Abstract
The majority of oncogenic drivers are intracellular proteins, thus constraining their immunotherapeutic targeting to mutated peptides (neoantigens) presented by individual human leukocyte antigen (HLA) allotypes1. However, most cancers have a modest mutational burden that is insufficient to generate responses using neoantigen-based therapies2,3. Neuroblastoma is a paediatric cancer that harbours few mutations and is instead driven by epigenetically deregulated transcriptional networks4. Here we show that the neuroblastoma immunopeptidome is enriched with peptides derived from proteins that are essential for tumourigenesis and focus on targeting the unmutated peptide QYNPIRTTF, discovered on HLA-A*24:02, which is derived from the neuroblastoma dependency gene and master transcriptional regulator PHOX2B. To target QYNPIRTTF, we developed peptide-centric chimeric antigen receptors (CARs) using a counter-panning strategy with predicted potentially cross-reactive peptides. We further hypothesized that peptide-centric CARs could recognize peptides on additional HLA allotypes when presented in a similar manner. Informed by computational modelling, we showed that PHOX2B peptide-centric CARs also recognize QYNPIRTTF presented by HLA-A*23:01 and the highly divergent HLA-B*14:02. Finally, we demonstrated potent and specific killing of neuroblastoma cells expressing these HLAs in vitro and complete tumour regression in mice. These data suggest that peptide-centric CARs have the potential to vastly expand the pool of immunotherapeutic targets to include non-immunogenic intracellular oncoproteins and widen the population of patients who would benefit from such therapy by breaking conventional HLA restriction.
Similar content being viewed by others
Main
The curative potential of CAR T cell-based cancer immunotherapies has been established in leukaemias, but their application in solid tumours has been limited by a paucity of known tumour-specific membrane proteins5. Although membrane proteins represent up to a quarter of the proteome, only a fraction of these are expressed specifically on tumour cells and not on normal tissues, and a smaller proportion are essential to tumour homeostasis6. Rather, most cancer driver proteins reside in the cytoplasm or nucleus of the cell, where they are accessible to the immune system only through presentation of peptides on the major histocompatibility complex (MHC).
MHC class I proteins, encoded by the highly polymorphic human leukocyte antigen (HLA)-A, -B and -C genes, present a snapshot of the intracellular proteome on the cell surface (immunopeptidome) where T cells surveil the peptide–MHC complexes (pMHC) for antigens derived from foreign pathogens. T cell recognition of mutation-derived pMHC neoantigens as non-self is the basis of curative responses achieved through immune checkpoint blockade and complete remissions using adoptive transfer of tumour-infiltrating lymphocytes (TILs)7. Nonetheless, only about 5% of these neoantigens are predicted to bind a given HLA allotype8, and just 1.6% of neoantigens are reported to be immunogenic3. Subclonal mutations and downregulation of mutated non-essential genes further constrain the pool of therapeutically relevant neoantigens, necessitating a mutational threshold for effective neoantigen-based therapies that is not surpassed in most cancers9. Tumour cells also present a wide range of unmutated self-peptides on MHC10 molecules, but these are largely immunogenically silent owing to negative thymic selection of T cells. We hypothesized that a subset of the immunopeptidome consists of tumour-specific peptides derived from essential oncoproteins and that these can be targeted using synthetic peptide-centric CARs (PC-CARs).
Peptides presented in the MHC groove make up only a small fraction of the extracellular pMHC molecular surface. The typical 8–14mer peptide presented on MHC class I comprises only around 2–3% of the amino acids in the pMHC complex and is spatially confined within the adjacent α-helices of the MHC groove, thus posing major challenges for engineering peptide-specific single-chain antibody variable fragment (scFv) binders11. Furthermore, cross-reactivity of engineered receptors with peptides of biophysically similar molecular surfaces presented in normal tissues have resulted in significant toxicity and death12,13.
Neuroblastoma is a childhood cancer derived from tissue of the developing sympathetic nervous system and is often lethal despite intensive cytotoxic therapy14. These tumours are low in mutational burden15 and MHC expression16, making neuroblastoma both a challenging tumour to target with MHC-based immunotherapies and an ideal model for addressing the major problems currently hindering the wider advancement of cancer immunotherapies. As a tumour derived from neural crest progenitor cells, neuroblastomas express a set of core-regulatory circuit (CRC) transcription factors involved in maintaining cell fate, metabolism, migration, epigenetic states, growth and proliferation4. These genes are epigenetically silenced upon terminal differentiation of normal neural tissues, but these developmental pathways are aberrantly hyperactivated in neuroblastoma. Here we present the discovery of recurring lineage-restricted oncoproteins presented on MHC, focusing on immunotherapeutic targeting of the neuroblastoma CRC master regulator PHOX2B using PC-CARs.
Identification of tumour-specific antigens
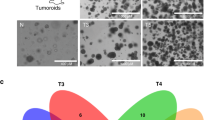
First, we surveyed the landscape of peptides accessible to T cells by performing MHC capture, peptide elution and liquid chromatography with tandem mass spectrometry17 (immunopeptidomics) on eight neuroblastoma cell-derived xenografts (CDX) and patient-derived xenografts (PDX) showing a wide range of MHC expression and also encompassing the array of rare recurring mutations found in high-risk neuroblastoma18,19 (Fig. 1a, Extended Data Table 1a). We identified a total of 7,608 peptides from 8 tumours (1% false discovery rate (FDR); Supplementary Table 1), finding none of the 4,105 potential 8–14mer neoantigens imputed from tumour mutational data in the immunopeptidome, consistent with expected rates of neoantigen presentation and limited detection using ligandomics in low-mutational tumours20. We first filtered the 7,608 peptides to select HLA binders with sufficient affinity to act as T cell epitopes using a predicted peptide–MHC (pMHC) half-maximal inhibitory concentration (IC50) threshold of 500 nM, yielding 2,286 predicted strong binders. We then filtered for peptides derived from differentially expressed parent genes as determined from RNA-sequencing (RNA-seq) data from 153 neuroblastoma tumours compared with 1,641 normal tissues (from the TARGET21 and GTEx17 databases, respectively), resulting in 61 peptides derived from genes with mRNA expression one log-fold higher in tumour for each normal tissue (P < 0.01). Finally, we filtered the remaining tumour peptides against a database of MHC peptides empirically characterized on 190 normal tissues22, removing any peptide with a parent gene that is represented in the normal tissue immunopeptidome. While this last step does not absolutely exclude potential presentation of a peptide in normal tissues, it allowed us to narrow our list to 13 peptides derived from 9 unique genes expressed in neuroblastoma that have not been previously detected in any normal tissue.
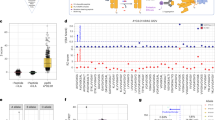
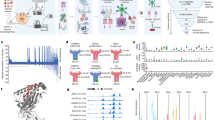
a, Summary of tumour-antigen discovery and CAR-engineering workflow: (1) integrated genomics and immunopeptidomics process; (2) target validation; (3) CAR engineering; and (4) cross-HLA tumour killing. b, Computational filtering of 9,117 peptide instances identified by immunopeptidomics in primary tumours (1% FDR) resulted in 56 neuroblastoma-specific peptides (33 unique peptides) derived from 29 distinct proteins. c, Primary neuroblastoma tumour immunopeptidome compared with 190 normal tissues. Each point on the x-axis represents one of 5,832 unique peptides identified in primary tumours, with the proportion of neuroblastoma tumours presenting a given peptide annotated above axis in dark blue and the proportion of normal tissue expressing the identical peptide below the axis in light blue. The green horizontal bar indicates 1,492 peptides not previously observed in the normal tissue immunopeptidome. Parent genes from neuroblastoma-specific peptides resulted in the top two GO enrichment terms: noradrenergic neuron differentiation and sympathetic nervous system development. The arrow denotes 351 recurring peptides presented in neuroblastoma that were not previously detected in normal tissues. d, Five antigens differentially expressed in PDX and primary tumours and further prioritized for analysis, HLA allele frequency, relative peptide abundance (percentile rank annotated below pMHC), predicted pMHC-binding affinity, and relevance to neuroblastoma tumourigenesis. e, PHOX2B expression in RNA-seq data from 153 neuroblastoma tumours versus 1,641 normal tissues in GTEx. PHOX2B expression is restricted to tumours, compared with the immunotherapy target HER2 and the neuroblastoma chemotherapy target TOP1 (note differences in the y-axis scales). FPKM, fragments per kilobase of transcript per million mapped reads. f, Crystal structure of PHOX2B 9mer QYNPIRTTF (red) refolded with HLA-A*24:02 (grey). g, Chromatin immunoprecipitation followed by DNA sequencing (ChIP-seq) of neuroblastoma tissue shows binding of all CRC proteins at the PHOX2B locus and association with a H3K27Ac super-enhancer mark. h, RNA-seq of fetal tissue shows that PHOX2B is expressed in early development and downregulated before birth across seven tissues. Created with BioRender.com.
We then performed immunopeptidomics on eight high-risk diagnostic neuroblastoma primary tumours, focusing on HLA-A*02:01 and HLA-A*24:02 allotypes (Extended Data Table 1a, Supplementary Table 1). Using the same filtering as described for xenograft tumours, we identified 56 peptides (33 unique peptides derived from 29 unique proteins) with strong HLA-binding affinity derived from differentially expressed genes in neuroblastoma and not previously detected in the benign tissue ligandome (Fig. 1b, Extended Data Fig. 1). We confirmed the presence of 7 of the 13 peptides from the xenografts in primary tumours, suggesting that CDX and PDX tumours can be a predictive model for primary tumour ligandomes. Notably, the most enriched gene ontologies of the peptide parent genes not previously observed in the normal-tissue ligandomic dataset were: noradrenergic neuron differentiation (P = 3.42 × 10−4, FDR = 0.0389) and sympathetic nervous system development (P = 2.95 × 10−5, FDR = 0.00665; Fig. 1c, green bar). These findings highlight the distinctiveness of the neuroblastoma ligandome and suggest that MHC-presented peptides are enriched for those derived from aberrantly expressed lineage-restricted genes.
