Abstract
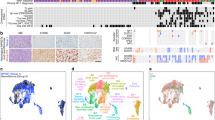
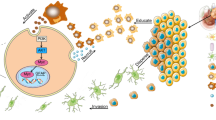
Fatty acid uptake and altered metabolism constitute hallmarks of metastasis1,2, yet evidence of the underlying biology, as well as whether all dietary fatty acids are prometastatic, is lacking. Here we show that dietary palmitic acid (PA), but not oleic acid or linoleic acid, promotes metastasis in oral carcinomas and melanoma in mice. Tumours from mice that were fed a short-term palm-oil-rich diet (PA), or tumour cells that were briefly exposed to PA in vitro, remained highly metastatic even after being serially transplanted (without further exposure to high levels of PA). This PA-induced prometastatic memory requires the fatty acid transporter CD36 and is associated with the stable deposition of histone H3 lysine 4 trimethylation by the methyltransferase Set1A (as part of the COMPASS complex (Set1A/COMPASS)). Bulk, single-cell and positional RNA-sequencing analyses indicate that genes with this prometastatic memory predominantly relate to a neural signature that stimulates intratumoural Schwann cells and innervation, two parameters that are strongly correlated with metastasis but are aetiologically poorly understood3,4. Mechanistically, tumour-associated Schwann cells secrete a specialized proregenerative extracellular matrix, the ablation of which inhibits metastasis initiation. Both the PA-induced memory of this proneural signature and its long-term boost in metastasis require the transcription factor EGR2 and the glial-cell-stimulating peptide galanin. In summary, we provide evidence that a dietary metabolite induces stable transcriptional and chromatin changes that lead to a long-term stimulation of metastasis, and that this is related to a proregenerative state of tumour-activated Schwann cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All raw data from gene expression RNA microarrays, ChIP–seq, RNA-seq, scRNA-seq, PRO-seq and spatial transcriptomics are available at the Gene Expression Omnibus (GEO) repository under accession code GSE148321. Source data are provided with this paper.
Code availability
R v.4.0.1 and Python v.3.6.0 were used. All of the code used to analyse the single-cell and positional RNA-seq data are available at GitHub (https://github.com/MarcElosua/10X-EPID).
References
Peck, B. & Schulze, A. lipid metabolism at the nexus of diet and tumor microenvironment. Trends Cancer 5, 693–703 (2019).
Pascual, G., Domínguez, D. & Benitah, S. A. The contributions of cancer cell metabolism to metastasis. Dis. Model. Mech. 11, dmm032920 (2018).
Boilly, B., Faulkner, S., Jobling, P. & Hondermarck, H. Nerve dependence: from regeneration to cancer. Cancer Cell 31, 342–354 (2017).
Zahalka, A. H. & Frenette, P. S. Nerves in cancer. Nat. Rev. Cancer 20, 143–157 (2020).
Solans, M., Chan, D. S. M., Mitrou, P., Norat, T. & Romaguera, D. A systematic review and meta-analysis of the 2007 WCRF/AICR score in relation to cancer-related health outcomes. Ann. Oncol. 31, 352–368 (2020).
Pascual, G. et al. Targeting metastasis-initiating cells through the fatty acid receptor CD36. Nature 541, 41–45 (2017).
Lee, C. et al. Tumor metastasis to lymph nodes requires YAP-dependent metabolic adaptation. Science 363, 644–649 (2019).
Rambow, F. et al. Toward minimal residual disease-directed therapy in melanoma. Cell 174, 843–855 (2018).
Haobin, Ye et al. Leukemic stem cells evade chemotherapy by metabolic adaptation to an adipose tissue niche. Cell Stem Cell 19, 23–37.
Abdelmagid, S. A. et al. Comprehensive profiling of plasma fatty acid concentrations in young healthy Canadian adults. PLoS ONE 10, e0116195 (2015).
Feng, R. et al. Free fatty acids profile among lean, overweight and obese non-alcoholic fatty liver disease patients: a case—control study. Lipids Health Dis. 16, 165 (2017).
Nyanbol, Kuol et al. Role of the nervous system in cancer metastasis. J. Exp. Clin. Cancer Res. 37, 5 (2018).
Wang, W. et al. Nerves in the tumour microenvironment: origin and effects. Front. Cell Dev. Biol. 8, 1630 (2020).
Venkatesh, H. S. et al. Electrical and synaptic integration of glioma into neural circuits. Nature 573, 539–545 (2019).
Amit, M. et al. Loss of p53 drives neuron reprogramming in head and neck cancer. Nature 578, 449–454 (2020).
Venkataramani, V. et al. Glutamatergic synaptic input to glioma cells drives brain tumour progression. Nature 573, 532–538 (2019).
Ubink, R., Calza, L. & Hökfelt, T. ‘Neuro’-peptides in glia: Focus on NPY and galanin. Trends Neurosci. 26, 604–609 (2003).
Gresle, M. M. et al. Galanin is an autocrine myelin and oligodendrocyte trophic signal induced by leukemia inhibitory factor. Glia 63, 1005–1020 (2015).
Avgustinova, A. & Benitah, S. A. The epigenetics of tumour initiation: cancer stem cells and their chromatin. Curr. Opin. Genet. Dev. 36, 8–15 (2016).
Schuettengruber, B., Bourbon, H.-M., Di Croce, L. & Cavalli, G. Genome regulation by polycomb and trithorax: 70 years and counting. Cell 171, 34–57 (2017).
Fawcett, J. W., Oohashi, T. & Pizzorusso, T. The roles of perineuronal nets and the perinodal extracellular matrix in neuronal function. Nat. Rev. Neurosci. 20, 451–465 (2019)
Quraishe, S., Forbes, L. H. & Andrews, M. R. The extracellular environment of the CNS: influence on plasticity, sprouting, and axonal regeneration after spinal cord injury. Neural Plast. 2018, 2952386 (2018).
Dzyubenko, E., Gottschling, C. & Faissner, A. Neuron-glia interactions in neural plasticity: contributions of neural extracellular matrix and perineuronal nets. Neural Plast. 2016, 5214961 (2016).
Bartus, K. et al. Large-scale chondroitin sulfate proteoglycan digestion with chondroitinase gene therapy leads to reduced pathology and modulates macrophage phenotype following spinal cord contusion injury. J. Neurosci. 34, 4822–4836 (2014).
McDonald, O. G. et al. Epigenomic reprogramming during pancreatic cancer progression links anabolic glucose metabolism to distant metastasis. Nat. Genet. 49, 367–376 (2017).
Shi, X. et al. The abundance of metabolites related to protein methylation correlates with the metastatic capacity of human melanoma xenografts. Sci. Adv. 3, 11 (2017).
Clements. M. P. et al. The wound microenvironment reprograms Schwann cells to invasive mesenchymal-like cells to drive peripheral nerve regeneration. Neuron 96, 98–114 (2017).
