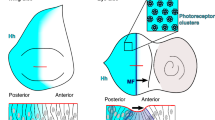
Abstract
Morphogen gradients are fundamental to establish morphological patterns in developing tissues1. During development, gradients scale to remain proportional to the size of growing organs2,3. Scaling is a universal gear that adjusts patterns to size in living organisms3,4,5,6,7,8, but its mechanisms remain unclear. Here, focusing on the Decapentaplegic (Dpp) gradient in the Drosophila wing disc, we uncover a cell biological basis behind scaling. From small to large discs, scaling of the Dpp gradient is achieved by increasing the contribution of the internalized Dpp molecules to Dpp transport: to expand the gradient, endocytosed molecules are re-exocytosed to spread extracellularly. To regulate the contribution of endocytosed Dpp to the spreading extracellular pool during tissue growth, it is the Dpp binding rates that are progressively modulated by the extracellular factor Pentagone, which drives scaling. Thus, for some morphogens, evolution may act on endocytic trafficking to regulate the range of the gradient and its scaling, which could allow the adaptation of shape and pattern to different sizes of organs in different species.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Source data are provided with this paper. Datasets generated during the parameter estimation are available in GitHub (https://github.com/zenah12/DppTrafficking-/blob/main/README.md). Source data are provided with this paper.
Code availability
Source codes are available in GitHub (https://github.com/zenah12/DppTrafficking-/blob/main/README.md). MATLAB code corresponding to the binning of control and pent mutant data in Fig. 2a is available upon request.
References
Rogers, K. W. & Schier, A. F. Morphogen gradients: from generation to interpretation. Annu. Rev. Cell Dev. Biol. 27, 377–407 (2011).
Wartlick, O., Julicher, F. & Gonzalez-Gaitan, M. Growth control by a moving morphogen gradient during Drosophila eye development. Development 141, 1884–1893 (2014).
Wartlick, O. et al. Dynamics of Dpp signaling and proliferation control. Science 331, 1154–1159 (2011).
Averbukh, I., Ben-Zvi, D., Mishra, S. & Barkai, N. Scaling morphogen gradients during tissue growth by a cell division rule. Development 141, 2150–2156 (2014).
Ben-Zvi, D., Pyrowolakis, G., Barkai, N. & Shilo, B. Z. Expansion-repression mechanism for scaling the Dpp activation gradient in Drosophila wing imaginal discs. Curr. Biol. 21, 1391–1396 (2011).
Fried, P. & Iber, D. Dynamic scaling of morphogen gradients on growing domains. Nat. Commun. 5, 5077 (2014).
Hamaratoglu, F., de Lachapelle, A. M., Pyrowolakis, G., Bergmann, S. & Affolter, M. Dpp signaling activity requires Pentagone to scale with tissue size in the growing Drosophila wing imaginal disc. PLoS Biol. 9, e1001182 (2011).
Zhu, Y., Qiu, Y., Chen, W., Nie, Q. & Lander, A. D. Scaling a Dpp morphogen gradient through feedback control of receptors and co-receptors. Dev. Cell 53, 724–739 (2020).
Inomata, H. Scaling of pattern formations and morphogen gradients. Dev. Growth Differ. 59, 41–51 (2017).
Romanova-Michaelides, M., Aguilar-Hidalgo, D., Julicher, F. & Gonzalez-Gaitan, M. The wing and the eye: a parsimonious theory for scaling and growth control? Wiley Interdiscip. Rev. Dev. Biol. 4, 591–608 (2015).
Entchev, E. V., Schwabedissen, A. & Gonzalez-Gaitan, M. Gradient formation of the TGF-β homolog Dpp. Cell 103, 981–991 (2000).
Kicheva, A. et al. Kinetics of morphogen gradient formation. Science 315, 521–525 (2007).
Stapornwongkul, K. S., de Gennes, M., Cocconi, L., Salbreux, G. & Vincent, J. P. Patterning and growth control in vivo by an engineered GFP gradient. Science 370, 321–327 (2020).
Akiyama, T. & Gibson, M. C. Decapentaplegic and growth control in the developing Drosophila wing. Nature 527, 375–378 (2015).
Matsuda, S. et al. Asymmetric requirement of Dpp/BMP morphogen dispersal in the Drosophila wing disc. Nat. Commun. 12, 6435 (2021).
Posakony, L. G., Raftery, L. A. & Gelbart, W. M. Wing formation in Drosophila melanogaster requires decapentaplegic gene function along the anterior–posterior compartment boundary. Mech. Dev. 33, 69–82 (1990).
Aguirre-Tamaral, A. & Guerrero, I. Improving the understanding of cytoneme-mediated morphogen gradients by in silico modeling. PLoS Comput. Biol. 17, e1009245 (2021).
Rojas-Rios, P., Guerrero, I. & Gonzalez-Reyes, A. Cytoneme-mediated delivery of hedgehog regulates the expression of bone morphogenetic proteins to maintain germline stem cells in Drosophila. PLoS Biol. 10, e1001298 (2012).
Roy, S., Huang, H., Liu, S. & Kornberg, T. B. Cytoneme-mediated contact-dependent transport of the Drosophila decapentaplegic signaling protein. Science 343, 1244624 (2014).
Aguilar Hidalgo, D., Hadjivasilou, Z., Romanova-Michaelides, M., González-Gaitán, M. & Jülicher, F. Dynamic modes of morphogen transport. Preprint at https://arxiv.org/abs/1909.13280 (2019).
Wolpert, M. K. A. L. Mechanisms for positional signalling by morphogen transport: a theoretical study. J. Theor. Biol. 191, 103–114 (1998).
Harmansa, S., Hamaratoglu, F., Affolter, M. & Caussinus, E. Dpp spreading is required for medial but not for lateral wing disc growth. Nature 527, 317–322 (2015).
Felder, S., LaVin, J., Ullrich, A. & Schlessinger, J. Kinetics of binding, endocytosis, and recycling of EGF receptor mutants. J. Cell Biol. 117, 203–212 (1992).
Hatta, T. et al. Identification of the ligand-binding site of the BMP type IA receptor for BMP-4. Biopolymers 55, 399–406 (2000).