To select those peptides with the highest potential as putative immunotherapeutic targets, we prioritized peptides on the basis of pMHC binding affinity, HLA allele frequency, degree of differential expression of parent genes, relative abundance on MHC as compared to other peptides, recurrence across multiple neuroblastoma tumours and relevance to neuroblastoma biology based on the published literature23,24,25. One peptide each from CHRNA3, GFRA2, HMX1, IGFBPL1, PHOX2B and TH were selected for preclinical development (Fig. 1d, e; Supplementary Table 1). The presence of these peptides in tumours was validated by performing liquid chromatography–mass spectrometry (LC–MS/MS) on synthetic peptides (Extended Data Fig. 2), yielding complete concordance across b and y ions. In addition, we validated the peptide binding to predicted HLA alleles by refolding the pMHC complex and solving the crystal structure of one PHOX2B peptide–HLA-A*24:02 (PDB ID 7MJA) and three IGFBPL1 peptide–HLA-A*02:01 (PDB IDs 7MJ6, 7MJ67 and 7MJ8) protein complexes (Fig. 1f, Extended Data Fig. 3b). Of note, we detected three unique IGFBPL1 peptides as a 9mer, 11mer and 12mer (all with the same core 9mer amino acids) with distinct thermal properties (Extended Data Fig. 3), underscoring the existence of multiple peptides with distinct conformations arising from the same region of a protein and presenting multiple opportunities for therapeutic targeting26.
We then inferred the ability of a tumour to evade the immune response via downregulation of target genes by examining the binding of neuroblastoma CRC transcription factors (MYCN, ASCL1, HAND2,ISL1, PHOX2B, GATA3 and TBX2; Extended Data Fig. 4a) to the parent gene locus of the prioritized antigens4. We found that all six CRC proteins bind regulatory elements at each parent gene locus within H3K27Ac super-enhancer elements (Fig. 1g, Extended Data Fig. 4b), suggesting that transcriptional redundancy and dependency should mitigate the risk of antigen loss in response to immunotherapy owing to downregulation of the parent gene. In addition, we found peptides from each of the six CRC proteins represented in the neuroblastomaimmunopeptidome (Extended Data Table 1b). Indeed, the most significantly enriched gene groups in the immunopeptidome are nucleic acid-binding proteins (P = 9.5 × 10−17; Extended Data Fig. 5c). Finally, we interrogated the dynamics of gene expression during development using temporal transcriptomic data27. Consistent with its function in orchestrating neural crest progenitor development28, PHOX2B is expressed exclusively during fetal development and is completely silenced in normal tissues before birth (Fig. 1h), as are IGFBPL1, TH and CHRNA3 (Extended Data Fig. 6). PHOX2B is a key CRC protein that is among the most specifically expressed genes in neuroblastoma29. PHOX2B expression is not detected in normal tissue, unlike many solid tumour immunotherapy targets, including HER2, and chemotherapy targets in neuroblastoma camptothecin, including TOP1, each of which exhibit significant normal tissue expression (Fig. 1e). PHOX2B expression is routinely used in neuroblastoma diagnostic assays30, is one of two highly penetrant neuroblastoma susceptibility genes31, and is the third most significant dependency in neuroblastoma, as reported in DepMap32. Together, these data suggest that PHOX2B is a highly specific tumour antigen in neuroblastoma and an ideal candidate for therapeutic targeting.
Before developing an immunotherapeutic construct targeting the PHOX2B peptide, we validated that low MHC expression in neuroblastoma16 did not prohibit T cell engagement and activation using an influenza antigen model (experimental details in Extended Data Fig. 7).
PC-CAR T cell engineering for PHOX2B
Owing to the low immunogenicity of self-antigens, we pursued development of scFv-based CARs rather than engineered T cell receptors (TCRs) for PHOX2B after no high affinity TCRs were identified in multiple screens (Extended Data Fig. 8). We reasoned that immunogenicity could be induced to otherwise immunogenically inert pMHCs using synthetic, peptide-centric receptors. Our first generation of pMHC-directed CARs showed significant cross-reactivity to the MHC, which we were able to abrogate using saturation mutagenesis techniques that also decreased binding affinity (Extended Data Figs. 9–11).
To screen for PHOX2B peptide-specific clones, we next adapted the retained display33 (ReD) system, a protein-display platform that enables the flow-cytometric selection of pMHC-binding scFvs in permeabilized bacterial cells, with a scFv library containing more than 1011 variants. Clones that demonstrated apparent selectivity by flow cytometry were further tested against a panel of 95 unrelated peptides and four highly similar peptides presented on the same HLA allotype to select for clones with the highest selectivity (for example, selective clone 10LH, shown in Fig. 2a), resulting in 25 scFv binders for screening in CAR T constructs. To address cross-reactivity with pMHC in normal tissues, we developed an algorithm to predict potential selective cross-reactive antigen presentation (sCRAP; https://marisshiny.research.chop.edu/sCRAP/) on the same HLA allotype (Extended Data Fig. 12), enabling pre-emptive selectivity filtering in early stages of scFv screening without the need of a prior receptor or alanine scan34. We benchmarked the sCRAP algorithm by testing its ability to predict the cross-reactivity of the MAGE-A3 peptide presented on HLA-A*01:01, whose targeting using an affinity-enhanced TCR previously resulted in the fatal cross-reaction with another peptide derived from the TITIN protein presented on HLA-A*01:01 in myocardial tissues12. We predicted the cross-reactivity of MAGE-A3 with the TITIN peptide as the 4th-ranked prediction out of 1,143,861 potential self-peptides presented in heart tissue (Extended Data Fig. 13).
a, Ranked binding affinity of 10LH scFv to PHOX2B (blue) and a panel of 95 peptides presented on HLA-A*24:02 peptides (orange) demonstrate high target binding and negligible binding to HLA-A*24:02 pMHCs. b, Cross-reactivity algorithm identifies CAR constructs with significant off-target binding and informs prioritization of highly selective receptors (selective receptors marked with arrow). Peptide score represents the predicted cross-reactivity based on the amino acid sequences of normal-tissue peptides; overall score is calculated on the basis of peptide score, binding affinity and normal tissue expression; T, peptides reported in the normal tissue immunopeptidome; F, peptides absent in normal immunopeptidome. ND, not determined. c, Example counterstaining of top CAR clones with target (x-axis) and off-target (y-axis) peptides on HLA-A*24:02 reveals selective target binding in 10LH and 302LH constructs. d, Left, flow cytometry plot of predicted cross-reactive peptides compared with PHOX2B shows cross-reactive binders ABCA8 and MYO7B. Right, flow mean fluorescence intensity (MFI) used to calculate degree of binding relative to PHOX2B in table. e, Functional screening of ABCA8 and MYO7B shows CAR killing through ABCA8 only at a supraphysiological concentration of 50 µM versus through PHOX2B at 0.1 µM. ABCA8 and MYO7B were not detected in the normal tissue immunopeptidome, and none of the peptides predicted by sCRAP that were detected in the normal immunopeptidome (FDFTI, SLC23A2 and TNS4) show binding to 10LH. The experiment was performed once on the entire panel of CAR constructs and repeated for 10LH and 302LH on an expanded panel of peptides. f, Representative BLItz plot at 200 nM PHOX2B pMHC shows fast on rate for 10LH and 302LH and exceptionally slow off-rate for 10LH (k d = 7.6 × 10−4 s−1). g, Alanine scan of QYNPIRTTF reveals that mutations in five residues (N3A, I5A, R6A, T7A and T8A) result in significant abrogation of binding to PC-CAR 10LH (n = 2; data are mean ± s.d.). h, PHOX2B–HLA-A*24:02 crystal structure paired with alanine scan of 10LH enables mapping of peptide–receptor interface, revealing spatial conformation of five receptor contact residues (left, top view; right, side view of pMHC complex). Created with BioRender.com.
We then screened our panel of PHOX2B-directed CARs against the top seven pMHC predicted by sCRAP (Fig. 2b), allowing us to eliminate cross-reactive CARs and prioritize those with the highest degree of target selectivity (Fig. 2c). Of 25 CARs screened, we selected clone 10LH—which had the highest-specificity profile, showing only two peptides with more than 10% relative binding to 10LH as compared to PHOX2B—for further development (Fig. 2d).
To test the functional significance of the binding to potential off-target pMHCs predicted by sCRAP, we pulsed HLA-matched PHOX2B− SW620 colon adenocarcinoma cells with the PHOX2B peptide and potential cross-reactive peptides across a range of concentrations. Pulsing with the PHOX2B peptide resulted in complete cytotoxicity when co-cultured with 10LH at the lowest tested concentration of 0.1 µM. The 10LH CAR T cells did not induce cytotoxicity with the most cross-reactive predicted peptide ABCA8 at 10 µM, and only induced killing at the supraphysiological concentration of 50 µM. The second most cross-reactive peptide with 10LH (MYO7B) showed no CAR cytotoxicity at concentrations of up to 50 µM (Fig. 2e). Neither ABCA8 nor MYO7B has been detected in normal tissue immunopeptidome10, and none of the peptides previously detected in the normal tissue immunopeptidome display any cross-reactivity with PC-CAR 10LH (Fig. 2d). These screens demonstrate the utility of sCRAP for pre-emptively identifying off-target effects, efficiently screening their functional consequences and identifying binders with highly selective binding to tumour targets.
The lead scFv 10LH bound PHOX2B pMHC with a dissociation constant K D of 13 nM and exceptionally slow off rate (k d) of 7.6 × 10−4 s−1 (Fig. 2f). We next performed an alanine scan for the 10LH CAR, characterizing binding to PHOX2B pMHC with amino acid substitutions at each non-anchor position of the peptide34. The alanine scan revealed significant interactions of the PC-CAR receptor with five out of seven non-anchor residues of the PHOX2B peptide, including key residues protruding from the MHC cleft (interaction interface of 10LH with PHOX2B pMHC mapped on crystal structures in Fig. 2h), highlighting the superior selectivity of PC-CARs compared with the three or four residues that typically interact with the TCR35.