Johnston, A. et al. Dedifferentiated Schwann cell precursors secreting paracrine factors are required for regeneration of the mammalian digit tip. Stem Cell 19, 433–448 (2016).
Ana-Maria, E. et al. Targeting CD36 as biomarker for metastasis prognostic: how far from translation into clinical practice? BioMed Res. Int. 2018, 7801202 (2018).
Gabriele, B. & Sarah-Maria, F. The metabolism of cancer cells during metastasis. Nat. Rev. Cancer 21, 162–180 (2021).
Sundaresan, S. & Abumrad, N. A. Dietary lipids inform the gut and brain about meal arrival via CD36-mediated signal transduction. J. Nutr. 145, 2195–2200 (2015).
Wang, L. et al. A cytoplasmic COMPASS is necessary for cell survival and triple-negative breast cancer pathogenesis by regulating metabolism. Genes Dev. 31, 2056–2066 (2017).
Bledau, A. S. et al. The H3K4 methyltransferase Setd1a is first required at the epiblast stage, whereas Setd1b becomes essential after gastrulation. Development 141, 1022–1035 (2014).
Liu, D., Archer, N., Duesing, K., Hannan, G. & Keast, R. Mechanism of fat taste perception: association with diet and obesity. Prog. Lipid Res. 63, 41–49 (2016).
Yang, P. et al. Dietary oleic acid-induced CD36 promotes cervical cancer cell growth and metastasis via up-regulation Src/ERK pathway. Cancer Lett. 438, 76–85 (2018).
Ubellacker, J. M. et al. Lymph protects metastasizing melanoma cells from ferroptosis. Nature 585, 113–118 (2020).
Vriens, K. et al. Evidence for an alternative fatty acid desaturation pathway increasing cancer plasticity. Nature 566, 403–406 (2019).
Myers, J. N., Holsinger, F. C., Jasser, S. A., Bekele, B. N. & Fidler, I. J. An orthotopic nude mouse model of oral tongue squamous cell carcinoma. Clin. Cancer Res. 8, 293–298 (2002).
Benaich, N. et al. Rewiring of an epithelial differentiation factor, miR-203, to inhibit human squamous cell carcinoma metastasis. Cell Rep. 9, 104–117 (2014).
Schneider, T. E. et al. Measuring stem cell frequency in epidermis: a quantitative in vivo functional assay for long-term repopulating cells. Proc. Natl Acad. Sci. USA 100, 11412–11417 (2003).
Shwartz, Y. et al. Cell types promoting goosebumps form a niche to regulate hair follicle stem cells. Cell 182, 578–593, (2021).
Bhandari, M. et al. Galanin receptor antagonist M35 but not M40 or C7 ameliorates cerulein-induced acute pancreatitis in mice. Pancreatology 10, 682–688 (2011).
Zuber, J. et al. Mouse models of human AML accurately predict chemotherapy response. Genes Dev. 23, 877–889 (2009).
Nowak, J. A. & Fuchs, E. Isolation and culture of epithelial stem cells. Methods Mol. Biol. 482, 215–232 (2009).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Soneson, C. & Robinson, M. D. Bias, robustness and scalability in single-cell differential expression analysis. Nat. Methods 15, 255–261 (2018).
Elosua-Bayes, M., Nieto, P., Mereu, E., Gut, I. & Heyn, H. SPOTlight: seeded NMF regression to deconvolute spatial transcriptomics spots with single-cell transcriptomes. Nucleic Acids Res. 49, e50 (2021).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map (SAM) Format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2018); http://www.R-project.org/
Stark, R. & Brown, G. DiffBind: differential binding ChIP-seq peak data. Bioconductor 1–27 (2011).
Yu, G., Wang, L. G. & He, Q. Y. ChIP seeker: An R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 128 (2013).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Douillet, D. et al. Uncoupling histone H3K4 trimethylation from developmental gene expression via an equilibrium of COMPASS, Polycomb and DNA methylation. Nat. Genet. https://doi.org/10.1038/s41588-020-0618-1 (2020).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1100 (2013).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Gonzalez-Roca, E. et al. Accurate expression profiling of very small cell populations. PLoS ONE 5, e14418 (2010).
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics, 4, 249–264 (2003).
Chenj, J. et al. ToppGene Suite for gene list enrichment analysis and candidate genes for priorizitation. Nucleic Acid Res. 37, W305–W311 (2009).
Berriz, G. F., King, O. D., Bryant, B., Sander, C. & Roth, F. P. Characterizing gene sets with FuncAssociate. Bioinformatics 19, 2502–2504 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Dougherty, J. D. et al. Analytical approaches to RNA profiling data for the identification of genes enriched in specific cells. Nucleic Acids Res. 38, 4218–4230 (2010).
Xu, X. et al. Cell type-specific expression analysis to identify putative cellular mechanisms for neurogenetic disorders. J. Neurosci. 34, 1420–1431 (2014).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Zambelli, F. et al. Pscan: finding over-represented transcription factor binding site motifs in sequences from co-regulated or co-expressed genes. Nucleic Acids Res. 37, W247–W252 (2009).
Nagy, A. et al. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci. Rep. 8, 11515 (2018).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2008).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A., affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Bolstad, B. M. et al. in Bioinformatics and Computational Biology Solutions Using R and Bioconductor. Statistics for Biology and Health (eds Gentleman R. et al.) 33–47 (Springer, 2005).
Eklund, A. C. & Szallasi, Z. Correction of technical bias in clinical microarray data improves concordance with known biological information. Genome Biol. 9, R26 (2008).
Smyth, G. in Bioinformatics and Computational Biology Solutions Using R and Bioconductor. Statistics for Biology and Health (eds Gentleman R. et al.) 397–420 (Springer, 2005).
Benjamini, Y. H. Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Wu, D. et al. ROAST: rotation gene set tests for complex microarray experiments. Bioinformatics 26, 2176–2182 (2010).
B., Efron, & R., Tibshirani, On testing the significance of sets of genes. Ann. Appl. Stat. 1, 107–129 (2007).
Goeman, J. J., & Bühlmann, P. Analyzing gene expression data in terms of gene sets: methodological issues. Bioinformatics 23, 980–987 (2007).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Carlson M. org.Hs.eg.db: genome wide annotation for Human. R package version 3.8.2. (2019).
Carlson M. org.Mm.eg.db: genome wide annotation for Mouse. R package version 3.8.2. (2019).