Kirchhausen, T., Owen, D. & Harrison, S. C. Molecular structure, function, and dynamics of clathrin-mediated membrane traffic. Cold Spring Harb. Perspect. Biol. 6, a016725 (2014).
Kural, C. et al. Dynamics of intracellular clathrin/AP1- and clathrin/AP3-containing carriers. Cell Rep. 2, 1111–1119 (2012).
Mineur, P. et al. Newly identified biologically active and proteolysis-resistant VEGF-A isoform VEGF111 is induced by genotoxic agents. J. Cell Biol. 179, 1261–1273 (2007).
Sigismund, S. et al. Clathrin-mediated internalization is essential for sustained EGFR signaling but dispensable for degradation. Dev. Cell 15, 209–219 (2008).
Wang, Y., Wang, X., Wohland, T. & Sampath, K. Extracellular interactions and ligand degradation shape the nodal morphogen gradient. Elife 5, e13879 (2016).
Waters, C. M., Oberg, K. C., Carpenter, G. & Overholser, K. A. Rate constants for binding, dissociation, and internalization of EGF: effect of receptor occupancy and ligand concentration. Biochemistry 29, 3563–3569 (1990).
Allendorph, G. P., Isaacs, M. J., Kawakami, Y., Izpisua Belmonte, J. C. & Choe, S. BMP-3 and BMP-6 structures illuminate the nature of binding specificity with receptors. Biochemistry 46, 12238–12247 (2007).
Schwank, G. et al. Formation of the long range Dpp morphogen gradient. PLoS Biol. 9, e1001111 (2011).
Hemalatha, A., Prabhakara, C. & Mayor, S. Endocytosis of Wingless via a dynamin-independent pathway is necessary for signaling in Drosophila wing discs. Proc. Natl Acad. Sci. USA 113, E6993–E7002 (2016).
Zhou, S. et al. Free extracellular diffusion creates the Dpp morphogen gradient of the Drosophila wing disc. Curr. Biol. 22, 668–675 (2012).
Khmelinskii, A. et al. Tandem fluorescent protein timers for in vivo analysis of protein dynamics. Nat. Biotechnol. 30, 708–714 (2012).
Shcherbo, D. et al. Far-red fluorescent tags for protein imaging in living tissues. Biochem. J. 418, 567–574 (2009).
Zerial, M. & McBride, H. Rab proteins as membrane organizers. Nat. Rev. Mol. Cell. Biol. 2, 107–117 (2001).
Vuilleumier, R. et al. Control of Dpp morphogen signalling by a secreted feedback regulator. Nat. Cell Biol. 12, 611–617 (2010).
Fujise, M. et al. Dally regulates Dpp morphogen gradient formation in the Drosophila wing. Development 130, 1515–1522 (2003).
Akiyama, T. et al. Dally regulates Dpp morphogen gradient formation by stabilizing Dpp on the cell surface. Dev. Biol. 313, 408–419 (2008).
Guha, A., Sriram, V., Krishnan, K. S. & Mayor, S. Shibire mutations reveal distinct dynamin-independent and -dependent endocytic pathways in primary cultures of Drosophila hemocytes. J. Cell Sci. 116, 3373–3386 (2003).
Gupta, G. D. et al. Analysis of endocytic pathways in Drosophila cells reveals a conserved role for GBF1 in internalization via GEECs. PLoS ONE 4, e6768 (2009).
Mayor, S., Parton, R. G. & Donaldson, J. G. Clathrin-independent pathways of endocytosis. Cold Spring Harb. Perspect. Biol. 6, a016758 (2014).
Sabharanjak, S., Sharma, P., Parton, R. G. & Mayor, S. GPI-anchored proteins are delivered to recycling endosomes via a distinct cdc42-regulated, clathrin-independent pinocytic pathway. Dev. Cell 2, 411–423 (2002).
Norman, M., Vuilleumier, R., Springhorn, A., Gawlik, J. & Pyrowolakis, G. Pentagone internalises glypicans to fine-tune multiple signalling pathways. Elife 5, e13301 (2016).
Yan, D. & Lin, X. Shaping morphogen gradients by proteoglycans. Cold Spring Harb. Perspect. Biol. 1, a002493 (2009).
Gratz, S. J. et al. Highly specific and efficient CRISPR/Cas9-catalyzed homology- directed repair in Drosophila. Genetics 196, 961–971 (2014).
Gratz, S. J. et al. Genome engineering of Drosophila with the CRISPR RNA-guided Cas9 nuclease. Genetics 194, 1029–1035 (2013).
Derivery, E. et al. Polarized endosome dynamics by spindle asymmetry during asymmetric cell division. Nature 528, 280–285 (2015).
Tanimoto, H., Itoh, S., ten Dijke, P. & Tabata, T. Hedgehog creates a gradient of DPP activity in Drosophila wing imaginal discs. Mol. Cell 5, 59–71 (2000).
Eugster, C., Panakova, D., Mahmoud, A. & Eaton, S. Lipoprotein–heparan sulfate interactions in the Hh pathway. Dev. Cell 13, 57–71 (2007).
Marois, E., Mahmoud, A. & Eaton, S. The endocytic pathway and formation of the Wingless morphogen gradient. Development 133, 307–317 (2006).
Loubery, S. & Gonzalez-Gaitan, M. Monitoring notch/delta endosomal trafficking and signaling in Drosophila. Methods Enzymol. 534, 301–321 (2014).
Montagne, C. & Gonzalez-Gaitan, M. Sara endosomes and the asymmetric division of intestinal stem cells. Development 141, 2014–2023 (2014).
Acknowledgements
We thank R. Mateus and I. Castanon Ortega for their feedback on the manuscript; F. Karch for Cre recombinase; K. Basler for the stock to make brkM68 tkv8 mutant clones; E. Doná for the p744_p3E_mkate2sfGFP plasmid; M. Affolter for the LexA inducible eGFP::DPP; K. Kruse for discussions; E. Derivery for the purified sfGFP-mKate2; V. Rasul-Kareeva for various contributions; and the Bioimaging Center of University of Geneva for microscopy support. Z.H. was supported by an HFSP long-term fellowship; and M.G.-G. was supported by the DIP of the Canton of Geneva, SNSF, the SystemsX EpiPhysX grant, the ERC (Sara and Morphogen) and the NCCR Chemical Biology program.