PC-CAR T cells break HLA restriction
Given the prerequisite of antigen processing and presentation necessary for detection of a given MHC peptide by immunopeptidomics, we hypothesized that identical peptides could be presented on additional HLA allotypes capable of binding a peptide’s anchor resides, and that some of these peptides could be presented in a similar enough conformation to be recognized by peptide-centric scFv binders (Extended Data Fig. 14a). We tested this hypothesis using PHOX2B-specific CARs engineered to bind the PHOX2B 9mer presented on HLA-A*24:02 in a peptide-centric fashion. We used our population-scale antigen presentation tool ShinyNAP8 to identify additional HLA allotypes that could present the same PHOX2B peptide, identifying 8 additional HLAs predicted to bind the PHOX2B 9mer (Extended Data Fig. 14b). We then used our pMHC structural modelling software, RosettaMHC36, to model the 3D conformation and binding free energy of peptides presented by additional HLA alleles, identifying HLA-A*23:01 and HLA-B*14:02 as top-scoring candidates for recognition by PC-CARs of the PHOX2B peptide originally discovered on HLA-A*24:02 (Fig. 3a, b). After validating binding of QYNPIRTTF to these alternate allotypes (Extended Data Fig. 14c), we measured the ability of 10LH to recognize these pMHCs, finding that in addition to HLA-A*24:02, 10LH binds with high affinity to the PHOX2B 9mer QYNPIRTTF presented by HLA-A*23:01 and HLA-B*14:02 (Fig. 3c). We also found that although QYNPIRTTF binds to HLA-C*07:02, 10LH exhibited 17.4-fold lower binding to HLA-C*07:02, probably owing to a distinct pMHC molecular surface in which the sidechains of the MHC α2-helix (R151 and Q155) protrude by 15Å at the position axially aligned with the key 10LH binding residues of QYNPIRTTF (I5 and R6) (Fig. 3b). To demonstrate functionally relevant recognition of our prediction of PHOX2B presentation on HLA-A*23:01, we pulsed the HLA-A*23:01 PHOX2B− melanoma cell line WM873 with the QYNPIRTTF peptide, showing induction of antigen-specific killing in peptide-pulsed cells and no cytotoxicity in cells pulsed with mismatched peptide (Extended Data Fig. 14d). HLA-A*23:01 is the most common non-A2 allele in people with African ancestry, highlighting the potential of PC-CARs to expand clinical application to underserved populations. Finally, we reanalysed our immunopeptidomics data, performing a matched peptide analysis to identify lower-confidence potential peptide matches to QYNPIRTTF in additional samples in which the peak was not fragmented. We identified m/z matches within 1 min of retention time across 6 out of 8 PDX lines and 7 out of 8 patient samples, each expressing one of the HLA alleles predicted by our analyses, suggesting that this peptide is ubiquitously expressed in neuroblastoma (Extended Data Fig. 14f). These findings warrant additional investigation into cross-HLA-targeting tumour self-antigens as well as neoantigens and demonstrate the potential to significantly expand the eligible patient population receiving peptide-centric scFv-based immunotherapies.
a, PHOX2B–HLA-A*24:02 crystal structure and models of PHOX2B in complex with HLA-A*23:01, HLA-B*14:02 and HLA-C*07:02. b, The charged and polar R151, Q155 and R69 residues of HLA-C*07:02 align with key 10LH interaction residues I5, R6 and I7 (MHC residues in blue and PHOX2B–10LH interaction residues in red). R151, Q155 and R69 create steric and charged hindrance of key peptide-binding residues. c, Staining of the PHOX2B PC-CAR 10LH reveals strong binding to HLA-A*24:02, HLA-A*23:01 and HLA-B*14:02, but not to HLA-C*07:02. CD19, CD19-directed CAR; 10LH, PHOX2B PC-CAR; UT, untransduced T cells. Created with BioRender.com.
PC-CAR T cells selectively eliminate tumours
We next tested the on-target killing potential of 10LH using available HLA-A*24:02 and HLA-A*23:01 neuroblastoma cell lines (SKNAS, NBSD and SKNFI) and demonstrated complete tumour cell killing and potent IFN-γ release after 24 h at 5:1 effector:target ratio (Fig. 4a–c, Supplementary Video 1). We tested the functional cross-reactivity of PC-CARs against the peptides presented by off-target tissues; PC-CARs showed no activity in three HLA-A*24:02 cell lines that do not express PHOX2B (SW620 colorectal adenocarcinoma, KATO III gastric adenocarcinoma and HEPG2 hepatocellular carcinoma) (Fig. 4a–c, Extended Data Fig. 10b; Supplementary Video 2). To validate the specificity of killing mediated by PC-CARs, we pulsed HLA-matched, PHOX2B-negative cancer cell lines with the PHOX2B peptide as well as forcibly over-expressing PHOX2B. We demonstrated specific killing only in cells pulsed with PHOX2B peptide and those transduced with full length PHOX2B mRNA, and not in cells pulsed with non-specific CHRNA3, ABCA8 and MYO7B peptides presented on the same HLA, nor in cells transduced with full length PRAME mRNA (Fig. 4d, e), demonstrating that native PHOX2B is processed and presented on MHC, where it is specifically recognized by PC-CARs. To detect PHOX2B pMHC on the cell surface, we generated a tetramerized 10LH scFv and stained on-target and off-target cell lines, which showed significant surface PHOX2B pMHC in neuroblastoma cells and not in HLA-matched controls (Fig. 4f, Extended Data Fig. 15a), suggesting that these reagents have the potential to be used to assess the presence of antigen in biopsied tissue samples. We also found that CARs flagged as cross-reactive by sCRAP demonstrated significant cross-reactivity, validating the functional consequences of cross-reactivities predicted by our algorithm (Extended Data Fig. 15b).
a–c, The CAR 10LH induces specific killing and IFN-γ release in neuroblastoma cells expressing HLA-A*24:02 and HLA-A*23:01 and PHOX2B (SKNAS, NBSD and SKNFI), but not in HLA-A*24:02 PHOX2B− non-neuroblastoma tumour cells (SW620, HEPG2 and KATO III), unless PHOX2B peptide is added. No T cell activity was observed in SW620 when pulsed with 10 μM of the predicted cross-reactive peptides ABCA8 or MYO7B (b, c). Cytotoxicity was visualized by T cell clustering and cleaved caspase (a), relative loss of confluence measured by loss of green fluorescence in GFP-transduced cancer cells in (b), and IFN-γ release measured by ELISA (c). UT denotes untransduced T cells. Assays performed using T cells from n = 3 donors, each in triplicate; data are mean ± s.d. d, Pulsing HLA-A*24:02 PHOX2B− cell line SW620 with 5 μM PHOX2B induces complete cell killing when co-cultured with 10LH CAR, but no killing when pulsed with 50 μM CHRNA3. Repeated across 3 experiments. e, 10LH CAR specifically and specifically kills SW620 control cells transduced with PHOX2B, but not with PRAME. f, Staining cancer cells with tetramerized 10LH scFv enables detection of PHOX2B pMHC on neuroblastoma cells but not in HLA-matched controls. g, PHOX2B-specific PC-CAR T cells induce potent tumour killing in mice engrafted with neuroblastoma PDX tumours, including the extremely fast-growing line COG-564x and HLA-A*23:01 line NBSD. n = 6 mice enrolled per arm (individual plots shown in Extended Data Fig. 17); data are representative from one of two independent in vivo studies for each PDX line; data are mean ± s.d. h, Treatment with 10LH and 302LH PC-CARs potently upregulate HLA expression in PDX tumours collected from lone mice in each arm reaching tumour burden compared with mice treated with untransduced T cells (COG-564x collected 11 days after treatment; NBSD collected 14 days after treatment for untransduced cells and 17 days after treatment for 10LH and 302LH; both tumours collected from one experiment). Created with BioRender.com.
We next treated immunodeficient mice engrafted with HLA-A*24:02 (SKNAS and COG-564x) and HLA-A*23:01 (NBSD) xenografts with 106 10LH- and 302LH-transduced CAR T cells once tumours reached 100–250 mm3. Both 10LH PC-CAR-treated and 302LH PC-CAR-treated mice showed complete tumour responses in both HLA-A*24:02 xenografts (Fig. 4g), but only 10LH-treated mice exhibited the response in the HLA-A*23:01 NBSD xenografts. This correlated directly with the relative affinity of these two constructs against the PHOX2B peptide presented on HLA-A*23:01 (Extended Data Fig. 14c), suggesting that a threshold affinity or distinct mode of binding by different scFvs may contribute to the ability to recognize the peptide in slightly altered conformations when presented by different HLA allotypes. We also observed that CAR treatment induced substantial upregulation of MHC in tumours. The COG-564x PDX model was generated from a post mortem blood draw from a patient with high-risk MYCN-amplified neuroblastoma who had suffered multiple relapses, and shows an extremely rapid tumour growth rate in mice. In this experiment, one mouse treated with the 10LH construct had a tumour reach endpoint size of 2 cm3 just one week after PC-CAR T cell therapy and was available for analysis, whereas all other tumours in this arm nearly reached endpoint size and then all regressed (Fig. 4g, Extended Data Figs. 16, 17). The lone COG-564x and NBSD tumours that reached endpoint showed significant PC-CAR T cell infiltrate and marked upregulation of MHC expression compared with endpoint tumours treated with non-transduced CAR T cells (Fig. 4h, Extended Data Fig. 17b). This upregulation is likely to be due to the potent IFN-γ release as measured in vitro, suggesting that these therapies can activate T cells at low antigen density to initiate a feed-forward cascade that increases MHC and antigen presentation.
Discussion
Here we have presented a process for identifying tumour-specific antigens derived from non-mutated oncoproteins, engineering PC-CARs against these tumour self-peptides and screening for cross-reactivity against MHC and the normal immunopeptidome. These methods have resulted in PC-CARs that can induce potent tumour killing across multiple HLA alleles in neuroblastoma and provide a roadmap for addressing the major challenges of therapeutic targeting of intracellular oncoproteins. These approaches demonstrate the value of pairing genomic, transcriptomic, epigenomic and immunopeptidomics datasets of normal and tumour tissues for the discovery of immunotherapy targets, as well as the utility of the ReD system paired with sCRAP to select ultra-rare scFv clones with desired binding and specificity profiles. Targeting of non-immunogenic self-antigens through pMHC-directed PC-CARs can vastly expand the landscape of actionable immunotherapeutic targets and enable the development of personalized immunotherapies in neuroblastoma and other cancers. Owing to the limitations in ionization and canonical search spaces of our immunopeptidomics, the prioritized peptides here are likely to represent only a fraction of potential pMHC complexes available to immunotherapeutic targeting. Neuroblastomas in general, and especially those harbouring MYCN amplification37, exist in a highly immunosuppressive tumour microenvironment such that future iterations of PC-CARs may require additional engineering to enable T cells to navigate to the pMHC target. However, our demonstration of significant upregulation of MHC (and thus target) in our models may help alleviate this therapeutic obstacle.