Acknowledgements
We thank the staff at the histology and genomics facilities of the IRB Barcelona for their assistance in this work; and V. Raker for manuscript editing. Research in the S.A.B. laboratory is supported in part by the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 787041), the Government of Cataluña (SGR grant), the Government of Spain (MINECO), the La Marató/TV3 Foundation, the Foundation Lilliane Bettencourt, the Spanish Association for Cancer Research (AECC), the Worldwide Cancer Research Foundation (WCRF) and the BBVA Foundation. D. Domínguez was supported by a La Caixa International Fellowship for Doctoral studies; C. Bigas by an FPI fellowship (MINECO); and I.H. by the EU Horizon 2020 Marie Skłodowska-Curie award (no. 754510). H.H. has received funding from the Ministerio de Ciencia, Innovación y Universidades (SAF2017-89109-P; AEI/FEDER, UE). Studies in A. Shilatifard laboratory related to COMPASS are supported by NCI’s Outstanding Investigator Award R35CA197569; other research, by funding from University of Miami Miller School of Medicine, Sylvester Comprehensive Cancer Center, grant R01 GM078455 and DP1 CA228041 from the National Institute of Health (to R.S.), and the National Cancer Institute of the National Institutes of Health (no. P30CA240139; note that the content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health). The IRB Barcelona is a Severo Ochoa Center of Excellence (MINECO award SEV-2015-0505).
Author information
Authors and Affiliations
Contributions
G.P., D. Domínguez and S.A.B. designed the study. C.G. helped to design some of the experiments. G.P. performed all of the in vivo palm oil and olive oil diet studies and analysed the innervation phenotype, including all in vivo transcriptome analyses of tumour and neural compartments (bulk arrays, 10x scRNA-seq and positional RNA-seq). D. Domínguez performed all the in vitro and in vivo epigenetic studies and the mechanistic studies for the Set1A, MLL1 and MLL2 proteins. D. Domínguez and D. Douillet carried out the COMPASS western blotting experiments. G.P. performed the experiments in melanoma. C.L. analysed the scRNA-seq and gene expression data. M.E.-B. analysed the 10x scRNA-seq and the positional RNA-seq data. C. Bigas performed the correlative analyses of the CD36+ signature and perineuronal nets in human tumours. M.A. helped to perform the sympathectomy experiments. I.H. and A. Symeodini analysed ChIP–seq data. D. Domínguez performed the in vitro scRNA-seq experiment. S.R.G., I.H. and H.H. contributed to and analysed scRNA-seq data. N.P. characterized histology samples. F.B and R.S. performed some transcriptome analyses. C. Bescós provided the tumour samples to establish the patient-derived VDH15 oral cancer cell line. S.A.B. wrote the manuscript with the input from G.P and D. Domínguez.
Corresponding authors
Ethics declarations
Competing interests
S.A.B. is a co-founder and scientific advisor of ONA Therapeutics. The other authors declare no competing interests.
Additional information
Peer review information Nature thanks Sarah-Maria Fendt and the other, anonymous, reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Palmitic acid, but not linoleic, oleic or stearic, enhances the metastatic potential of OSCC long term after removing the stimulus.
a, Flow cytometry analysis of in vitro cultured SCC-25 cells after 4 days of fatty acid stimulation. The percentage of CD36 membrane expression and cell viability (as measured by DAPI incorporation) are shown. b, BLI quantification of lymph node (LN) metastases, showing number and size, from mice inoculated with untreated SCC-25 cells (n = 17) or in vitro treated with 300 µM PA (n = 17), 50 µM OA (n = 10) or 50 µM LA (n = 10). BLI signals are expressed as the relative normalized photon flux. P = 0.05, two-tailed t-test. c, Frequency of developed LN metastases from animals in b. *P = 0.01, **P = 0.001; two-tailed Fisher’s exact test. d, Flow cytometry analysis of in vitro cultured SCC-25 cells immediately after 4 days (4D) PA stimulation or at 14D after PA withdrawal (wdl). The numbers indicate [CD44brightCD36bright] and [CD44brightCD36dim] populations in the represented gate, expressed as percentage of total DAPI-negative cells from in vitro cultured cells (samples are representative of n = 5 independent experiments). e, BLI quantification of primary tumours generated from mice injected with SCC-25 cells in vitro 14D after PA withdrawal. The BLI signal is expressed as the relative normalized photon flux. Data are given as the mean and s.e.m. (untreated, n = 14; 14D after PA wdl, n = 14, *P = 0.05, two-tailed t-test). f, Frequency of developed LN metastases from animals in e (*P = 0.02, two-tailed Fisher’s exact test). g, ex vivo BLI lung metastasis quantification of mice injected with VDH-15 cells at 14D fatty acid withdrawal (PA, palmitic acid; SA, stearic acid; OA, oleic acid; LA, linoleic acid). BLI signals are expressed as the relative normalized photon flux. Images are representative of n = 15 mice per group. h, Frequency of developed lung metastasis of mice injected with SCC-25 after FA withdrawal expressed as percentages (n = 10 mice per group, lung metastases: *P = 0.05; two-tailed Fisher’s exact test). PA/OA denotes a 4-day treatment with palmitic acid followed by a 4-days treatment with oleic acid, followed by 14-day with no fatty acid. i, BLI quantification of FACS-sorted and serially-transplanted PT populations (as indicated) from SCC-25 primary recipients. BLI signals are shown as the normalized photon flux (n = 10 mice per group. *P = 0.03, **P = 0.05, ***P = 0.01; two-tailed t-test). j, Frequency of developed LN metastases of mice in i (*P = 0.05, two-tailed Fisher’s exact test).
Extended Data Fig. 2 Tumour cells from palm-oil-fed primary recipient mice display a prometastatic memory in secondary recipient mice in a CD36 dependent manner.
a, Schematic diagram representing the in vivo diet experiments in which OSCC that were exposed to a high-fat diet in primary NSG mice recipients were injected into secondary recipients that were maintained on a control diet. At the final point primary tumour (PT) cells were purified by FACS-SORT for molecular characterization (created with BioRender.com). b, Flow cytometry analysis of PTs from VDH-15– and SCC-25–injected primary recipients fed a high-fat or control diet, as schematized in a. Numbers indicate [CD44brightCD36bright] and [CD44brightCD36dim] populations in the represented gate, expressed as percentages of total DAPI–GFP+Lin– cells. Samples are representative of n = 3 independent experiments. c, d, BLI tumour monitoring of secondary recipients injected with VDH-15–derived cells from primary recipient mice fed with palm oil–rich, olive oil–rich diet and normal diet (as control). BLI signals are shown as the normalized photon flux (n = 20 mice per group, two independent experiments. LN metastasis, P = 0.04; lung metastasis, *P = 0.04, **P = 0.007; two-tailed t-test). e, Frequency of developed LN metastases from animals in c expressed as percentage (for LN metastasis, *P = 0.003; **P = 0.02; for lung metastasis: *P = 0.01, ***P = 0.0004, two-tailed Fisher’s exact test). e, f, BLI metastasis quantification of secondary recipient mice injected with control pLKO.1 or shCD36 VDH-15- (f) and SCC-25- (g) derived cells from primary recipient mice fed with a high-fat palm oil or a control diet. BLI signals are expressed as the normalized photon flux (for VDH-15, n = 10 mice per group; for lymph node and lung met, *P = 0.04; n.s., not significant; two-tailed t-test, data are mean ± s.e.m.; for SCC-25, n = 20 mice per group, data from two independent experiments; *P = 0.03, ****P < 0.0001; two-tailed t-test, data are mean ± s.e.m.). g, Frequency of developed metastases from animals in g expressed as percentage (for lymph node met, ***P = 0.003, **P = 0.0004; for lung met, ****P < 0.0001, two-tailed Fisher’s exact test). h, Ex vivo BLI lung metastasis monitoring from secondary recipient mice injected with control pLKO.1 or shCD36 melanoma-derived cells from primary recipient mice fed with a high-fat palm oil or a control diet. Pictures are representative of n = 5 mice per group. i, BLI metastasis quantification of animals in i. BLI signals are expressed as the normalized photon flux. (***P = 0.0009, ****P <0.0001, two-tailed t-test, data are mean ± s.e.m.).