Author information
Authors and Affiliations
Contributions
M.R.-M. performed most experiments and quantifications and performed data analysis. D.A.-H., Z.H. and F.J. developed the theory and performed data analysis. Z.H. and D.A.-H. performed numerical simulations. C.S. cloned and made fly stocks to express DppTimer and UAS Pentagone::GFP, performed immunoprecipitations and purified the GBP. D.B. performed the photoconversion experiment, labelled the purified GBP with Alexa555, cloned and purified GBP–Dendra2 and developed the acid wash. M.D. made the DppCRISPR stock. M.R.-M. and M.G.-G. conceived and designed experiments. M.R.-M. and Z.H. prepared figures. M.R.-M., Z.H., F.J. and M.G.-G. prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Dagmar Iber, Timothy Saunders and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Photoconversion assay controls and two extreme regimes of Dpp transport.
a–d, Photoconversion assay. Test of efficiency of the acid wash in the photoconversion experiment using GBP-Dendra2: GFP-Dpp expressing discs have been incubated in GBP-Dendra2 for 50 min at 4 °C (the nanobody is only bound to the extracellular pool) and subsequently acid-washed to remove the label of the extracellular pool. Confocal image of eGFP-DppLOP expressing disc (a) and corresponding images of Dendra2* (b) before photoconversion (left) and 40 min following photoconversion (right; see Materials and Methods). Note that no detectable Dendra2* signal is observed 40 min after the acid wash, indicating that the extracellular pool of nanobodies has been efficiently removed and that the potential extracellular leftover (below the detection limit) cannot lead to an observable recovery in intracellular compartments. c, Comparison between eGFP-DppGal4 gradient profiles and gradient profiles formed by photoconverted Dendra2* propagated into the posterior compartment of the discs (photoconversion experiments as in Fig. 1a). Bar plot showing ϕ=λ/l of eGFP-DppGal4 gradient profiles and photoconverted Dendra2* gradient profiles for large discs. Bars, standard deviations. Two-tailed two sample t-test, p-value = 0.2353. d, Fluorescence intensity of Dendra2* in a ROI of 6x35 µm at the source boundary in the photoconversion experiment in Fig. 1a. Measured Dendra2* fluorescence (blue dots) is plotted as a function of time after the photoconversion event. The red line represents the theoretical dynamics of Dendra2* fluorescence signal considering the parameterized values for large discs. n = 4 biologically independent samples. Data represented as mean values ± s.e.m. e–h, Acid wash efficiently removes the extracellular pool. Confocal images of eGFP-DppLOP gradient (green in e, f), and extracellular eGFP-DppLOP pools monitored by means of an extracellular immunostaining (see Materials and Methods, Supplementary Information section 2.3.2) by using a GBP-Alexa555 nanobody against GFP (g, h; red in e, f) before (e, g) and after (f, h) acid wash. Acid wash in these conditions largely reduces the extracellular staining down to 9% of the signal. Scale bar: 10 µm. i, Acid wash does not affect internalized GBP-Alexa555. Confocal images of eGFP-DppLOP (top, green) and GBP-Alexa555 internalized for 40 min (bottom, red) before (left) and after acid wash (right). The GBP-Alexa555 signal decreases by 2.3 ± 0.6% after acid wash. j, Acid wash: effect of pH on GBP binding to GFP from larval extracts. Immunoblot of GFP which was bound to GFP-Trap beads (Chromotek, GFP-Trap beads, lanes 3-7) and GFP dissociated from GFP-Trap beads (supernatant, lanes 8-12) following treatment at different pH. FT, flowthrough (lane 1), PD, pulldown (lane 2). For gel source data, see Supplementary Fig. 1a. k, Stacked bar chart showing the relative contribution of the different modules to Dpp transport in the two theoretical extreme regimes of morphogen transport: extracellular diffusion (ExD20) and transcytosis (Tr) regimes. The relative contribution of different modules is expressed as the ratio λi2/λ2 with the index i corresponding to each of the four modules (i = u,b,r,t). Note that the unbound module contributes almost exclusively to λ2 in ExD and the transcytosis module, in Tr. l, Theoretical values of the 8 transport rates characteristic for ExD (rate values as in reference20) and Tr regimes of morphogen transport. m, n, FRAP recovery with respect to the two extreme theoretical regimes. Red lines, calculated recovery curves in a FRAP experiment for a set of parameter values corresponding to the extreme Tr (m) and ExD17 regimes (n). Blue dots, average of the experimental recovery curves in discs of l = 144 µm average posterior length. n = 9 biologically independent samples. Data represented as mean values ± s.e.m. The coefficient of determination R2 characterizes how well the calculated curves fit the experimental FRAP data. λ, decay length of the Dpp gradient profile calculated using equation (1) and the set of parameter values corresponding to Tr and ExD (see Supplementary Information section 4.2). Bars, s.e.m. Scale bar, 10 µm (a, h, i).
Extended Data Fig. 2 Analysis of Dpp leakage and effects of growth on Dpp gradient profile.