We also highlight the utility of pairing ShinyNAP with RosettaMHC in identifying HLA allotypes capable of presenting identical peptides in a similar conformation. We suggest that these tools, in addition to matched peptide searches of immunopeptidomics data across multiple tumour samples, have the potential to appreciably expand the identification of tumour-specific peptides presented on multiple HLAs. The potential ability to target these antigens beyond canonical HLA restriction can substantially expand the population of patients eligible to receive each PC-CAR construct and reach underserved populations, but this will need to be verified with other PC-CARs that are in development. The use of the sCRAP algorithm can rapidly exclude constructs with safety liabilities and prioritize those constructs withoptimal safety profiles, in contrast to alanine scanning which determines construct-specific cross-reactivities post hoc34.
We demonstrate several distinct advantages of PC-CARs for targeting of MHC peptides compared with TCRs: (1) PC-CARs can be used to target essential unmutated oncoproteins by inducing immunogenicity using synthetic peptide-centric receptors; (2) in contrast to TCRs, peptide-centric receptors are not constrained by the germline-encoded CDR1 and CDR2 interactions with MHC and therefore can be engineered such that these binding loops are able to form additional contacts with target peptides independently of the HLA, allowing us to generate receptors with superior peptide selectivity tothat of TCRs; and (3) owing to the lack of CDR1 and CDR2 loop interactions with MHC, PC-CARs can target a peptide presented by multiple HLA alleles, opening up the potential to substantially increase the clinical reach of each construct. Furthermore, PC-CARs may have an advantage over CARs in targeting membrane proteins in their ability to initiate a feed-forward loop of MHC upregulation and increased antigen density. These findings build on recent studies that demonstrate the ability to target neoantigens from mutated TP53 and RAS on MHC using scFv-based approaches and further demonstrate the utility of these approaches in targeting intracellular proteins38,39. We expect that the methods presented here will facilitate the discovery of tumour-specific targets in other human cancers with high unmet need and envision a library of scFv-based synthetic immunotherapies that provides population-scale coverage of HLA genotypes for both neoantigens and self-antigens.
Methods
Neuroblastoma samples and cell lines
Five neuroblastoma cell line xenografts and three patients derived xenografts showing a range of HLA expression by RNA sequencing and immunohistochemistry were selected for the initial immunopeptidomics experiment (Extended Data Table 1). All had whole exome sequencing and single nucleotide polymorphism array data available in addition to RNA-seq data18. Eight high-risk tumours with a mean mass of 0.56 g ranging from 0.17–1.7 g were obtained from Children’s Oncology Group (COG; https://childrensoncologygroup.org/) with matched sequencing from TARGET (https://ocg.cancer.gov/programs/target). Informed consent from each research subject or legal guardian was obtained for each deidentified tumour and blood sample used in this study through the COG neuroblastoma biobanking study ANBL00B1.
Human-derived neuroblastoma cell lines, including SK-N-AS, SK-N-FI and NB-SD were obtained from the Maris Lab cell line bank. Neuroblastoma cell lines were cultured in RPMI supplemented with 10% fetal bovine serum (FBS), 100 U ml−1 penicillin, 100 µg ml−1 streptomycin, and 2 mM l-glutamine. Other human cancer cell lines, including Jurkat,SW620, HEPG2 and KATO III were obtained from American Type Culture Collection (ATCC). Jurkat cells were cultured in Iscove’s modified Dulbecco’s medium (IMDM) supplemented with 10% FBS, 100 U ml−1 penicillin, 100 µg ml−1 streptomycin and 2 mM l-glutamine. SW620 cells were cultured in RPMI supplemented with 10% FBS, 100 U ml−1 penicillin, 100 µg ml−1 streptomycin, and 2 mM l-glutamine. HEPG2 cells were cultured in Eagle’s minimum essential medium (EMEM) supplemented with 10% FBS, 100 U ml−1 penicillin, 100 µg ml−1 streptomycin and 2 mM l-glutamine. KATO III cells were cultured in IMDM supplemented with 20% FBS, 100 U ml−1 penicillin, 100 µg ml−1 streptomycin and 2 mM l-glutamine. Packaging cell lines including Platinum-A cells and HEK 293T cells were obtained from Cell BioLabs and ATCC, respectively. Both packaging cell lines were cultured in DMEM supplemented with 10% FBS, 100 U ml−1 penicillin, 100 µg ml−1 streptomycin and 2 mM l-glutamine. All cell lines were grown under humified conditions in 5% CO2 at 37 °C, and samples were regularly tested for mycoplasma contamination.
Primary human T cells
Primary human T cells were obtained from anonymous donors through the Human Immunology Core at the University of Pennsylvania (Philadelphia, PA) under a protocol approved by the Children’s Hospital of Philadelphia Institutional Review Board. Cells were cultured using AIM-V (Thermo Fisher Scientific) supplemented with 10% FBS, 100 U ml−1 penicillin, 100 µg ml−1 streptomycin and 2 mM l-glutamine under humified conditions in 5% CO2 at 37 °C. T cell donors provided informed consent through the University of Pennsylvania Immunology Core
Isolation of HLA ligands by immunoaffinity purification
HLA class I molecules were isolated using standard immunoaffinity purification as described previously,53,54. In brief, cell pellets were lysed in 10 mM CHAPS/PBS (AppliChem/Lonza) containing 1× protease inhibitor (Complete; Roche). Mouse MHC molecules were removed using a 1 h immunoaffinity purification with H-2K-specific monoclonal antibody 20-8-4S, covalently linked to CNBr-activated sepharose (GE Healthcare). Remaining HLA molecules were purified overnight using the pan-HLA class I-specific monoclonal antibody W6/32 or a mix of the pan-HLA class II-specific monoclonal antibody Tü39 and the HLA-DR-specific monoclonal antibody L243, covalently linked to CNBr-activated Sepharose. MHC–peptide complexes were eluted by repeated addition of 0.2% trifluoroacetic acid (Merck). Elution fractions E1–E4 were pooled, and free MHC ligands were isolated by ultrafiltration using centrifugal filter units (Amicon; Merck Millipore). MHC ligands were extracted and desalted from the filtrate using ZipTip C18 pipette tips (Merck Millipore). Extracted peptides were eluted in 35 µl of acetonitrile (Merck)/0.1% trifluoroacetic acid, centrifuged to complete dryness and resuspended in 25 µl of 1% acetonitrile/0.05% trifluoroacetic acid. Samples were stored at −20 °C until analysis by LC–MS/MS.
Analysis of HLA ligands by LC–MS/MS
Peptide samples were separated by reversed-phase liquid chromatography (nanoUHPLC, UltiMate 3000 RSLCnano, Dionex) and subsequently analysed in an on-line coupled Orbitrap Fusion Lumos (Thermo Fisher Scientific). Samples were analysed in three technical replicates. Sample volumes of 5 µl (sample shares of 20%) were injected onto a 75 µm × 2 cm trapping column (Acclaim PepMap RSLC, Dionex) at 4 µl min−1 for 5.75 min. Peptide separation was subsequently performed at 50 °C and a flow rate of 300 nl min−1 on a 50 µm × 25 cm separation column (Acclaim PepMap RSLC, Dionex) applying a gradient ranging from 2.4–32.0% of acetonitrile over the course of 90 min. Eluting peptides were ionized by nanospray ionization and analysed in the mass spectrometer implementing the TopSpeed method. Survey scans were generated in the Orbitrap at a resolution of 120,000. Precursor ions were isolated in the quadrupole, fragmented by collision induced dissociation (CID) in the dual-pressure linear ion trap for MHC class I-purified peptides. Finally, fragment ions were recorded in the Orbitrap. An AGC target of 1.5x105 and a max injection time of 50ms was used for MS1. An AGC target of 7x104 and a max injection time of 150ms was used for MS2. The collision energy for CID fragmentation was 35%. For fragmentation mass ranges were limited to 400-650 m/z with charge states 2+ and 3+ for MHC class I.
Synthetic peptides were analyzed using a thirty minute gradient due to the simplicity of the sample. Full scan was acquired in the Orbitrap with a scan range of 300-1200 at 120,000 resolution. The automatic gain control (AGC) target was 5.0 × 105 with a maximum injection time of 50ms. Precursor ions were isolated in the quadropole, fragmented (CID, HCD and ETD) and analysed in the Orbitrap. MS2 were also acquired in the Orbitrap with 30,000 resolution, collision energy of 35%, AGC of 5 × 104, and maximum injection time of 150 ms. Since the discovery analysis were completed using CID, synthetic peptides fragmented with CID were compared for validation.
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE40 partner repository with the dataset identifier PXD027182 and 10.6019/PXD027182
HLA typing
FASTQ files from TARGET DNA and RNA-seq and RNA-seq data from cell lines were processed using PHLAT algorithm, as we previously validated on 15 HLA alleles8.
Database search and spectral annotation
Data were processed against the human proteome as compiled in the Swiss-Prot database (https://uniprot.org; 27 May 2021; 20,395 reviewed protein sequences contained) using the SequestHT algorithm3 in Proteome Discoverer (v2.1, Thermo Fisher Scientific) software. Precursor mass tolerance was set to 5 ppm, fragment mass tolerance to 0.02 Da. Search was not restricted to an enzymatic specificity. Oxidized methionine was allowed as a dynamic modification. FDR was determined by the Percolator algorithm based on processing against a decoy database consisting of shuffled sequences. FDR was set to 1%. Peptide lengths were limited to 8–14 amino acids for MHC class I. HLA annotation was performed using NetMHC-4.0 for HLA class I. For peptide matching, data was reprocessed using Proteome Discoverer (v2.4, Thermo FisherScientific) using the same parameters but with the addition of the feature mapper node to allow peptide matching between samples. Synthetic peptides were searched using a similar approach but Percolator was replaced with the fixed value PSM validator due to the simplicity of the synthetic peptide sample. Gene Ontology analyses were performedusing PANTHER41 (http://geneontology.org/) and P-values were calculated using Fischer’s exact test.