Extended Data Fig. 3 CD36 blocking metastatic assays and assessment of the impact of PA on the OSCC chromatin landscape.
a, Frequency of developed lung metastasis from secondary mice injected with SCC-25–derived from primary recipient mice fed a normal or palm oil–rich diet; Primary recipients were treated with an anti-CD36 neutralizing antibody (JC63.1) or a control isotype (IgA). Data is expressed as the percentages. (n = 13 mice per group; *P = 0.01, *P = 0.03, two-tailed Fisher’s exact test). b, c, ex vivo BLI imaging and quantification of lung metastasis from VDH-15 secondary recipients injected with a doxycycline-inducible shCD36 (shCD36 #23 or #76) VDH-15–derived tumour cells from primary recipient mice fed a normal or palm oil–rich diet. The secondary mice were untreated or continuously doxycycline-treated. (Data are mean ± s.e.m. Images are representative of n = 10 mice per group; *P = 0.01; **P = 0.001; ****P < 0.0001; two-tailed t-test). d, e, TSS-centred heat maps of d) H3K4me1 and e) H3K27me3 non-memory peaks for SCC-25 pLKO.1 14D Untreated and 14D post-PA conditions. f, TSS-centred density plot showing the signal of three histone marks of interest (H3K4me3, H3K4me1 and H3K27me3) in the genomic areas where H3K4me3 memory peaks were identified. g, ChIP–seq signal TSS-centred distribution plots of H3K4me1 and H3K27ac histone marks used to map enhancer regions in SCC-25 pLKO.1 cells, both at 4D (upper panel) and 14D (bottom panel) time points for Untreated and PA-treated conditions. A total of 2,183 enhancers were mapped. h, Volcano plots showing the enhancer regions displaying differential transcription of eRNAs (FC>+/-1.6, P<0.05) at 4D PA (upper panel) or 14D post-PA (bottom panel) according to our Pol II travelling ratio analysis (PRO-seq). Up-/down-regulated eRNAs are coloured in red and blue respectively. i, PCA plots of 4D (left) and 14D (right) Untreated/PA-treated H3K9me3 ChIP–seq sSCC-25 pLKO.1 samples. j, Images showing representative H3K9me3 neural-related peaks (NRXN3 and GABRG3 genes) lost in 4D PA and 14D post-PA SCC-25 pLKO.1 cells, as compared to control samples.
Extended Data Fig. 4 Surveying in vitro and in vivo histone methylation PA-driven changes in OSCC cells.
a, PCA plots of 4D/14D Untreated/PA-treated H3K4me3 (left), H3K4me1 (centre) and H3K27me3 (right) ChIP–seq samples of CTRL pLKO.1 SCC-25 cells. On the side, PCA plot of 14D Untreated/PA-treated H3K4me1 ChIP–seq samples of CTRL pLKO.1 VDH-15 cells. b, PCA plots of 14D Untreated/OA-treated H3K4me3 ChIP–seq samples of CTRL pLKO.1 SCC-25 (left)/VDH-15 (right) cells. The bottom figures show representative H3K4me3 peaks (NEFM and CHRDL2 genes) at 14D post-OA for CTRL pLKO.1 SCC-25/VDH-15 cells. c, H3K4me3 ChIP–seq from secondary (2ary) primary tumours (PTs) of CTRL pLKO.1 VDH-15 cells upon in vivo exposure to Control (CTRL)/Palm oil (PALM)-enriched diets. On the bottom, a table displaying the number of total, differentially up-/down-regulated H3K4me3 peaks identified when comparing CTRL and PALM diet samples. d, Left, TSS-centred heat maps of H3K4me3 memory and non-memory representative peaks for 2ary CTRL/PALM PTs; middle plot, TSS-centred density plot showing the H3K4me3 signal in differential peaks for both conditions assessed; right, representative in vivo memory peaks (FABP3, DRGX and CNTFR genes) for both CTRL/PALM tumours (Diff.Bind FDR≤0.1). e, Venn diagrams showing the overlap between genes harboring total (non-differential) H3K4me3 ChIP–seq peaks for VDH-15 pLKO.1 (left) Untreated 14D and CTRL diet 2ary recipient in vivo samples or (right) 14D post-PA and PALM diet 2ary recipient in vivo conditions. f, Venn diagrams showing the overlap between differential H3K4me3 ChIP–seq peaks-bearing genes for VDH-15 pLKO.1 14D post-PA and PALM diet 2ary recipient in vivo conditions. (a,b: a representation factor >/<1 indicates a higher/lower enrichment than expected by chance in gene overlap; P-Values were calculated using a hypergeometric test). g, h, Bar plots showing the top biological processes GO terms built by the g) common or h) unique genes indicated in f). Unique refers to the PALM diet 2ary recipient in vivo condition. Neural-related terms are highlighted in purple.
Extended Data Fig. 5 Characterizing PA-triggered epigenetic and transcriptional changes and their correlation.
a, Left, PCA plot of 4D/14D Untreated/PA-treated H3K27ac ChIP–seq samples of SCC-25 pLKO.1 cells; middle, TSS-centred density plot showing the H3K27ac signal of all differential peaks in 4D/14D PA samples as compared to their control counterparts; right, Image showing representative H3K27ac peaks (ANGPTL4 gene) gained upon 4D PA treatment in SCC-25 pLKO.1 cells, as compared to the corresponding 14D samples. b, Left, plot displaying the three main clusters (0, 1 and 2) detected upon scRNA-seq data t-SNE analysis of CTRL pLKO.1 SCC-25 cells after 4D PA exposure. On the side, trajectory plot displaying the predicted Cluster distribution of 4D Untreated/PA-treated individual cells; right, trajectory plot showing the distribution of 4D Untreated (blue)/PA-treated (red) cells and their corresponding clusters. c, t-SNE plot showing distribution of cells enriched in the 4D PA transcriptional signature; on the side, trajectory plot displaying the predicted distribution of the PA response score of each cell analyzed; right, bar plot showing the quantification of the Cluster distribution of 4D Untreated/PA-treated cells shown as the proportion of total cells per condition. d, Cell cycle Analysis using Propidium Iodide; PI staining. Representative FACS plots displaying the PI cell cycle profiles of both 4/14D Untreated/ PA-treated cells. e, Box plots showing the mRNA expression detected by RNA-seq in Untreated and PA-treated SCC-25 pLKO.1 cells at 4D /14D in those regions displaying H3K4me3 PA-driven changes (UP/DOWN in 4D/14D PA box plots ****P<0.0001 of a two-tailed t-test).