a, Confocal images of GBP-Alexa555 labelling extracellular GFP-Dpp in a control extracellular staining (top) and following a chase of living discs for 7 h at 4 °C. b, Total GBP-Alexa555 fluorescence in the conditions in a. Two-tailed two sample t-test, p-value = 0.7787. n, number of biologically independent samples. Bars, s.e.m. c, d, Schemes of sGFPPtc-Wg, see reference31 (c) and sGFPDpp constructs (d). Sizes of fragments represented in the scheme do not correspond to the nucleotide sequences. e, Confocal images of sGFPDpp (top), phalloidin staining (middle) and merge (bottom). Left panels, orthogonal views; right panels, xy plane. f, Normalized average spatial profile of sGFPDpp fluorescence (green) compared to the normalized profiles of gradients with decay lengths λ=λDpp; λ = 6L; λ = 3L and λ = 2L with λDpp = 28.9 µm and L = 144.6, average posterior size of eGFP-DppLOP third instar discs. g, Orthogonal views of confocal images of sGFPDpp fixed immediately after dissection (0 h) and following a chase of living discs for 1h at 25 °C and 4 °C. h, Total sGFP fluorescence in the conditions in g, normalized for each temperature to the value at t = 0 h. Two-tailed two sample t-test for unequal variances, p-values: 0.9792 (25 °C) and 0.7543 (4 °C). n, number of biologically independent samples. Bars, s.e.m. i, Effect of leakage on parameterization of Dpp transport rates. Average estimated parameters considering leakage rates kL = 0s−1; 0.00001 s−1; 0.0005 s−1 and 0.001 s−1. Simulations represent 3.7 x 106 randomly chosen parameter sets per condition. j, Stacked bar chart showing the relative contribution of the different modules to λ2 (described in Fig. 1e,f) for conditions in i. n, sample size; bars, s.d. k, Long-term FRAP assay. Dynamics of fluorescence recovery in conventional FRAP for one hour (red) and long-term FRAP for ten hours (blue). Fluorescence recovery is normalized to the signal in the ROI before bleaching. Note that recovery of conventional FRAP overlays the dynamics of long-term FRAP at short time scales. Bars, s.e.m. l, n, Dynamics of long-term FRAP recovery and fit to double (l, blue line) and single exponential dynamics (n, blue line) to the dataset (both early and late). Box in l, late recovery (after 5,000 s) analysed in m. m, Dynamics of long-term FRAP recovery (late recovery) and single exponential fit (blue line) to the late slow dynamics. o, Wing disc area plotted as a function of disc age in staged larvae (hours after egg laying) and fit to an exponential growth in which growth rate decays exponentially over time (red line). See Supplementary Information section 2.9. Orange and blue lines correspond to area and age of discs of l = 144 µm and l = 80 µm posterior length, respectively, as determined by the plot in p. p, Posterior compartment length (l) as a function of wing disc area (A). Black line, power-law fit. Growth anisotropy \(m={g}_{x}/g=\frac{\dot{{\ell }}/{\ell }}{\dot{A}/A}\). Using m, the area of discs of l = 144 µm and l = 80 µm posterior length can be determined (orange and blue lines). q, Wing disc growth rate (g), relaxation rate of the slow dynamics (that of the immobile fraction, IF) in long-term FRAP (kIF) and degradation rate of the immobile pool (k2) estimated according to k2 = kIF − g. The timescales corresponding to these rates are indicated on top of bars. r, s, Measurement of the mobile pool decay length. r, Confocal images of eGFP-DppLOP before (top) and at indicated times after bleaching (middle and bottom). s, Correlation between the decay length of the total pool of eGFP-Dpp at steady state (λT) measured before bleaching and the mobile pool decay length measured 30 min after bleaching (λM). Black line, linear regression. Note the slope close to 1, indicating that for discs of different sizes λM ≃ λT. Scale bar, 10 µm (a, e, g, r).
Extended Data Fig. 3 Parameterization assay controls I: steady-state decay length and nanobody internalization.
a, Immunoprecipitation of eGFP-Dpp under different expression systems. See Methods. Input (I) and immunoprecipitate (IP) from eGFP-DppCRISPR/+ (lanes 1,2), eGFP-DppCRISPR/CyO,Dpp+ (lanes 3,4), dppLG/+ ; LOP-eGFP-Dpp/+ (lanes 5,6; eGFP-DppLOP) and Dpp-Gal4/UAS- sfGFP-mKate2-Dpp larval head extracts (lanes 7,8). Mature GFP-Dpp fragment after Furin cleavages is marked by an asterisk. Note that GFP-Dpp amounts when expressed using LexA/LOP system are similar to the amounts of GFP-Dpp endogenously expressed (1.1 fold), whereas Gal4/UAS system expresses almost 400 fold more GFP-Dpp. For gel source data, see Supplementary Fig. 1b. b, Confocal image of eGFP-DppLOP in the background of overexpression of Dpp by dppGal4. c, d, Dynamics of FRAP recovery (c) and nanobody uptake (d) in this condition (red lines) as compared to control (blue).Bars, s.e.m. e, Average decay length λ of the gradients considered in the three datasets, corresponding to the three conditions considered in this report: large discs (average posterior length l = 144 µm in the dataset), small discs (average l = 80 µm) and in a pent2 mutant disc (average l = 130 µm). Bars, standard error to the mean (s.e.m.). The average decay length for the average l corresponding to the three experimental conditions was estimated using the linear regression of eGFP-DppLOP control (sample size n = 157 discs) and pentagone mutant (n = 63 discs) datasets (see Fig. 2a). f, Confocal images (maximum projections) of the eGFP-DppLOP gradient (red box, region of interest (ROI) in the posterior compartment) in representative discs from the three conditions described in b. The source is to the left. g, Average spatial distribution of eGFP-DppLOP in these datasets. Shaded areas, s.e.m. Black line, exponential fit. h, i, Left, normalized eGFP-DppLOP profiles in large control discs (h; l = 144 µm) and pent2 mutant disc (i; l = 130 µm); right, average residuals of the fits of these profiles to an exponential function. Bars, s.e.m. j, Scaling plot of eGFP-DppLOP. Decay length (λ, from the exponential fit) of the eGFP-DppLOP gradient versus l. Red line, linear regression. ϕL = λ/l determined from the linear regression. k, GBP-Alexa555 signal intensity as a function of time in 13 different discs. Lines, fits to the phenomenological \({c}_{{\rm{T}}}^{i}(t)\,\)equation for the internalized signal intensity (left equation in m; red/green boxes as in l). l, Average dynamics of the GBP-Alexa555 fluorescence signal in the three conditions. Bars, s.e.m. m, Parameterization of kN, ko and kr based on the dynamics of GBP-Alexa555 signal. Left, phenomenological \({c}_{{\rm{T}}}^{i}(t)\,\)equation which captures the exponential (red box; see also l) and linear dynamics (green box) of the accumulation of the GBP-Alexa555 signal. Right, relationship between the phenomenological parameters A, B and p and kN, ko and kr (see Supplementary Information section 2.2.1). n, Scheme of the GBP-Alexa555 internalization assay. Rates and pools indicated, like in Fig. 1d. Note that the fluorophore (Alexa555; star) degrades on a time scale which is much longer than the duration of the experiment. o, Confocal images of internalized GBP-Alexa555 in a disc expressing eGFP-DppLOP (top) and a control disc (bottom) at indicated timepoints of nanobody internalization using the same nanobody concentration as in Fig. 2b–f. Note that, under these conditions, fluid-phase internalization of the nanobody in the absence of eGFP-DppLOP (bottom, control) is negligible compared to the internalization when bound to eGFP-DppLOP (top, eGFP-Dpp). p, Dynamics of internalized GBP-Alexa555 in the disc expressing