS-CoV-2 RBD mutations that escape the monoclonal antibody LY-CoV555 and its cocktail with LY-CoV016. Cell Rep. Med. 2, 100255 (2021).
Amanat, F. et al. SARS-CoV-2 mRNA vaccination induces functionally diverse antibodies to NTD, RBD, and S2. Cell 184, 3936–3948 (2021).
Annavajhala, M. K. et al. Emergence and expansion of SARS-CoV-2 B.1.526 after identification in New York. Nature 597, 703–708 (2021).
Zhang, W. et al. Emergence of a novel SARS-CoV-2 variant in southern California. JAMA 325, 1324–1326 (2021).
Faria, N. R. et al. Genomics and epidemiology of the P.1 SARS-CoV-2 lineage in Manaus, Brazil. Science 372, 815–821 (2021).
Tegally, H. et al. Detection of a SARS-CoV-2 variant of concern in South Africa. Nature 592, 438–443 (2021).
Schmidt, F. et al. Plasma neutralization of the SARS-CoV-2 Omicron variant. New Engl. J. Med. https://doi.org/10.1056/NEJMc2119641 (2021).
Garcia-Beltran, W. F. et al. mRNA-based COVID-19 vaccine boosters induce neutralizing immunity against SARS-CoV-2 Omicron variant. Cell, https://doi.org/10.1016/j.cell.2021.12.033 (2022).
Cameroni, E. et al. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature https://doi.org/10.1038/s41586-021-04386-2 (2021).
Doria-Rose, N. A. et al. Booster of mRNA-1273 vaccine reduces SARS-CoV-2 Omicron escape from neutralizing antibodies. Preprint at https://doi.org/10.1101/2021.12.15.21267805 (2021).
Planas, D. et al. Considerable escape of SARS-CoV-2 variant Omicron to antibody neutralization. Nature https://doi.org/10.1038/s41586-021-04389-z (2021).
Andrews, N. et al. Effectiveness of COVID-19 vaccines against the Omicron (B.1.1.529) variant of concern. Preprint at https://doi.org/10.1101/2021.12.14.21267615 (2021).
Wroughton, L. Omicron variant more resistant to vaccine but causes less severe covid, major South African study concludes. The Washington Post (14 December 2021).
Kuhlmann, C. et al. Breakthrough infections with SARS-CoV-2 Omicron variant despite booster dose of mRNA vaccine. SSRN https://doi.org/10.2139/ssrn.3981711 (2021).
Cromer, D. et al. Neutralising antibody titres as predictors of protection against SARS-CoV-2 variants and the impact of boosting: a meta-analysis. Lancet Microbe 3, e52–e61 (2021).
Baym, M. et al. Inexpensive multiplexed library preparation for megabase-sized genomes. PLoS ONE 10, e0128036 (2015).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Chu, H. et al. Comparative tropism, replication kinetics, and cell damage profiling of SARS-CoV-2 and SARS-CoV with implications for clinical manifestations, transmissibility, and laboratory studies of COVID-19: an observational study. Lancet Microbe 1, e14–e23 (2020).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Acknowledgements
We thank the staff at Regeneron Pharmaceuticals, B. Zhang and P. Kwong (NIAID), and B. Dekosky (MIT) for antibodies. This study was supported by funding from the Gates Foundation, the JPB Foundation, A. and P. Cherng, S. Yin, C. Ludwig, D. and R. Wu, Health@InnoHK and the National Science Foundation (MCB-2032259).
Author information
Authors and Affiliations
Contributions
D.D.H. conceived this project. Lihong Liu, S.I. and M.W. conducted pseudovirus neutralization experiments. J.F.-W.C., H. Chu, K.K.-H.C., T.T.-T.Y., C.Y., K.K.-W.T. and H. Chen conducted authentic virus neutralization experiments. Y.G. and Z.S. conducted bioinformatic analyses. Liyuan Liu and Yiming Huang constructed the spike expression plasmids. Y.L. managed the project. J.Y. expressed and purified antibodies. M.T.Y. and M.E.S. provided clinical samples. M.S.N. and Yaoxing Huang contributed to discussions. H.H.W., K.-Y.Y. and D.D.H. directed and supervised the project. Lihong Liu, S.I. and D.D.H. analysed the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Lihong Liu, S.I., M.S.N., J.Y., Yaoxing Huang and D.D.H. are inventors on patent applications (WO2021236998) or provisional patent applications (63/271,627) filed by Columbia University for a number of SARS-CoV-2 neutralizing antibodies described in this manuscript. Both sets of applications are under review. D.D.H. is a co-founder of TaiMed Biologics and RenBio, consultant to WuXi Biologics and Brii Biosciences, and board director for Vicarious Surgical.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Mutations within B.1.1.529 denoted on the full SARS-CoV-2 spike trimer.
The SARS-CoV-2 spike structure was downloaded from PDB 6ZGE.
Extended Data Fig. 2 Individual neutralization curves for pseudovirus neutralization assays by serum.
Neutralization by a, convalescent sera. b, Pfizer (BNT162b2) vaccinee sera. c, Moderna (mRNA-1273) vaccinee sera. d, J&J (Ad26.COV2.S) vaccinee sera. e, AstraZeneca (ChAdOx1 nCoV-19) vaccinee sera. f, boosted (three homologous BNT162b2 or mRNA-1273 vaccinations) vaccinee sera. Error bars denote mean ± standard error of the mean (SEM) for three technical replicates.
Extended Data Fig. 3 Individual neutralization curves for pseudovirus neutralization assays by monoclonal antibodies.
Error bars denote mean ± standard error of the mean (SEM) for three technical replicates.
Extended Data Fig. 4 Individual neutralization curves for pseudovirus neutralization assays by monoclonal antibodies against individual SARS-CoV-2 mutations.
Error bars denote mean ± standard error of the mean (SEM) for three technical replicates.
Extended Data Fig. 5 Individual neutralization curves for pseudovirus neutralization assays by monoclonal antibodies against SARS-CoV-2 variants.
Error bars denote mean ± standard error of the mean (SEM) for three technical replicates.
Supplementary information
Rights and permissions
About this article
Cite this article
Liu, L., Iketani, S., Guo, Y. et al. Striking antibody evasion manifested by the Omicron variant of SARS-CoV-2. Nature 602, 676–681 (2022). https://doi.org/10.1038/s41586-021-04388-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04388-0
This article is cited by
-
Computational design and engineering of self-assembling multivalent microproteins with therapeutic potential against SARS-CoV-2
Journal of Nanobiotechnology (2024)
-
Single-domain antibodies against SARS-CoV-2 RBD from a two-stage phage screening of universal and focused synthetic libraries
BMC Infectious Diseases (2024)
-
Potent and broadly neutralizing antibodies against sarbecoviruses induced by sequential COVID-19 vaccination
Cell Discovery (2024)
-
COVID-19 drug discovery and treatment options
Nature Reviews Microbiology (2024)
-
SARS-CoV-2-specific cellular and humoral immunity after bivalent BA.4/5 COVID-19-vaccination in previously infected and non-infected individuals
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.