Extended Data Fig. 6 Tumour cells with a Palm Diet-induced metastatic memory display a neural-related signature.
a, Principal component analysis (PCA) from microarray data of secondary (2ary) recipient-primary tumours (PTs)-sorted [CD36brightCD44bright] or [CD36dimCD44bright]. Diets are indicated as (B) primary recipient mice fed a normal (red), olive oil–rich (green) or palm oil–rich (blue) diet, and secondary mice fed with a normal diet and (A) primary mice fed with a normal diet and secondary mice fed with a palm oil–rich diet (fuchsia). The axis shows the percent variability covered by each of the represented components. b, Heat map displaying the DGE levels of 2ary VDH-15–derived PTs with a palm oil memory. CD36b, [CD36brightCD44bright] and CD36d, [CD36dimCD44bright]. c, GO and Gene Set Enrichment (GSEA) analysis showing top categories from biological processes that are upregulated in palm oil diet–memory from [CD36brightCD44bright]-sorted populations from secondary PTs. For GO analysis, FC > 1.5, P < 0.05). d, Principal component analysis (PCA) from microarray data of secondary recipient-PTs control pLKO.1– or shCD36 sorted [CD36brightCD44bright] or [CD36dimCD44bright], ±palm oil memory. e, Gene Set Enrichment (GSEA) analysis showing top categories from biological processes that were upregulated in palm oil diet–memory from [CD36brightCD44bright]-sorted populations from secondary PTs. f, g, Principal component analysis (PCA) and heatmap plot from the microarray data of cells FACS-sorted from secondary PTs from control (pLKO.1) or shCD36 melanoma ± palm oil memory. In f, the axis shows the percent variability covered by each of the represented components. The heat-map plot (g) shows the DEGs in control (pLKO.1)–palm oil memory PTs as compared to control–normal diet and their correspondence with the gene expression levels from the shCD36–palm oil. h, GO and GSEA analysis showing the top biological process categories that were upregulated by a palm oil memory of [GFP+]-sorted populations from secondary recipient, melanoma-derived PTs, as analysed in RNA microarrays. For GO analysis, FC > 1.5, P < 0.05.
Extended Data Fig. 7 Impact of chemical sympathectomy on metastasis and identification of EGR2 and galanin signalling in tumour cells with a Palm Diet-induced metastatic memory.
a, Diagram of the experimental setup. Secondary (2ary) recipient mice were injected on day 0 with OSCC from primary tumours (PTs) ± palm oil memory. 2ary recipients were treated with the neurotoxin 6-hydroxydopamine (6-OHDA; to induce the apoptosis of dopaminergic neurons) or vehicle on days –6, –3 and +3 relative to the day of injection. b, Frequency of developed lung metastasis from animals treated as shown in a, expressed as the percentages (*P = 0.03, **P = 0.001; two-tailed Fisher’s exact test). c, Tyrosine hydroxylase (TH) immunofluorescence analysis of VDH-15 primary tumours (PTs) from secondary recipients ± palm oil memory after vehicle or 6-OHDA treatment. Nuclei are stained with DAPI. KT-14, cytokeratin 14. Note that 6-OHDA-lesioned-tumours display a marked reduction in the expression level of TH. Images are representative of n = 3 biological replicates per group. d, Top common predicted binding site motifs in promoter regions of the co-regulated neural-related genes in SCC-25 or VDH-15 primary tumours with a palm memory. Z-scores and P values are shown for each cell line. e, Integrative gene set enrichment analysis (GSEA) from PTs of secondary recipients SCC-25–, VDH-15– or melanoma tumour–derived cells ± palm oil memory. The graph shows the biological processes enrichment in palm oil memory compared to control diet. On the right side, detail of the neuropeptide signalling pathway–associated gene-set assayed by integrative GSEA showing galanin (GAL) as the top significantly represented gene in palm oil memory tumours. f, g, PCA plots of differentially bound signal (DBS) regions detected for 4D Untreated/PA-treated EGR2 ChIP–seq samples in f) SCC-25 pLKO.1 and g) SCC-25 CD36-KD cells. h, Heat maps showing the EGR2 DBS regions (differential peaks) detected for 4D SCC-25 pLKO.1 (left) and SCC-25 CD36 KD (right) shown in f) and g) respectively (FDR<0.05).
Extended Data Fig. 8 Modulating EGR2 expression to determine its relevance in modulating the OSCC epigenome, transcriptome and metastatic potential.
a, PCA plots of differentially bound signal (DBS) regions detected for 4D (left) and 14D (right) Untreated/PA-treated H3K4me3 ChIP–seq samples in SCC-25 EGR2-KD cells. b, Bar plots showing the top biological processes GO terms uniquely down-regulated in 4D PA (upper panel) and 14D post-PA (bottom panel) H3K4me3 ChIP–seq SCC-25 EGR2-KD samples. Neural-related terms are highlighted in purple. c, Venn diagrams showing the overlap between in vivo proneural-induced gene signatures in secondary OSCC primary tumours with a palm oil memory and the knockdowns, as indicated. d, Gene set enrichment analysis (GSEA) showing the negative enrichment of biological processes in secondary PTs derived from SCC-25–shEGR2/PALM vs pLKO.1/control diet or shGAL/PALM vs pLKO.1/control diet. e, BLI metastasis quantification of lymph node (LN) and lung of secondary recipient mice injected with primary recipient SCC-25 cells, derived from control (pLKO.1), shRNA-knockdown of EGR2 (shEGR2 #38_9 and shEGR2 #40_9) or shRNA-knockdown of GAL (shGAL #74_4 shRNAs) ±palm oil memory (pLKO.1/control diet, n = 22; pLKO.1/PALM, n = 21; shEGR2/control diet and shEGR2/PALM, n = 20; shGAL/control diet and shGAL/PALM, n = 10; in the knockdowns, n reflects number of mice per shRNA used; *P = 0.01, two-tailed t-test. Data are the mean ±s.e.m.).