eGFP-DppLOP (green curve) and a control disc (blue curve), in the same experimental conditions (e.g. same nanobody concentration) as in the nanobody uptake experiments in o. Note that, in these conditions, internalization of GBP-Alexa555 by fluid phase in the absence of GFP-Dpp is negligible. q–r, Dynamics of fluid-phase internalization of GBP-Alexa555. q, Confocal image of fluid-phase internalized GBP-Alexa555 (40 min of nanobody incubation) showing that, at high concentration of the nanobody, a signal can be detected at low levels which is homogenous in space (there is no gradient). Five-fold higher concentration of the nanobody than in o was used to reliably detect the signal of the fluid-phase internalized nanobody. r, Dynamics of fluid-phase internalized GBP-Alexa555 signal intensity, averaged over 3 independent experiments. Same concentration as in p. Shaded area, s.e.m. Note that the dynamics do not show the early exponential regime seen in the presence of eGFP-Dpp, indicating that the nanobody by itself is not significantly recycled. s, Top, confocal image of fluid-phase internalized Alexa555 (40 min of Alexa555 incubation). Also here, internalization of the fluorophore is homogeneous in space. Bottom, high magnification of the ROI area shown in the top. t, Dynamics of fluid-phase internalized Alexa555, showing a linear increase without saturation in the timescale of the experiment, which reflects a lack of degradation in the lysosome of the Alexa555 fluorophore. u, Confocal images of the eGFP-DppLOP gradient (left) and internalized GBP-Alexa555 (right) after 45 min of incubation with the nanobody in a control large disc. The source is to the left. In contrast to the situation for fluid phase internalization (p, r), internalized eGFP-DppLOP with GBP-Alexa555 is distributed as a gradient. v, Spatial profiles of the gradients in u in the posterior compartment. The decay length is determined by fitting the spatial profiles to an exponential function with an offset. The decay length is given with its confidence interval. n, number of biologically independent samples. Bars, s.e.m (c, g, h, l, r). Scale bars, 10 µm (b, f, o, s, u) and 50 µm, (q).
Extended Data Fig. 4 Parameterization assay controls II: FRAP, extracellular fraction determination and parameter estimation by ABC.
a, Left, confocal image of the eGFP-DppLOP gradient in a FRAP experiment (source and posterior compartment). Red box, region to be photobleached. Right, eGFP-DppLOP fluorescent signal in the red box region before photobleaching (−1 min) and at different times (as indicated below) after photo-bleaching. b, Average dynamics of fluorescence recovery in the bleached area in the three experimental conditions (discs of l = 144 µm and l = 80 µm posterior length and in a pent2 mutant disc). Data represented as mean values. Bars, s.e.m. Lines, calculated recovery using the five-pool theoretical framework for a set of parameter values. The coefficient of determination R2 characterizes how well the theoretical curves fit the FRAP data. n, sample size. c, d, Robustness analysis of the FRAP assay. The average FRAP trace was fitted by a single dynamic equation3. Dependence of the goodness of the fit (R2) to this single dynamic equation (c) and the effective diffusivity (Deff) estimated by this fit (d) on the number of individual recovery curves (n) considered for the average FRAP trace. The analysis was performed for the three experimental conditions of this report: large discs (average posterior length l = 144 µm in the dataset; left), small discs (average l = 80 µm; centre) and pent2 discs (right). Bars, confidence intervals (d). In d data are represented as Deff estimated by fit for varying number of independent recovery curves, n. Bars, confidence intervals of fit. e, Effective diffusivity (Deff, left) and effective degradation rate (keff, right) plotted against the average posterior length of discs within two datasets: small (average l = 80 µm) and large (average l = 144 µm). The average FRAP recovery curve was fitted by a single dynamic equation3 to determine Deff and keff. Note, that as discs grow, Deff does not change significantly, whereas keff decreases significantly, as previously reported23. Data is represented as Deff and keff estimated by fit. Bars, confidence intervals of fit. n, number of biologically independent samples. One-tailed two sample t-test with unequal variances; p-values: 0.1765 (Deff, left) and 0.0038 (keff, right). f, Simulated intensity profile of eGFP-DppLOP at indicated times after photobleaching in the ROI in the posterior compartment (experiment as in a). x, distance from the edge of the anterior compartment. Parameter values used in the simulations are those of our parmeterization for l = 144 µm. g, Confocal images of the eGFP-DppLOP gradient (left; total pool), and the extracellular eGFP-DppLOP pools monitored by means of an extracellular immunostaining (see Supplementary Information section 2.3) by using a GBP-Alexa555 nanobody against GFP (right; extracellular pool). Higher magnification of the fluorescent signal of the area boxed in the images are shown to the right. h, Expression of the extracellular fraction (ρ) as function of Dpp transport rates. i, Equimolarity of the GBP-Alexa555 and eGFP solutions used for calibration of the Alexa555 versus GFP fluorescent signal (see Methods, Supplementary Information section 2.3.2; relevant to the extracellular fraction determination assay). The concentrations of GBP-Alexa555 and eGFP was first roughly determined by means of a BCA assay (Supplementary Information section 2.3.2). Plot of GFP fluorescence intensity as a function of the ratio of GBP-Alexa555 and GFP concentrations (determined by BCA) in the solutions. The relative concentration of GFP and GBP-Alexa555 can be determined from the relative concentration at which the minimum value (rmin) of GFP fluorescence has been reached. Note that rmin ≃ 1 confirms that the BCA estimation was already accurate. j, Parameter value sets determined by the parameterization procedure (see Supplementary Information section 2.5.2) are represented in the (kon, koff) plane. Light orange area represents the full space of 3 × 107 parameter value sets considered (l = 144 µm dataset). Dark orange dots represent sets of parameter values within those which satisfy the constraints given by the steady-state decay length, the long-term FRAP assay, the nanobody internalization and the FRAP assay. Calculated FRAP recovery curves using these sets of values fit the experimental FRAP data with R2>0.92. Note that the solutions are separated into two clusters (clouds): the upper cloud, with higher kon, koff, is characterized by a low extracellular fraction ρ<0.10 and a lower cloud, by a high ρ<0.25. k, Selected sets of parameter values from j for which the calculated extracellular fraction is within the experimentally determined range of ρ values (0.08<ρ<0.18). l, Sets of parameter values which satisfy all the constraints given by our assays (see Supplementary Information section 2.5.2), represented in (koff, kon), (koff, k), (D0, kon) and (ko, kon) planes. The parameter values corresponding to the two extreme theoretical cases discussed in Supplementary Information section 4.2 (Extracellular diffusion regime, ExD, yellow and Transcytosis regime, Tr, purple) are represented by circles for comparison. m, Average estimated parameters in the three experimental conditions compared to the theoretical values of parameters in ExD and Tr. Bars, s.d. N, number of parameterized sets of values. Scale bars: 10 µm (a, g).