HLA binding predictions
HLA binding predictions were performed using NetMHC-4.042, NetMHCpan 4.143, and HLAthena44 on HLA class I.
scFv biopanning and CAR design
scFv binders against MHC presented peptides were retrieved from a large (2 × 1010) naive phage display scFv library45. A competitive panning process was developed to identify specific binders targeting the pMHC based on previous protocols46. Biotinylated pMHC monomers (target antigens) and non-biotinyated tetramers (decoy competitors) were obtained from the NIH tetramer core facility. 1012 copies of phages were depleted against magnetic beads (Invitrogen Dynabeads MyOne Streptavidin T1) for 30 min before incubation for 1.5 h with 5 µg biotinylated pep–MHC conjugated beads in the presence of 20 µg irrelevant decoy competitors. After incubation, the beads were washed with PBS containing 0.05% Tween-20 (PBST buffer) 5 times followed by two PBS washes. The remaining bound phage were recovered by log-phase TG1 and rescued by M13KO7 helper phage. The amplified phage was collected the next day by PEG–NaCl precipitation and used for the next round of panning. The target antigen input was decreased from 5 µg for the first round panning to 2 µg and 0.5 µg for the second and third rounds, respectively, and the washing conditions were more stringent along with the panning rounds. After three rounds of panning, polyclonal phage ELISA was performed to evaluate the enrichment. The TG1 cells from the second and third rounds were randomly picked into 96-well plates for soluble expression-based monoclonal ELISA (semELISA), as described previously46,47. Clones producing signals when binding to target antigens and not producing signals when binding the decoy competitors were amplified and sequenced. For protein preparation, these clones were transformed into HB2151 cells for expression, and proteins were purified by one-step Ni-NTA resin affinity. Protein purity and homogeneity were analysed by SDS–PAGE. Protein concentration was measured spectrophotometrically (NanoVue, GE Healthcare). Second-generation CAR constructs were synthesized using scFv sequences with 4-1BB and CD3ζ co-stimulatory domains and cloned into pMP71 vector for screening.
ReD library panning
The Ruby scFv library (>1011 diversity) was constructed using fully-germline IGLV3-1 and IGLV6-57 scaffolds paired with the IGHV3-23 scaffold as described33, with fully synthetic amino acid diversity in both VL and VH CDR3 loops.
The Ruby library was panned for two rounds using PHOX2B (43-51) MHC complex bound to MyOne Streptavidin C1 Dynabeads (ThermoFisher, Cat: 65002). Panned library output were transferred into the ReD cell-display platform33 and cells were permeabilized using 0.5% n-octyl β-d-thioglucopyranoside (Anatrace, catalogue (cat.) no. 0314) and labelled using recombinant PHOX2B pMHC complex ligated to fluorophores excitable by 405 nm and 488 nm lasers. Cells that were positive for target binding were isolated using the FACSMelody sorter (Becton-Dickinson).
After two rounds of positive selection for binding to PHOX2B MHC complex, two further rounds of fluorescence-activated cell sorting (FACS) were conducted using counter-labelled A*24:02 MHC complexes with unrelated peptides. After four rounds of FACS, individual colonies were picked and grown in 96-well plates before scFv induction, cell permeabilization, and PHOX2B MHC labelling and detection by CytoFLEX (Beckman Coulter).
Clones that were identified as binding specifically to the PHOX2B (43–51)–MHC complex were sequenced and unique scFvs were expressed as fusions to the AviTag biotinylation motif in Escherichia coli. Biotinylated scFv protein was released by permeabilization with 0.5% n-octyl β-d-thioglucopyranoside and purified to ~90% purity on Nickel NTA agarose resin (ABT, cat. no. 6BCL-NTANi).
Binding kinetics
Affinity measurements were performed using a BLItz system (ForteBio) and analysed using BLItz Pro software. Streptavidin biosensors (ForteBio, cat. no. 18-5019) were loaded with AviTag-biotinylated scFv, blocked with biotin, washed in PBS and then associated with pMHC ligand in PBS.
Steady-state binding assay
An equilibrium binding assay to target pMHCs was also established using MyOne Streptavidin C1 Dynabeads. In brief, 50 µg of Streptavidin C1 Dynabeads were incubated with excess biotinylated scFv before being blocked with free biotin and washed in PBS. Fluorophore-labelled pMHC complex was added to a concentration of 3.5 nM and incubated for 1 h at 4 °C followed by 10 min at 25 °C. Binding of the free MHC complex to the beads was quantified by the CytoFLEX at 488 nm (excitation)/525 nm (emission). Binding was normalized to beads without scFv and with unrelated control MHC complex.
This bead-binding assay was used to quantitate the binding of scFv to MHC complexes with alanine-scan substitutions of the PHOX2B peptides as well as to a plate of 95 unrelated 9mer peptide A*24:02 MHC complexes and the degree of cross-reactivity of binding of MHC complexes with peptides identified as having high homology to the PHOX2B peptide by eXpitope 2.0.
Viral production and transduction of Jurkat and primary T cells
Retrovirus for transduction of Jurkats and primary CD4/8 T cells was produced using Platinum-A (Plat-A) cells, a retroviral packaging cell line. Cells were plated in 6-well plates at 7 × 105 cells per well and transfected with 2.5 µg of the appropriate TCR or CAR construct in the retroviral vector pMP71 using Lipofectamine 3000 (Life Technologies, Invitrogen). After 24 h, medium was replaced with IMDM-10% FBS or AIM-V-10% FBS for Jurkat cells or primary cells, respectively. Supernatants were collected and filtered with 0.2-µm filters after 24 h incubation.
A second-generation lentiviral system was used to produce replication-deficient lentivirus. The day preceding transfection, 15 million HEK 293T cells were plated in a 15-cm dish. On the day of transfection, 80 µl Lipofectamine 3000 (Life Technologies, Invitrogen) was added to 3.5 ml room-temperature Opti-MEM medium (Gibco). Concurrently, 80 µl P3000 reagent (Thermo Fisher Scientific), 12 µg psPAX2 (Gag/Pol), 6.5 µg pMD2.G (VSV-G envelope), and a matching molar quantity of transfer plasmid were added to 3.5 ml room-temperature Opti-MEM medium. Virus supernatant was collected after 24 and 48 h later and briefly centrifuged at 300g and passed through a 0.45-µm filter attached to a syringe.
Jurkat cells were plated in 6-well plates pre-treated with 1 ml per well Retronectin (20 mg ml−1, Takara Bio) at 1 × 106 cells per well and spinoculatedwith 2 ml retroviral supernatant at 800g for 30 min at room temperature. After 24 h, cells were collected, and grown in IMDM-10% FBS.
Primary T cells were thawed and activated in culture for 3 days in the presence of 100 U ml−1 IL-2 and anti-CD3/CD28 beads (Dynabeads, Human T-Activator CD3/CD28, Life Technologies) at a 3:1 bead:T cell ratio. On days 4 and 5, activated cells were plated in 6-well plates pre-treated with 1 ml per well Retronectin (20 mg ml−1, Takara Bio) at 1 × 106 cells per well and spinoculated with 2 ml retroviral supernatant at 2,400 rpm for 2 h at 32 °C. On day 6, cells were collected and washed, beads were magnetically removed, and cells were expanded in AIM-V-10% FBS supplemented with 25 U ml−1 IL-2.
Primary human T cells were thawed and activated in culture for 1 day in the presence of 5 ng ml−1 recombinant IL-7, 5 ng ml−1 recombinant IL-15, and anti-CD3/CD28 beads (Dynabeads, Human T-Activator CD3/CD28, Life Technologies) at a 3:1 bead:T cell ratio in G-Rex system vessels (Wilson Wolf). On day 2, thawed lentiviral vector was added to cultured T cells with 10 µg ml−1 Polybrene (Millipore Sigma), and 24 h later vessels were filled with complete AIM-V medium supplemented with indicated concentrations of IL-7 and IL-15. On day 10, cells were collected and washed. Activation beads were magnetically removed, and cell viability was determined before freezing.
Human neuroblastoma cell lines were plated in 6-cm dishes, and 2 ml of thawed lentiviral vector produced with transfer plasmid pLenti-CMV-eGFP-Puro (Addgene plasmid #17448) was added with 10 µg ml−1 Polybrene (Millipore Sigma). Cells were selected for eGFP expression using flow-assisted cell sorting (BD FACSJazz, BD Biosciences) followed by 1 µg ml−1 puromycin selection.
sCRAP prediction
Tumour antigens were compared against the entire normal human proteome on the matched HLA (85,915,364 total normal peptides among 84 HLAs). Each residue in the same position of the tumour and human peptides was assigned a score for perfect match, similar amino acid classification or different polarity, scoring 5, 2 or −2, respectively (Extended Data Fig. 12). Similarity scores were calculated based on amino acid classification and hydrophobicity was determined using residues one and three through eight and excluding MHC anchor residues. Next, the maximum normal tissue RPKM values were identified from 1,643 normal tissues in GTEx. Normal peptides were compared to a database of normal tissue immunopeptidomes48. The overall cross-reactivity score for each normal peptide was then calculated using the following equation:
where n is the peptide length, \(P\) is the score of each amino acid of the normal peptide as compared to the tumour antigen, b is the pMHC binding affinity of the normal peptide, and E max is the maximum normal tissue expression
The algorithm is available at https://marisshiny.research.chop.edu/sCRAP/.
Tetramer and dextramer staining and flow cytometric analysis
Surface expression and binding of TCR- and CAR-transduced Jurkat cells and primary T cells was measured by staining with PE- or APC-conjugated dextramers carrying NB antigen peptide–MHC (Immudex). Cells were collected from culture, washed with 2 ml PBS at 800g for 5 min, incubated with 1 µl dextramer for 10 min in the dark, washed again, and resuspended in 300 µl PBS for analysis. Typically 5 × 105 cells were used for staining, and analysed on a BD LSR II (BD Biosciences) or an Attune Acoustic Focusing Cytometer (Applied Biosystems, Life Technologies). Flow cytometry data was collected using CytExpert (Beckman Coulter) and FACSDiva (BD Biosciences). Gating strategy for all tetramer and dextramer staining shown in Extended Data Fig. 9f.