Extended Data Fig. 9 Metastatic functional assay after systemic inhibition of galanin signallling through intraperitoneal administration of the pan-galanin receptor inhibitor galantide (M15).
a, Experimental setup of the experiment for galanin receptor inhibition. b, c, Ex vivo BLI lung metastasis (b) and frequency of lung metastasis development (c), expressed as the percentages, from secondary (2ary) recipient mice injected with VDH-15 cells derived from primary recipient mice fed a palm oil–rich diet or a control diet. 2ary recipients were treated with the galanin receptor antagonist galantide (M15) or vehicle as explained in (a) (vehicle-treated, n = 12 mice; M15-treated, n = 15 mice. *P = 0.03; *P = 0.04; two-tailed Fisher’s exact test).
Extended Data Fig. 10 Elucidating the role of COMPASS methyltransferases in PA-driven OSCC chromatin changes and metastatic abilities.
a, b, Western blot analysis of the COMPASS family of methyltransferases (Set1A/B and MLL1/2) in VDH-15 or SCC-25 pLKO.1 cells that were Untreated, a) 4D after PA treatment or b) 14D after removal of PA. Cells were infected with a non-targeting (nt)-shRNA (pLKO.1). Hsp90 was used as a loading control. c, d, PCA plots displaying ChIP–seq for c) MLL1or d) MLL2 from untreated or 14D post-PA VDH-15 pLKO.1 cells. e, Representative peaks from MLL1/MLL2 ChIP–seq of MLL1/MLL2-regulated genes (HOX gene cluster) and non-regulated genes (neural-related CHRDL2 gene) for all conditions tested (Untreated/4D PA and 14D post-PA pLKO.1 VDH-15 cells). f, PCA plot showing 14D Untreated/PA-treated H3K4me3 ChIP–seq samples of CTRL pLKO.1 and Set1A #42 KD VDH-15 cells. g, GO analysis of PA-treated Set1A KD VDH-15 H3K4me3 ChIP–seq samples. The plot shows the top biological processes GO terms significantly down-regulated in Set1A KD #42 samples 14D post-PA exposure when compared to the 14D Untreated samples (Diff. Bind FDR≤0.05). On the side, H3K4me3 peaks of Set1A-regulated neural genes (CHRDL1 and GRIP2 genes). H3K4me3 representative examples are shown at 4/14D time points for all assessed conditions (CTRL pLKO.1 and Set1A #42 KD VDH-15 cells). h, Left, secondary (2ary) recipient orthotopic injections of CTRL pLKO.1 and Set1A #42 KD VDH-15 cells upon in vivo exposure to Control/Palm-oil enriched diets in primary recipient mice. The frequency of developed LNmets (at the end-point of the experiment is shown for all conditions (n= 10 mice/group; LNmets: CTRL pLKO.1 CTRL Diet vs CTRL pLKO.1 PALM Diet n.s= 0.065, CTRL pLKO.1 PALM Diet vs Set1A KD PALM Diet n.s= 0.091; n.s. is not significant Fisher’s exact test). Right, frequency of developed LNmets at the end-point of the experiment in 1ary recipient orthotopic injections of CTRL pLKO.1/Set1A KD #07 VDH-15 cells 14D post-PA in vitro exposure (n= 20 mice/group; LNmets: CTRL pLKO.1 UNTR vs CTRL pLKO.1 14D post-PA *P=0.038, CTRL pLKO.1 14D post-PA vs Set1A KD 14D post-PA *P=0.031 of Fisher’s exact test). For gel source data, see Supplementary Figure 1.
Extended Data Fig. 11 Tumour cells with a Palm Diet-induced metastatic memory alter the tumour-associated stroma in a CD36-dependent manner.
a, Flow cytometry analysis of SCC-25-primary tumours (PTs) from secondary (2ary) recipients ± palm oil memory. [GFP–/CD31–/CD45–] tumour stroma–selected cells were purified and processed to perform RNA microarrays. Data are representative of n = 3 independent experiments. Numbers indicate the population in the represented gate, expressed as the percentages. b, Principal component analysis (PCA) of the microarray data of VDH-15–associated bulk stroma purified from secondary (2ary) recipients. Diet samples are indicated as (B) primary recipient mice fed a normal (red), olive oil–rich (green) or palm oil–rich (blue) diet, and 2ary mice fed with a normal diet and (A) primary mice fed with a normal diet and 2ary mice fed with a palm oil–rich diet (fuchsia). The axis shows the percent variability covered by each of the represented components. c, Principal component analysis (PCA) from microarray data of 2ary recipient tumour–associated stroma derived from injection in primary recipients of control (pLKO.1)– or shCD36–VDH-15 or SCC-25 cells, as indicated by the colour-code. The axis shows the percent variability covered by each of the represented components. d, Volcano plots for the differential gene expression microarray analysis and GO analysis from a purified tumour stroma from 2ary recipient mice injected with OSCC from PTs ± palm oil memory. Volcano plots show the up- or downregulated genes in control–palm oil memory tumours compared to control–normal diet tumours (top) and the expression of these genes in the shCD36–palm oil (lower plot). Data from n = 2 biological replicates; fold-change, FC > 2, P < 0.01. The GO analysis shows the top biological processes categories upregulated in the palm oil memory tumour stroma, FC > 2, P < 0.05. (Integrated analysis, derived from VDH-15 or SCC-25 cells). e, Neural–mouse stromal signature induced in 2ary recipients after orthotopic injection of primary recipient–PT cells, derived from control (pLKO.1), shRNA-CD36, shRNA-EGR2 or shRNA-GAL SCC-25 cell. Each dot in the plot represents a neural gene. f, g, GSEA showing the negative enrichment of biological processes in neural–mouse stroma associated with 2ary PTs from shEGR2/PALM or shGAL/PALM vs pLKO.1/control diet conditions.
Extended Data Fig. 12 Schwann cell development and neuron projection regeneration are processes enhanced in the stroma of palm-oil derived tumours.
a, Functional network interaction between tumour–mouse stromal palm oil (PALM)–controlled genes and the human OSCC primary tumours with a palm memory. The network graph shows the co-regulated functional nodes of interaction between the two compartments (fold-change, FC > 1.5; P value < 0.05). b, c, 10X single-cell (sc) RNA-seq clustering analysis of the tumour-associated stroma purified from secondary (2ary) oral SCC-25 PTs ±palm oil memory. The principal component UMAP plot is shown in which the specific cell types have been annotated to each respective cluster. d, Overlap analysis represented by Venn diagrams showing the intersection between bulk-stroma palm memory-controlled genes and 10X single-cell (sc) RNA-seq clusters from SCC-25–derived stromal cells in 2ary recipients. Representation factor (RF) and P values are shown for the overlap. Hypergeometric test; estimated number of protein-encoding genes = 25,000.