Extended Data Fig. 5 Quantitative considerations: robustness analysis and decay length boosts.
a, Cluster of parameter value sets in the (kon, koff) plane corresponding to three different ranges of R2 to the experimental FRAP recovery for the three experimental conditions. The coefficient of determination R2 characterizes the goodness of the fit between the FRAP data and the calculated recovery curves. Relaxing the quality of fit down to R2>0.85 (from R2>0.93) does not populate the lower cloud, and therefore does not affect the assignment to the ExD-type versus Combined transport regimes. Points that populate the lower cloud as in the l = 80 µm and pent2 conditions) require that R2< \({{\rm{R}}}_{{\rm{th}}}^{2}\) (see Supplementary Information section 3.7 for details). b, Cluster of parameter value sets in the (kon, koff) plane corresponding to different ranges of calculated extracellular fraction ρ for the three experimental conditions. An increase in ρ beyond ρ* is required to shift the solutions to the “lower” cloud. The lower cloud is characteristic of the ExD-type regime. c, d, Sets of parameter values (clouds of points) compatible with all the assays considered in this report in the (kon, koff) plane. Isolines for Boost kr (c) and Boost koff (d) are also represented (see look up table). See Supplementary Information section 3.5 for definition of the Boosts. The three conditions considered in this work are shown: large discs (average posterior length l = 144 µm in the dataset; left), small discs (average l = 80 µm; centre) and pent2 discs (right). e, Average calculated Boost kr, Boost koff and Boost D0 for the three experimental conditions compared to the calculated Boosts for the theoretical values of parameters in the ExD and Tr regimes. N, number of parameterized sets of values. Data represented as mean values over N parameterized value sets. Bars, s.e.m. f–i, iFRAP assay. f, Scheme of the iFRAP assay (see Supplementary Information section 2.7). g, h, Test of efficiency of the acid wash in the iFRAP (and photoconversion) experiment: GFP-Dpp expressing discs have been incubated in GBP-Alexa555 for 50 min at 4 °C (the nanobody is only bound to the extracellular pool) and subsequently acid-washed to remove the label of the extracellular pool. Confocal image of eGFP-DppLOP expressing disc (g) and corresponding images of GBP-Alexa555 (h) at indicated times after the acid wash (see Materials and Methods). Note that no detectable GBP-Alexa555 signal is observed 40 min after the acid wash, indicating that the extracellular pool of nanobodies has been efficiently removed and that the potential extracellular leftover (below the detection limit) cannot lead to an observable recovery in intracellular compartments. i, Theoretical dynamics of GBP-Alexa555 fluorescence recovery in the iFRAP experiment normalized to the pre-photobleaching levels. Recovery was calculated numerically using the set of values determined experimentally for large (top) and small discs (bottom). The dashed lines indicate the estimated fraction of recovery 2,000s after photobleaching in large and small discs to compare with the experimental conditions in the iFRAP experiments (Fig. 4g). Scale bar, 10 µm (g, h).
Extended Data Fig. 6 Internalized Dpp is recycled and spreads in the tissue: DppTimer and recycling Rab proteins.