Cross-reactivity pMHC screen
Potential cross-reactive peptides (GenScript) were suspended at a 200 µM working concentration. For each test, 0.5 µl of peptide was added to 5 µl HLA-A*24:02 empty loadable tetramer (Tetramer Shop) before incubating on ice for 30 min, or using TAPBR peptide exchange as previously described49. Following preparation, pMHC tetramers were used to stain cells (described above). CAR construct cross-reactivity values were determined using Jurkat cells transduced with CAR clones followed by staining with HLA-A*24:02 tetramers loaded with cross-reactive peptides. Mean fluorescent intensity was compared across peptides to determine cross reactivity.
Antigen-specific CD8 T cell enrichment and expansion
Normal donor monocytes were plated on day 1 in 6-well plates at 5 × 106 cells per well in RPMI-10 FBS supplemented with 10 ng ml−1 IL-4 (Peprotech) and 800 IU ml−1 GM-CSF (Peprotech) and incubated at 37 °C overnight. On day 2, fresh medium supplemented with 10 ng ml−1 IL-4 and 1,600 IU ml−1 GM-CSF was added to the monocytes and incubated at 37 °C for another 48 h. On day 4, non-adherent cells were removed, and immature dendritic cells washed and pulsed with 5 µM peptide in AIM-V-10% FBS supplemented with 10 ng ml−1 IL-4, 800 IU ml−1 GM-CSF, 10 ng ml−1 lipopolysaccharide (Sigma-Aldrich), and 100 IU ml−1 IFN-γ (Peprotech) at 37 °C overnight. Day 1 was repeated on days 4 and 8 to generate dendritic cells for the second and third stimulations on days 8 and 12, respectively.
On day 5, normal donor-matched CD8+ T cells were enriched using protein kinase inhibitor dasatinib (Sigma-Aldrich), dextramers, and anti-PE or anti-APC beads (Miltenyi Biotec) as previously described50. Enriched T cells were co-incubated with the appropriate pulsed dendritic cells in AIM-V-10% FBS. The day 5 protocol was repeated on days 8 and 12 using dendritic cells generated on days 4 and 8 for the second and third stimulation, respectively. Expanded T cells were validated for antigen-specificity by staining with the appropriate dextramers and for activation marker 41BB/CD137 (BioLegend).
Antigen-specific T cell sorting, sequencing and cloning
Expanded T cells were stained with CD3, CD8, CD14, CD19, live/dead and matched and mismatched dextramer an single-antigen-specific T cells were sorted using a FACSAria Fusion (BD Biosciences).
Sorted cells were loaded onto 10× Genomics 5′ V(D)J chips and libraries prepared according to manufacturer protocols. TCRα/β amplicons were run on MiSeq using 5,000 reads per cell. Sequencing data were processed using Cell Ranger and analysed using Loupe VDJ Browser. TCR alpha and beta chains were codon optimized and synthesized into bicistronic expression cassettes using engineered cysteine residues in the TCR constant domains, using F2A ribosomal skip sites and furin cleavage sites51. TCR cassettes were cloned into pMP71 retroviral vector.
Incucyte cytotoxicity assay
A total of 0.5 × 105 tumour cell targets were co-incubated with varying ratios of transduced primary cells (5 × 105, 2.5 × 105, 1 × 105, 0.5 × 105 and 2.5 × 104 for 10:1, 5:1, 2:1, 1:1 and 1:2 effector:target (E:T) ratios, respectively) in 96-well plates at 37 °C in the presence of 0.05 µM caspase-3/7 red (Incucyte, Essence BioScience). Plates ran on the Incucyte Zoom or S3 for 24–72 h and measured for apoptosis activity via caspase cleavage and comparison of relative confluency. Following the assay, supernatants were collected for ELISA. Total GFP integrated intensity (Total green calibrated units × μm2 per image) was assessed as a quantitative measure of live, GFP+ tumour cells. Values were normalized to the t = 0 measurement.
Cytokine secretion assays
Cell supernatant collected from cell cytotoxicity assays was thawed and plated in triplicate for each condition. IFN-γ and IL-2 levels were determined using ELISA kits according to the manufacturer’s protocol (BioLegend).
Antigen processing and presentation
Neuroblastoma cell lines were titrated with H1N5 influenza virus and infectivity was measured by flow cytometry using virus nucleoprotein (NP) antibody. HLA-A2 neuroblastoma cell lines were cultured with either 5 μM CEF1 or 50 HAU of H1N5 virus then co-cultured with M1 antigen-specific T cell hybridoma kindly provided by David Canaday52. T cell activation was measured using IL-2 ELISA (Abcam).
Expression, refolding, and purification of recombinant peptide and HLA molecules
HLA-A*02:01 and HLA-A*24:02 constructs for bacterial expression were cloned into pET24a+ plasmid. DNA plasmids encoding HLA-A*02:01 (heavy chain), HLA-A*24:02 (heavy chain), and human β2M (light chain) were transformed into E. coli BL21-DE3 (Novagen), expressed as inclusion bodies and refolded using previously described methods53. E. coli cells were grown in autoinduction medium54 for (16–18 h). Afterward, the E. coli cells were collected by centrifugation and resuspended with 25 ml BugBuster (Milipore Sigma) per litre of culture. The cell lysate was sonicated and subsequently pelleted by centrifugation (5,180g for 20 min at 4 °C) to collect inclusion bodies. The inclusion bodies were washed with 25 ml of wash buffer (100 mM Tris pH 8.0, 2 mM EDTA, and 0.01% v/v deoxycholate), sonicated, and pelleted by centrifugation. A second wash was done using 25 ml of Tris-EDTA buffer (100 mM Tris pH 8.0 and 2 mM EDTA). The solution was once again resuspended by sonication then centrifuged. The inclusion bodies were then solubilized by resuspension with 6 ml resuspension buffer (100 mM Tris pH 8.0, 2 mM EDTA, 0.1 mM DTT, and 6 M guanidine-HCl). Solubilized inclusion bodies of the heavy and light chain were mixed in a 1:3 molar ratio and then added dropwise over 2 days to 1 l of refolding buffer (100 mM Tris pH 8.0, 2 mM EDTA, 0.4 M arginine-HCl, 4.9 mM reduced l-glutathione, and 0.57 mM oxidized l-glutathione oxidized) containing 10 mg of synthetic peptide at >98% purity confirmed by mass-spectrometry (Genscript). Refolding was allowed to proceed for 4 days at 4 °C without stirring. Following this incubation period, the refolding mixture was dialyzed into the size-exclusion buffer (25 mM Tris pH 8.0 and 150 mM NaCl). After dialysis, the sample was concentrated first using a Labscale tangential flow filtration system and then using an Amicon Ultra-15 Centrifugal 10 kDa MWCO Filter Unit (Millipore Sigma), to a final volume of 5 ml. Purification was performed using size-exclusion chromatography on a HiLoad 16/600 Superdex 75 column. After size exclusion, the sample was further purified by anion exchange chromatography using a MonoQ 5/50 GL column and a 0–100% gradient of buffer A (25 mM Tris pH 8.0 and 50 mM NaCl) and buffer B (25 mM Tris pH 8.0 and 1 M NaCl). The purified protein was exhaustively exchanged into 20 mM sodium phosphate pH 7.2 and 50 mM NaCl. The final sample was validated using SDS–PAGE to confirm the formation of a pMHC complex containing both the heavy and light chains.
Differential scanning fluorimetry
To measure the thermal stability of the pMHc-I molecules, 2.5 μM of protein was mixed with 10× Sypro Orange dye in matched buffer (20 mM sodium phosphate pH 7.2, 100 mM NaCl) in MicroAmp Fast 96-well plates (Applied Biosystems) at a final volume of 50μl. Differential scanning fluorimetry was performed using an Applied Biosystems ViiA quantitative PCR machine with excitation and emission wavelengths at 470 nm and 569 nm, respectively. Thermal stability was measured by increasing the temperature from 25 °C to 95 °C at a scan rate of 1 °C min−1. Melting temperatures (T m) were calculated in GraphPad Prism 7 by plotting the first derivative of each melt curve and taking the peak as the T m.
Protein crystallization
Purified HLA-A*02:01–LLLPLLPPL, HLA-A*02:01–LLPLLPPLSP, HLA-A*02:01–LLPLLPPLSPS, HLA-A*02:01–LLPRLPPL and HLA-A*24:02–QYNPIRTTF complexes were used for crystallization. Proteins were concentrated to 10–12 mg ml−1 in 50 mM NaCl, 25 mM Tris pH 8.0, and crystal trays were set up using a 1:1 protein-to-buffer ratio at room temperature. Optimal crystals for HLA-A*02:01–LLLPLLPPL, HLA-A*02:01–LLPLLPPLSP and HLA-A*02:01–LLPLLPPLSPS were obtained with 1 M sodium citrate dibasic, 0.1 M sodium cacodylate pH 6.5. For HLA-A*02:01–LLPRLPPL, diffracting crystals were obtained with 0.2 M magnesium chloride, 0.1 M HEPES pH 7.0, 20 % PEG 6000. HLA-A*24:02–QYNPIRTTF diffracting crystals were obtained with 0.1 M HEPES pH 7.0, 10 % PEG 6000. Diffraction-quality crystals were collected and incubated from the above conditions plus glycerol as a cryoprotectant and flash-frozen in liquid nitrogen before data collection. All crystals used in this study were grown using the hanging-drop vapour-diffusion method. Data were collected from single crystals under cryogenic condition at Advanced Light Source (beam lines 8.3.1 and 5.0.1). Diffraction images were indexed, integrated, and scaled using MOSFLM and Scala in CCP4 Package55. Structures were determined by Phaser56 using previously published structures of HLA-A*02:01 (PDB ID 5C07)57 and HLA-A*24:02 (PDB ID 3VXN)58. Model building and refinement were performed using COOT59 and Phenix60, respectively.