Extended Data Fig. 13 ECM components related to tumour-associated Schwann cells are increased in palm-oil derived tumours and correlate with CD36+ metastatic signature.
a, b, c, Integrated UMAP cluster visualization, annotated for cell types, of 10X single-cell (sc) RNA-seq data of the tumour-associated stroma purified from secondary (2ary) oral SCC-25 primary tumours (PTs) ±palm oil memory. The UMAP plot shows the expression level and cluster distribution of selected gene markers relative to glial cells and progenitor glia a), specialized extracellular matrix constituent (ECM) b) and nerve injury/ nerve regenerative processes c). d, Regression analysis showing the correlation between the long-lasting palm oil–related signature of the OSCC tumours (derived from SCC-25 or VDH-15 cells) or melanoma (501-mel), and the expression of markers related with perineuronal nets (GO:0072534). R-squared coefficients (R) and P values are shown for each analysis. e, Immunofluorescence analysis of PTs from 2ary recipient mice injected with VDH-15–derived cells from primary recipient ± palm oil memory. Cytokeratin-14 (KT-14) is shown as epithelial marker of OSCCs, and the specialized ECM markers Hyaluronidase 1 (Has1), Tenascin R (TNR) and the glial/Schwann marker s100 are shown as tumour stroma markers. Yellow arrows indicate areas of Has1 positive cells. Dashed lines delimitate the interface between the tumour and the tumour-associated stroma. f, Magnification from b) (palm diet memory condition). Note the double labelling of TNR and S100 in the close proximity of the tumour front. Images are representative of n = 4 biological replicates.
Extended Data Fig. 14 Spatial transcriptomic analysis of secondary recipient OSCC primary tumours.
a, Images showing Haematoxylin-Eosin (H&E) staining of primary tumours (PTs) from secondary (2ary) recipients injected with control (pLKO.1) and shCD36 SCC-25 cells derived from primary recipient mice fed a normal diet or a palm oil–rich diet. e, Spatial transcriptomics analysis of tumours in a) showing the proportion content and spatial distribution of mouse and human transcriptome per spot, the analysis stratification of the tissue as healthy/tumour invasive front/tumour and the proportion content and spatial distribution of the tumour-associated Schwann cells. c, Quantitative analysis of the proportional content of tumour-associated Schwann cells within each compartment (healthy/tumour front/tumour) of the PTs from 2ary recipients injected with control (pLKO.1) and shCD36 SCC-25 cells derived from primary recipient mice fed a normal diet or a palm oil–rich diet.
Extended Data Fig. 15 Immunofluorescent analysis of secondary recipient OSCC primary tumours.
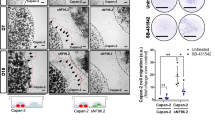
a, b. e, Immunofluorescence analysis and quantification of primary tumours (PTs) from secondary (2ary) recipient mice injected with VDH-15– or SCC-25-derived cells, as indicated, derived from primary recipient mice fed a normal diet, a palm oil–rich or an olive oil–rich diet. Cytokeratin-14 (KT-14) is shown as an epithelial marker of OSCC, and the glial/Schwann markers s100 or GAP43 as a tumour stroma marker. Nuclei are stained with DAPI. Images are representative of n = 3 independent experiments. c, Graphs showing the values of integrated density of (a, b). (n = 3 biological replicates per group; *P = 0.05, ****P < 0.0001; two-tailed t-test). f, Immunofluorescence analysis of PTs from M15-treated or vehicle-treated mice ± palm oil memory. The expression of cytokeratin-14 (KT-14) is shown as an epithelial marker of OSCC, and the glial Schwann cell marker s100 is shown in the tumour-associated stroma. Nuclei are stained with DAPI. Note that the increased expression of s100 in the stroma of palm memory tumours is slightly reduced in the case of M15-treated palm condition.
Extended Data Fig. 16 Enzymatic digestion through chondroitinase ABC (chABC) prevents the increase in metastatic competency induced by palmitic acid in OSCC cells.
a, Western blot analysis of total protein extracted from in vitro cultured VDH-15 cells infected with the lentiviral LV-chondroitinase ABC (ch-ABC) vector, showing the overexpression of the chondroitinase ABC enzyme in the infected cells (n = 2 replicates per group). b, c, Immunofluorescence analysis of primary tumours from secondary recipient mice injected with wild-type VDH-15 (WT) or LV-chondroitinase ABC enzyme (chABC)–derived cells from primary recipient mice fed a normal diet or a palm oil–rich diet. Cytokeratin-14 (KT-14) is shown as epithelial marker of OSCCs, and the specialized ECM markers Versican (Vcan), Tenascin R (TNR) and the glial/Schwann marker s100 as tumour stroma markers d) or collagen 5A1 (Col5A1) as ECM marker e). Nuclei are stained with DAPI. Images are representative of n = 3 biological samples.
Supplementary information
Supplementary Figure 1
Raw images of the western blot experiments presented in Extended Data Figure 10. Pictures of exposed blots for the detection of COMPASS proteins expression in SCC-25 and VDH-15 pLKO.1 cell lines are provided for both the 4 d and 14 d time points, together with information on the molecular mass ladder that was used.
Supplementary Table 1
This table is complementary and related to Fig. 1 and Extended Data Figs. 1–3. It contains data about VDH-15 and SCC-25 bioluminiscence (BLI) quantification of primary tumours, lymph node and lung metastasis and frequencies of developed primary tumours and metastasis; CD36 membrane expression by FACS profile of primary tumours and in vitro cells.
Supplementary Table 2
This table contains H3K4me3 ChIP–seq differential peaks and annotated genes as well as the corresponding GO analyses for: SCC-25 and VDH-15 pLKO.1 cells 14d after PA (14 d post-PA) samples; VDH-15 pLKO.1 secondary primary tumour samples.
Supplementary Table 3
This table contains H3K4me1 and H3K27me3 ChIP–seq as well as PRO-seq information, including: gene overlap of untreated and PA-treated conditions at both the 4 d and 14 d time points for both H3K4me1 and H3K27me3 ChIP–seq samples; H3K4me1 ChIP–seq differential peaks and annotated genes for SCC-25 and VDH-15 pLKO.1 cells 14 d post-PA; H3K27me3 ChIP–seq differential peaks and annotated genes for SCC-25 pLKO.1 cells 14 d post-PA; and total and differentially expressed eRNAs as detected by PRO-seq data analysis at both 4 d PA and 14 d post-PA for SCC-25 pLKO.1 cells.
Supplementary Table 4
This table contains information on PA-treated and OA-treated H3K4me3 ChIP–seq samples of both SCC-25 and VDH-15 pLKO.1 cells, including: GO analysis 14 d post-PA H3K4me3 ChIP–seq samples for both SCC-25 and VDH-15 cells; GO analysis 14 d post-OA for H3K4me3 ChIP–seq SCC-25 pLKO.1 samples; GO analysis of H3K4me3 ChIP–seq secondary palm-oil-enriched diet exposed primary tumours; gene overlap of H3K4me3 OA-treated and PA-treated ChIP–seq conditions at the 14 d time point, together with the comparison of total number of differentially expressed peaks 14 d post-PA/OA; gene overlap of H3K4me3 and H3K4me1 or H3K27me3 14 d post-PA ChIP–seq SCC-25 pLKO.1 samples; comparison of the total number of differentially expressed H3K4me1 peaks 14 d post-PA between SCC-25 and VDH-15 pLKO.1 samples; H3K4me3 ChIP–seq differential peaks and annotated genes for SCC-25 and VDH-15 pLKO.1 cells 14 d post-OA and their corresponding GO analyses; and a list of differentially expressed mRNAs as detected by RNA-seq at 4 d and 14 d post-PA as well as the 14 d post-PA corresponding GO analysis.