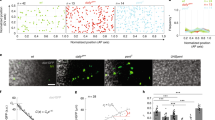
a, Functionality of DppTimer. Left, control disc, expressing sfGFP-mKate2-Dpp under the control of the GAL4/UAS expression system (DppTimer). Centre, dpp mutant disc, the wing imaginal disc is outlined with the white dashed line. Right, dpp mutant disc expressing DppTimer. Note that the mutant phenotype seen in the central image is rescued. b, Scatter plot of sfGFP and mKate2 pixel intensities and linear fit to obtain the calibration factor F (see Supplementary Information section 2.6.3). n = 23 beads. c, Confocal images of the DppTimer gradient in the wing disc (sfGFP, top and mKate2, bottom). d, Relative concentration profiles of mature sfGFP and mKate2 plotted against the distance from the Dpp source (see Supplementary Information section 2.6.3), corresponding to the intensity profiles measured from the images in c. These intensity profiles represent the relative amounts of sfGFP and mature mKate2 molecules. e, Adjusted fluorescence intensity profiles for sfGFP (g*(x)) and mature mKate2 (r*(x)) which are proportional to the respective concentration profiles. X-axis represents the distance from the source. Red dashed line is positioned at the anterior-posterior boundary. Note that both in the source and in the region of the target closer to the source, there are less mature mKate2 molecules, confirming that Dpp molecules are younger closer to the source. f, Plotted relative age (A(x)) of Dpp molecules as a function of position calculated from the calibrated profiles in e. Note that as molecules move away from the source they become older on average: A(x) increases to plateau at values close to 1. n, number of biologically independent samples. Shaded areas, s.e.m (e, f). g–j, Effect of pH on the Timer. g, Control of the bafilomycin treatment. Confocal images of a ROI in discs incubated with a LysoSensorTM probe for 30 min before (top) and after (bottom) incubation in control Clone 8 medium (right) or bafilomycin solution (left). h, Effect of pH on sfGFP and mKate2 in the DppTimer. Confocal images of sfGFP (left) and mKate2 (right) of DppTimer before (top) and after (bottom) neutralization of pH to 7 following bafilomycin treatment for 30 min. i, Fluorescence signal decrease of sfGFP and mKate2 owing to acidic pH in intracellular compartments. Percentage decrease of fluorescence from pH 7 (discs after bafilomycin treatment) to the acidic environment in intracellular compartments (discs before bafilomycin treatment). Note that the decrease is very similar for both fluorophores. j, Normalized fluorescence intensity of sfGFP (blue) and mKate2 (orange) in purified Timer molecules in solutions at different pH. Data normalized to the intensity at pH 7.4. The number of biologically independent samples for this analysis: npH5.86 = 8; npH6.4 = 7; npH7.4 = 7; npH7.9 = 5. Data represented as mean values ± s.e.m. Note, that the difference between the normalized intensity of sfGFP and mKate2 at the different pH value is not significant (p-value>0.05; two-tailed two sample t-test). k, Confocal images of eGFP-DppLOP in control condition (top) and after RNAi through expression of dsRNA for the recycling Rab proteins, Rab11 (middle) and Rab4 (bottom) in posterior target cells. l, Spatial fluorescence profiles of eGFP-DppLOP corresponding to control (top), Rab11RNAi (middle) and Rab4RNAi (bottom) conditions in k. m, Decay length λ of eGFP-DppLOP gradient versus posterior compartment length l for control (n = 157), pent2 discs (n = 63) and Rab4RNAi (n = 39). Dots, binned data; bars, s.e.m. Control and pent2 data as in Fig. 2a, Extended Data Fig. 7f. n, Average eGFP-DppLOP decay length in control and Rab11RNAi conditions. Difference between the two conditions is significant as determined by a two-tailed, two sample t-test with unequal variances, p-value = 0.0034. o, Recycling rate in control and Rab4RNAi conditions, determined by the nanobody uptake assay. Number of curves for each condition is n = 4. Difference between the two conditions is significant; two-tailed, two sample t-test with unequal variances, p-value < 0.0001. Rab4RNAi expression was driven by means of the thermosensitive Gal4Gal80ts system (29 °C). p–r, Scaling of eGFP-DppLOP. p, Dpp gradient profiles of discs from 40 to 160 µm posterior length. Each individual profile was fitted to an exponential function with an offset (see Supplementary Information section 2.1.2) and the offset returned from the fit was subtracted. q, Normalized Dpp gradient profiles. Each profile was normalized to the amplitude C0 of its exponential fit in the ordinates (C(r)/C0) and to the posterior length l of the corresponding wing disc in the abscissas (r=x/l). Shaded area, s.e.m. Black line, average normalized profile. r, Density plot of q: Colour-code corresponds to the fraction of the number of gradients passing through a certain r, C(r)/C0 bin. Scale bars, 100 µm (a) and 10 µm (c, g, h, k).
Extended Data Fig. 7 Gradient scaling by recycling: Pentagone.
a, Continuous and monotonic transition from λ ≈ 15 μm (black dashed line) to λ ≈ 27 μm (red dashed line). Left: decay length (λ) versus a parameter b that captures monotonic and continuous changes in kon, koff and D0 as shown in the right. Right: Variations in kon, koff and D0 with b as defined by the equations shown in the plot. Black and red dashed lines indicate initial (small discs) and final (large discs) values for kon, koff and D0. b, Top: expression for the ratio of the recycling to the unbound module (λr2/λu2, see Fig. 1c). Bottom: Sets of parameter values (clouds of points) compatible with all the assays considered in this report in the (kon, koff) plane. Isolines for (λr2/λu2 are also shown (see look-up table). The three conditions considered in this work are shown: large discs (average posterior length l = 144 µm in the dataset; left), small discs (average l = 80 µm; centre) and pent2 discs (right). These isolines convey the relative importance of the recycling and the unbound modules to the Dpp transport. c, PMAD scaling analysis for control and pentagone mutants. Left, Decay length λ of PMad gradients plotted as a function of posterior compartment length l. Raw and binned data (Bar, s.e.m) are shown together with a linear regression to the raw data. Right, bar plots showing the slopes ϕ of corresponding linear regressions for control (blue) and pentagone mutant experimental conditions (red). Number of biologically independent samples: n = 45 (control) and n = 25 (pent2). ****p-value < 0.00001; two-tailed two sample t-test with unequal variances. Bars, confidence intervals at 95%. d, UAS-GFP-Pentagone expression driven by ap-Gal4. In the right, higher magnification of the area boxed in the image to the left. Scale bars, 10 µm. e, GFP-Pentagone gradient profile in the ventral compartment. The profile is fitted to an exponential function (red) to determine the decay length shown. x, distance from the dorso-ventral boundary. f, eGFP-DppLOP scaling analysis for control and pentagone mutants. Left, Decay length λ of eGFP-Dpp gradients plotted as a function of posterior compartment length l. Raw and binned data (Bar, s.e.m) are shown together with a linear regression to the raw data. Right, bar plot showing the slopes ϕ of corresponding linear regressions from these plots. Control experimental condition (blue) compared to pentagone mutant experimental condition (red). Number of biologically independent samples: n = 157 (control) and n = 63 (pent2). ****p-value < 0.00001; two-tailed two sample t-test with unequal variances. Bars, confidence intervals at 95%. g, Sets of parameter values satisfying the constraints given by all the experimental assays represented in (kon, koff), (kon, D0) and (k, koff) planes in the four experimental conditions: eGFP-DppLOP-expressing discs of 144 µm and 80 µm average posterior length and pent2 mutant discs of 130 µm and 85 µm average posterior length. h, Stacked bar chart showing the relative contribution of the different modules to λ2 (described in Fig. 1e,f) in the four experimental conditions in e compared to the theoretical values of parameters in the extracellular diffusion (ExD) and transcytosis regimes of transport (Tr). i, Average extracellular fraction in control discs of 144 µm and 80 µm average posterior length and pent2 mutant discs of 130 µm and 85 µm average posterior length. Box plot represents the minimum and the maximum, median, 25th and 75th percentile. n, number of biologically independent samples. j, Confocal images of PentGFP from the endogenous gene in discs of different sizes. Scale bar, 10 µm. Dotted lines, contour of discs. k, PentGFP average intensity in its expression domain as a function of the squared posterior length of the wing disc; Black, binned data. Orange dots, raw data. Bars, s.e.m. Vertical boxes indicate posterior width sizes l = 144 µm (orange) and l = 80 µm (blue).