Homology modelling of p/HLA complexes using RosettaMHC
Three-dimensional structural models of HLA-A*23:01, HLA-B*14:02 and HLA-C*07:02 bound to the peptide QYNPIRTTF were generated using RosettaMHC, an in-house method for modeling the α1 and α2 peptide-binding domains of pMHC-I molecules61. In brief, the amino acid sequences of HLA-A*23:01, HLA-B*14:02 and HLA-C*07:02 were first obtained from the IPD-IMGT/HLA Database62. The sequence of HLA alleles of interest was aligned against the sequences of 318 HLA curated template structures available in RosettaMHC. For each allele, all candidate templates were selected according to a 70% sequence identity criterion between aligned residues within the peptide-binding groove (within 3.5 Å of any peptide heavy atom). Generation of 3D models was performed using a Monte Carlo sampling of sidechain rotamer conformations, followed by gradient-based optimization of all backbone and sidechain degrees of freedom. For each peptide–HLA complex, the top 5 models with the lowest Rosetta binding energy were selected as the final structural ensemble. The quality of the final models was assessed using the Molprobity webserver63. Analysis of polar contacts and surface area were performed using the PyMOL Molecular Graphics System, version 2.4.1.
Immunohistochemistry
CD3 (Dako A0452), PHOX2B (Abcam ab183741), and HLA-ABC (Abcam ab70328) antibodies were used to stain formalin fixed paraffin embedded tissue slides. Staining was performed on a Bond Max automated staining system (Leica Biosystems). The Bond Refine polymer staining kit (Leica Biosystems, DS9800) was used. The standard protocol was followed with the exception of the primary antibody incubation which was extended to 1 h at room temperature. CD3, PHOX2B and HLA-ABC antibodies were at 1:100, 1:500 and 1:1,200 dilutions, respectively. Antigen retrieval was performed with E1 (Leica Biosystems) retrieval solution for 20 min (E2 for PHOX2B). Slides were rinsed, dehydrated through a series of ascending concentrations of ethanol and xylene, then coverslipped. Stained slides were then digitally scanned at 20× magnification on an Aperio CS-O slide scanner (Leica Biosystems).
Murine PC-CAR T cell preclinical trials
NOD SCID Gamma (NSG) female (6–8 weeks of age) mice from Jackson Laboratories (stock no. 005557) were used to propagate subcutaneous xenografts. All mice were maintained under barrier conditions and experiments were conducted using protocols and conditions approved by the IACUC at the Children’s Hospital of Philadelphia. Treatment was initiated via lateral tail intravenous injection. Dose administered was 100 µl per animal of vehicle or CAR T cells as a single treatment. Treatment was administered at weeks 8–10 when tumour volumes reached 150 mm3 to 250 mm3. Six mice were enrolled per arm based on previous experience and randomized based on tumour size. Mouse technician was blinded to T cell engineering. Tumour volume and survival were monitored bi-weekly measurements until the tumours reached a size of 2.0 cm3 or mice showed signs of graft versus host disease. Animals were removed from study and studies terminated following onset of graft-versus-host disease (GVHD) when animals display hunched posture, rapid breathing, urine staining, weight loss and a body condition score of 2, as determined by visual inspection. Onset of GVHD defined as urine staining and weight loss of 20% or weight loss of 10–15% if accompanied by hunched posture, laboured breathing, or poor body condition.
Statistics and reproducibility
Box and whisker plot representations of data show the median as centre, and 25th percentile and 75th percentile as bounds of boxes for plots shown in Fig. 1e, Extended Data Figs. 1, 13b.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Neuroblastoma HLA class I immunopeptidomics data is available through PRIDE under accession PXD027182. Sample files are annotated in Supplementary Table 2. Protein structures are available in the Protein Data Bank under accession codes 7MJ6 (HLA-A*02:01–LLLPLLPPL), 7MJ7 (HLA-A*02:01–LLPLLPPLSP), 7MJ8 (HLA-A*02:01–LLPLLPPLSPS), 7MJ9 (HLA-A*02:01–LLPRLPPL) and 7MJA (HLA-A*24:02–QYNPIRTTF). All other data are available within the article and supplementary information files, or by request from corresponding author.
Code availability
The sCRAP algorithm is accessible to the scientific community through a web portal (https://marisshiny.research.chop.edu/sCRAP/).
Change history
08 November 2023
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1038/s41586-023-06731-z
References
Pearlman, A. H. et al. Targeting public neoantigens for cancer immunotherapy. Nat. Cancer 2, 487–497 (2021).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Leko, V. & Rosenberg, S. A. Identifying and targeting human tumor antigens for T cell-based immunotherapy of solid tumors. Cancer Cell 38, 454–472 (2020).
Durbin, A. D. et al. Selective gene dependencies in MYCN-amplified neuroblastoma include the core transcriptional regulatory circuitry. Nat. Genet. 50, 1240–1246 (2018).
Majzner, R. G. & Mackall, C. L. Clinical lessons learned from the first leg of the CAR T cell journey. Nat. Med. 25, 1341–1355 (2019).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576.e516 (2017).
Yossef, R. et al. Enhanced detection of neoantigen-reactive T cells targeting unique and shared oncogenes for personalized cancer immunotherapy. JCI Insight 3, e122467 (2018).
Yarmarkovich, M. et al. Immunogenicity and immune silence in human cancer. Front. Immunol. 11, 69 (2020).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Shao, W. et al. The SysteMHC Atlas project. Nucleic Acids Res. 46, D1237–D1247 (2018).
Maus, M. V. et al. An MHC-restricted antibody-based chimeric antigen receptor requires TCR-like affinity to maintain antigen specificity. Mol. Ther. Oncolytics 3, 1–9 (2016).
Linette, G. P. et al. Cardiovascular toxicity and titin cross-reactivity of affinity-enhanced T cells in myeloma and melanoma. Blood 122, 863–871 (2013).
Morgan, R. A. et al. Cancer regression and neurological toxicity following anti-MAGE-A3 TCR gene therapy. J. Immunother. 36, 133–151 (2013).
Matthay, K. K. et al. Neuroblastoma. Nat. Rev. Dis. Primers 2, 16078 (2016).
Pugh, T. J. et al. The genetic landscape of high-risk neuroblastoma. Nat. Genet. 45, 279–284 (2013).
Burr, M. L. et al. An evolutionarily conserved function of polycomb silences the MHC class I antigen presentation pathway and enables immune evasion in cancer. Cancer Cell 36, 385–401.e388 (2019).
Aguet, F. et al. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Rokita, J. L. et al. Genomic profiling of childhood tumor patient-derived xenograft models to enable rational clinical trial design. Cell Rep. 29, 1675–1689.e1679 (2019).
Kowalewski, D. J. & Stevanović, S. Biochemical large-scale identification of MHC class I ligands. Methods Mol. Biol. 960, 145–157 (2013).
Bassani-Sternberg, M. et al. Direct identification of clinically relevant neoepitopes presented on native human melanoma tissue by mass spectrometry. Nat. Commun. 7, 13404 (2016).
Ma, X. et al. Pan-cancer genome and transcriptome analyses of 1,699 paediatric leukaemias and solid tumours. Nature 555, 371–376 (2018).
Freudenmann, L. K., Marcu, A. & Stevanović, S. Mapping the tumour human leukocyte antigen (HLA) ligandome by mass spectrometry. Immunology 154, 331–345 (2018).
Raabe, E. H. et al. Prevalence and functional consequence of PHOX2B mutations in neuroblastoma. Oncogene 27, 469 (2007).
Lee, N. H. et al. Clinical significance of tyrosine hydroxylase mRNA transcripts in peripheral blood at diagnosis in patients with neuroblastoma. Cancer Res. Treat. 48, 1399–1407 (2016).
Marachelian, A. et al. Expression of five neuroblastoma genes in bone marrow or blood of patients with relapsed/refractory neuroblastoma provides a new biomarker for disease and prognosis. Clin. Cancer Res. 23, 5374–5383 (2017).
Toor, J. S. et al. A recurrent mutation in anaplastic lymphoma kinase with distinct neoepitope conformations. Front. Immunol. 9, 99 (2018).
Cardoso-Moreira, M. et al. Gene expression across mammalian organ development. Nature 571, 505–509 (2019).
Pattyn, A., Morin, X., Cremer, H., Goridis, C. & Brunet, J.-F. The homeobox gene Phox2b is essential for the development of autonomic neural crest derivatives. Nature 399, 366–370 (1999).
Wang, L. et al. ASCL1 is a MYCN- and LMO1-dependent member of the adrenergic neuroblastoma core regulatory circuitry. Nat. Commun. 10, 5622 (2019).
Hata, J. L. et al. Diagnostic utility of PHOX2B in primary and treated neuroblastoma and in neuroblastoma metastatic to the bone marrow. Arch. Pathol. Lab. Med. 139, 543–546 (2015).
Mosse, Y. P. et al. Germline PHOX2B mutation in hereditary neuroblastoma. Am. J. Hum. Genet. 75, 727–730 (2004).
Dharia, N. V. et al. A first-generation pediatric cancer dependency map. Nat. Genet. 53, 529–538 (2021).
Beasley, M. D., Niven, K. P., Winnall, W. R. & Kiefel, B. R. Bacterial cytoplasmic display platform retained display (ReD) identifies stable human germline antibody frameworks. Biotechnol. J. 10, 783–789 (2015).
Kunert, A., Obenaus, M., Lamers, C. H. J., Blankenstein, T. & Debets, R. T-cell receptors for clinical therapy: in vitro assessment of toxicity risk. Clin. Cancer Res. 23, 6012–6020 (2017).
Rudolph, M. G., Stanfield, R. L. & Wilson, I. A. How TCRs bind MHCs, peptides, and coreceptors. Annu. Rev. Immunol. 24, 419–466 (2006).
Nerli, S. & Sgourakis, N. G. Structure-based modeling of SARS-CoV-2 peptide/HLA-A02 antigens. Front. Med. Technol. https://doi.org/10.3389/fmedt.2020.553478 (2020).
Asgharzadeh, S. et al. Clinical significance of tumor-associated inflammatory cells in metastatic neuroblastoma. J. Clin. Oncol. 30, 3525–3532 (2012).
Douglass, J. et al. Bispecific antibodies targeting mutant RAS neoantigens. Sci. Immunol. 6, eabd5515 (2021).