Supplementary Table 5
This table contains information on PA-treated and OA-treated H3K4me3 ChIP–seq samples of both SCC-25 and VDH-15 pLKO.1 cells, including: GO analysis 14 d post-PA H3K4me3 ChIP–seq samples for both SCC-25 and VDH-15 cells; GO analysis 14 d post-OA for H3K4me3 ChIP–seq SCC-25 pLKO.1 samples; GO analysis of H3K4me3 ChIP–seq secondary palm-oil-enriched diet exposed primary tumours; gene overlap of H3K4me3 OA-treated and PA-treated ChIP–seq conditions at the 14 d time point, together with the comparison of the total number of differentially expressed peaks 14 d post-PA/OA; gene overlap of H3K4me3 and H3K4me1 or H3K27me3 14 d post-PA ChIP–seq SCC-25 pLKO.1 samples; comparison of the total number of differentially expressed H3K4me1 peaks 14 d post-PA between SCC-25 and VDH-15 pLKO.1 samples; H3K4me3 ChIP–seq differential peaks and annotated genes for SCC-25 and VDH-15 pLKO.1 cells 14 d post-OA and their corresponding GO analyses; and a list of the differentially expressed mRNAs as detected by RNA-seq at 4 d and 14 d post-PA as well as the 14 d post-PA corresponding GO analysis.
Supplementary Table 6
This table contains microarray data, GO and GSEA, and integrative analysis from VDH-15, SCC-25 and melanoma primary tumours from secondary recipients ± palm-oil memory.
Supplementary Table 7
This table contains microarray data, GO and GSEA from melanoma (501mel) primary tumours from secondary recipient ± palm-oil memory.
Supplementary Table 8
This table is complementary and related to Extended Data Fig. 7. It contains data about VDH-15 BLI quantification of primary tumours, lymph node and lung metastasis and the frequencies of developed primary tumours and metastasis.
Supplementary Table 9
This table contains data about: transcription factor binding site analysis in the pro-neural signature of VDH-15 and SCC-25 primary tumours with a palm memory; microarray data and GSEA of SCC-25 shEGR2 and SCC-25 shGAL primary tumours ± palm-oil memory.
Supplementary Table 10
This table contains information on EGR2 ChIP–seq for SCC-25 pLKO.1 and CD36-KD samples, including a comparison of the total number of differentially expressed EGR2 peaks at 4 d PA between SCC-25 pLKO.1 and SCC-25 CD36-KD samples; a comparison of the total number of differentially expressed H3K4me3 peaks at 4 d and 14 d post-PA between SCC-25 CD36-KD samples; gene overlap of H3K4me3 SCC-25 pLKO.1 and EGR2-KD ChIP–seq samples at 4 d and 14 d post-PA, together with the corresponding GO analyses; EGR2 ChIP–seq differential peaks and annotated genes for SCC-25 pLKO.1 at 4 d PA and 14 d post-PA; H3K4me3 ChIP–seq differential peaks and annotated genes for SCC-25 EGR2-KD cells at 4 d PA and 14 d post-PA as well as their corresponding GO analyses; and H3K4me3 ChIP–seq differential peaks and annotated genes for SCC-25 CD36-KD cells at the 4 d PA time point.
Supplementary Table 11
This table is complementary and related to Extended Data Fig. 9. It contains data about VDH-15 BLI quantification of primary tumours, lymph node and lung metastasis, and frequencies of developed primary tumours and metastasis.
Supplementary Table 12
This table contains data of the correlation of EGR2 or GAL expression and the overall survival in different types of tumours.
Supplementary Table 13
This table contains data on MLL1 and MLL2 ChIP–seq experiments as well as in vivo MLL1-KD assays, including information on the total number of differential MLL1 and MLL2 ChIP–seq peaks 14 d post-PA as well as at the corresponding list of differential peaks and annotated genes for SCC-25 pLKO.1 samples; and data on VDH-15 BLI quantification for lymph node metastases 14 d post-PA exposure.
Supplementary Table 14
This table contains data regarding H3K4me3 ChIP–seq experiments together with in vivo assays involving VDH-15 Set1A-KD cells, including H3K4me3 ChIP–seq differential peak GO analysis of VDH-15 Set1A-KD cells 14 d post-PA; and BLI quantification data of VDH-15 pLKO.1 and Set1A-KD primary tumours 14 d post-PA exposure.
Supplementary Table 15
This table contains microarray data, GO and GSEA of bulk tumour-associated stroma from VDH-15 and SCC-25 secondary recipients ± palm-oil memory.
Supplementary Table 16
This table contains data relating to the 10x single-cell analysis of bulk tumour stroma from pLKO.1 and shCD36 SCC-25 secondary recipients ± palm-oil memory; overlap analysis showing the intersection between bulk-stroma palm memory-controlled genes; the 10x scRNA-seq clusters from SCC-25-derived stromal cells in secondary recipients; Palm-oil diet-ECM-related genes.
Supplementary Table 17
This table is complementary and related to Extended Data Fig. 16. It contains data about VDH-15 BLI quantification of primary tumours, lymph node and lung metastasis, and frequencies of developed primary tumours.
Supplementary Table 18
This table contains data about VDH-15 BLI quantification and frequencies of lung metastasis in secondary recipients.
Supplementary Table 19
This table contains Pearson correlation analysis between the CD36+ metastatic signature and perineuronal net gene sets in different tumour types.
Supplementary Table 20
This table contains methods information relating to shRNA sequences, antibodies and Taqman gene expression probes.
Rights and permissions
About this article
Cite this article
Pascual, G., Domínguez, D., Elosúa-Bayes, M. et al. Dietary palmitic acid promotes a prometastatic memory via Schwann cells. Nature 599, 485–490 (2021). https://doi.org/10.1038/s41586-021-04075-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04075-0
This article is cited by
-
Association of saturated fatty acids with cancer risk: a systematic review and meta-analysis
Lipids in Health and Disease (2024)
-
Lipids as mediators of cancer progression and metastasis
Nature Cancer (2024)
-
Immunosurveillance encounters cancer metabolism
EMBO Reports (2024)
-
Transient loss of Polycomb components induces an epigenetic cancer fate
Nature (2024)
-
Metabolic heterogeneity in cancer
Nature Metabolism (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.