Extended Data Fig. 8 Gradient scaling by recycling: HSPGs.
a, b, Scaling analysis for control and dally mutants. Left, decay length λ of eGFP-Dpp (a) and PMad gradients (b) plotted as a function of posterior compartment length l. Raw and binned data (bars, s.e.m.) are shown together with a linear regression to the raw data. Right, bar plots showing the slopes ϕ of corresponding linear regressions for control experimental conditions (blue) compared to dally mutant experimental conditions (red). Number of biologically independent samples: n = 93 (control) and n = 39 (dallygem) (a); n = 43 (control) and n = 36 (dallygem) (b). ****p-value < 0.00001; two-tailed two sample t-test with unequal variances. Bars, confidence intervals at 95%. c, Sets of parameter values satisfying the constraints given by all the experimental assays represented in (kon, koff), (kon, D0) and (k, koff) planes in the four experimental conditions: eGFP-Dpp discs of 144 µm and 80 µm average posterior length, pent2 (average length, 130 µm) mutant and dallygem mutant discs (average length, 174 µm). d, Stacked bar chart showing the relative contribution of the different modules to λ2 (described in Fig. 1e,f) in the four experimental conditions compared to the theoretical values of parameters in the extracellular diffusion (ExD) and transcytosis regimes of transport (Tr). e, GBP-Alexa555 signal intensity as a function of time in discs expressing eGFP-DppGal4 in control discs (left), dallygem mutant discs (middle) and control discs following treatment with PI-PLC for 1h (right). Lines, fits to the phenomenological equation describing the internalized signal intensity dynamics CT(t). f, Values of kN, kr and k0 estimated by the nanobody uptake assay in control discs, dallygem mutant discs and PI-PLC treated discs expressing eGFP-DppGal4. g, Internalized GBP-Alexa555 fluorescence as a function of time in discs expressing eGFP-DppCRISPR (control), discs expressing eGFP-DppCRISPR and sflRNAi (sflRNAi) and control discs (no GFP-Dpp). Number of biologically independent samples: n = 3 for each condition. Data represented as the average curve. Shaded area, s.e.m. h, i, Confocal images of eGFP-DppCRISPR (left) and internalized GBP-Alexa555 (right) after 85 min of incubation with the nanobody in control discs (h) and discs expressing sflRNAi in the posterior compartment (i). Posterior compartment, to the right from the GFP-Dpp source boundary. j, Decay length of the eGFP-DppCRISPR gradient λ as a function of the posterior compartment width l. Red line, linear regression to the raw data. bars, s.e.m. eGFP-DppCRISPR was visualized by means of a nanobody uptake assay (Methods). Number of biologically independent samples n = 38. k, Slope ϕ of the linear regressions for scaling plots corresponding to eGFP-DppLOP (LOP) and eGFP-DppCRISPR (CRISPR). Bars, confidence intervals of the fitted slope. l, Confocal images of photoconverted GBP-Dendra2* in eGFP-DppCRISPR-expressing discs at different times after photoconversion (post-conversion). Before photoconversion, discs were incubated in GBP-Dendra2* solution for 45 min and extracellular GBP-Dendra2 was removed by an acid wash, so that only internalized GBP-Dendra2 is remaining. PhotoconvOgradient outside of the photoconverted region. m, The values of kN, kr and k0 estimated by the nanobody uptake parameters for large discs expressing eGFP-DppCRISPR versus eGFP-DppLOP. Bars, confidence intervals of the fits. Number of biologically independent samples n = 10 (eGFP-DppCRISPR) and n = 13 (eGFP-DppLOP). Scale bar, 10 µm (h, l).
Supplementary information
Supplementary Information
This file contains the Supplementary Methods, Supplementary Notes, Supplementary Discussion, Supplementary Fig. 1, Supplementary Tables 1–2 and Supplementary References.
41586_2021_4346_MOESM4_ESM.avi
Supplementary Video 1 Photoconversion assay. Left, a movie combining sequentially first, a confocal image of eGFP-DppLOP, then an image of photoconverted endosomal GBP–Dendra2 (Dendra2*) before (pre-conversion) and finally images of Dendra2* at indicated times after photoconversion (post-conversion). Before conversion, following pulse-chase and acid wash, only internalized GBP–Dendra2 remains. Photoconversion, to the left of the red dotted line. Note build-up of a Dendra2* gradient outside the photoconverted region. Right, average spatial distribution of GBP–Dendra2* fluorescence signal at indicated times after photoconversion. Shaded areas, s.e.m.; (n = 7 independent experiments).
Source data
Rights and permissions
About this article
Cite this article
Romanova-Michaelides, M., Hadjivasiliou, Z., Aguilar-Hidalgo, D. et al. Morphogen gradient scaling by recycling of intracellular Dpp. Nature 602, 287–293 (2022). https://doi.org/10.1038/s41586-021-04346-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04346-w
This article is cited by
-
Precise and scalable self-organization in mammalian pseudo-embryos
Nature Structural & Molecular Biology (2024)
-
Highly conserved and extremely evolvable: BMP signalling in secondary axis patterning of Cnidaria and Bilateria
Development Genes and Evolution (2024)
-
Single-molecule tracking of Nodal and Lefty in live zebrafish embryos supports hindered diffusion model
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.