Hsiue, E. H.-C. et al. Targeting a neoantigen derived from a common TP53 mutation. Science, eabc8697 (2021).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Mi, H., Muruganujan, A., Ebert, D., Huang, X. & Thomas, P. D. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 47, D419–D426 (2019).
Nielsen, M. et al. Reliable prediction of T-cell epitopes using neural networks with novel sequence representations. Protein Sci. 12, 1007–1017 (2003).
Reynisson, B., Alvarez, B., Paul, S., Peters, B. & Nielsen, M. NetMHCpan-4.1 and NetMHCIIpan-4.0: improved predictions of MHC antigen presentation by concurrent motif deconvolution and integration of MS MHC eluted ligand data. Nucleic Acids Res. 48, W449–W454 (2020).
Sarkizova, S. et al. A large peptidome dataset improves HLA class I epitope prediction across most of the human population. Nat. Biotechnol. 38, 199–209 (2020).
Zhu, Z. & Dimitrov, D. S. Construction of a large naive human phage-displayed Fab library through one-step cloning. Methods Mol. Biol. 525, 129–142 (2009).
Zhang, M.Y. & Dimitrov, D. S. Sequential antigen panning for selection of broadly cross-reactive HIV-1-neutralizing human monoclonal antibodies. Methods Mol. Biol. 562, 143–154 (2009).
Chen, W., Xiao, X., Wang, Y., Zhu, Z. & Dimitrov, D. S. Bifunctional fusion proteins of the human engineered antibody domain m36 with human soluble CD4 are potent inhibitors of diverse HIV-1 isolates. Antiviral Res. 88, 107–115 (2010).
Shao, W., Caron, E., Pedrioli, P. & Aebersold, R. in Bioinformatics for Cancer Immunotherapy: Methods and Protocols (ed. Boegel, S.) 173–181 (Springer, 2020).
Overall, S. A. et al. High throughput pMHC-I tetramer library production using chaperone-mediated peptide exchange. Nat. Commun. 11, 1909 (2020).
Dolton, G. et al. Optimized peptide–MHC multimer protocols for detection and isolation of autoimmune T-cells. Front. Immunol. 9, 1378–1378 (2018).
Yang, S. et al. Development of optimal bicistronic lentiviral vectors facilitates high-level TCR gene expression and robust tumor cell recognition. Gene Ther. 15, 1411–1423 (2008).
Canaday, D. H. Production of CD4+ and CD8+ T cell hybridomas. Methods Mol. Biol. 960, 297–307 (2013).
Garboczi, D. N., Hung, D. T. & Wiley, D. C. HLA-A2–peptide complexes: refolding and crystallization of molecules expressed in Escherichia coli and complexed with single antigenic peptides. Proc. Natl Acad. Sci. USA 89, 3429–3433 (1992).
Studier, F. W. Stable expression clones and auto-induction for protein production in E. coli. Methods Mol. Biol. 1091, 17–32 (2014).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Cole, D. K. et al. Hotspot autoimmune T cell receptor binding underlies pathogen and insulin peptide cross-reactivity. J. Clin. Invest. 126, 2191–2204 (2016).
Shimizu, A. et al. Structure of TCR and antigen complexes at an immunodominant CTL epitope in HIV-1 infection. Sci. Rep. 3, 3097 (2013).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Nerli, S. & Sgourakis, N. G. Structure-based modeling of SARS-CoV-2 peptide/HLA-A02 antigens. Front. Med. Technol. https://doi.org/10.3389/fmedt.2020.553478 (2020).
Maccari, G. et al. IPD-MHC 2.0: an improved inter-species database for the study of the major histocompatibility complex. Nucleic Acids Res. 45, D860–D864 (2017).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Andreatta, M., Lund, O. & Nielsen, M. Simultaneous alignment and clustering of peptide data using a Gibbs sampling approach. Bioinformatics 29, 8–14 (2012).
Alvarez, B. et al. NNAlign_MA; MHC peptidome deconvolution for accurate MHC binding motif characterization and improved T-cell epitope predictions. Mol. Cell Proteomics 18, 2459–2477 (2019).
Sarkizova, S. et al. A large peptidome dataset improves HLA class I epitope prediction across most of the human population. Nat. Biotechnol. 38, 199–209 (2020).
Acknowledgements
This work was supported by a St Baldrick’s Foundation-Stand Up to Cancer Dream Team Translational Research Grant (SU2C-AACR-DT-27-17). The St. Baldrick’s Foundation collaborates with Stand Up To Cancer. Research Grants are administered by the American Association for Cancer Research, the Scientific Partner of SU2C. Stand Up To Cancer is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research (J.M.M.). This work was also supported by NIH grants U54 CA232568 as part of the Beau Biden Cancer Moonshot Program (J.M.M.) and NIH R35 CA220500 (J.M.M.), R01 AI143997 (N.G.S.), R35 GM125034 (N.G.S), the Science Center Quod Erat Demonstrandum (QED) program at the Philadelphia Science Center (J.M.M. and M.Y.), the Children’s Hospital of Philadelphia Cell and Gene Therapy Collaborative (J.M.M. and M.Y.), and the Giulio D’Angio Endowed Chair (J.M.M.). Diffraction data for protein crystallography were collected at the Advanced Light Source, which is supported by the Director, Office of Science, Office of Basic Energy Sciences, and U.S. Department of Energy, under contract DE-AC02-05CH11231. We thank D. Canaday, A. Wolpaw, E. Hooijberg and H. Goldstein for providing reagents. BioRender.com was used in generating figures.
Author information
Authors and Affiliations
Contributions
Conceptualization: M.Y. Methodology: M.Y., Q.F.M., J.W., H.S., T.N., E.R., M.d.M., D.M., K.R.B., H.T., L.E., S.T., S.N., A.W., R.W., M.C., D.M., S.S. and D.G. Immunopeptidomics: M.Y. and M.d.M. scFv screening: R.P., N.L.C., W.L., D.S.D. and B.R.K. Algorithm development: M.Y. and A.F. X-ray crystallography: J.S.T., S.T. and M.Y. Structural modelling, H.T. and M.Y. Analysis: M.Y., Q.F.M., H.T., N.K. and J.M.M. Investigation: M.Y. and J.M.M. Funding acquisition: M.Y., N.G.S. and J.M.M. Writing: M.Y. and J.M.M. Supervision: M.Y. and J.M.M.
Corresponding author
Ethics declarations
Competing interests
J.M.M. and M.Y. have equity stake in Tantigen Bio Inc. and HuLA Therapeutics. Both companies have interest in commercializing technologies described herein. B.R.K. is an employee of and has equity interests in Myrio Therapeutics. R.P. and N.L.C. are employees of Myrio Therapeutics. J.M.M., M.Y. and B.R.K. are co-inventors on a patent filed regarding the PHOX2B PC-CAR.
Additional information
Peer review information Nature thanks Lélia Delamarre, Naomi Taylor and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article has been retracted. Please see the retraction notice for more detail: https://doi.org/10.1038/s41586-023-06731-z
Extended data figures and tables
Extended Data Fig. 1 Prioritized peptides from immunopeptidome are highly differentially expressed in neuroblastoma.
Differential expression between 153 neuroblastoma tumors from TARGET compared to 1643 normal tissues in GTEx, as described in Fig. 1e.
Extended Data Fig. 2 Validation of antigens discovered by immunopeptidomics using LC/MS/MS of synthetic peptide.
Peptide sequences imputed from immunopeptidomics spectra were synthesized and LC/MS/MS was under matched conditions. Synthetic peptides show complete concordance with tumor-eluted peptides across all b and y ions; masses highlighted in green represent detected peaks corresponding to matched b and y ions found in both tumor-eluted peptide spectra and synthetic peptide spectra.
Extended Data Fig. 3 Crystal structures solved for IGFBPL1 presented as three distinct peptides on HLA-A2.
Crystallographic analysis of IGFBPL1 peptide bound to HLA-A*02:01. a. X-ray structure of HLA-A*02:01 presenting the IGFBPL1 nonameric peptide (LLLPLLPPL), where yellow lines represent polar contacts between the HLA groove and peptide. b. Differential Scanning Fluorimetry (DSF) of HLA-A*02:01 refolded with IGFBPL1 peptides of different lengths. The legend indicates the sequence of the IGFBPL1 peptide and the corresponding melting temperature of the resulting peptide/MHC-I complexes. Mean of triplicate samples reported with error bars representing SD. c. Overlay of IGFBPL1 9mer, 11mer, and 12mer in MHC groove reveals that core peptides and anchor residues are maintained across peptides of varying length, and that additional amino acids in the 11mer and 12mer protrude at C terminus downstream of the L9 anchor position.
Extended Data Fig. 4 Tumor antigens derived from parent genes are under control of neuroblastoma core regulatory circuit.
a. Illustration of core-regulatory circuit in which transcription factors bind to one another’s promoters, acting as feed-forward loop b. ChIP Seq data at prioritized neuroblastoma antigens parent gene loci are bound by each of the 7 core-regulatory circuit (CRC) proteins MYCN, ASCL1, GATA3, HAND2, PHOX2B, ISL1, and TBX2. All CRC binding sites are associated with a H3K27Ac super-enhancer element.
Extended Data Fig. 5 Properties of neuroblastoma immunopeptidome.
a. Logo plots of peptides eluted from HLA-A2 and HLA-A24 show canonical binding motifs for both alleles64. b. Peptides detected by immunopeptidomics are enriched for highly expressed parent genes as compared to the entire cellular transcriptome. c. Gene ontology analysis of the parent genes of peptides of the neuroblastoma immunopeptidome compared to the cellular transcriptome. The most significantly enriched ontology groups are nucleic acid binding proteins (RNA binding protein ontology p = 1.22e−25 and nucleic acid binding ontology p = 9.47e−17; Fischer’s Exact test).
Extended Data Fig. 6 Prioritized antigen parent genes are expressed during tissue development and downregulated in normal tissue after birth.
a-f. Temporal transcriptomic analysis of prioritized neuroblastoma target antigens reveals high expression during development and downregulation prior to birth in the majority of target genes. Expression shown for each tissue from shown from 4 weeks post-conception to 58-63 years with birth marked by red arrows. PHOX2B (a), TH (b), IGFBPL1 (c), and CHRNA3 (d) exhibit developmentally restricted expression patterns. HMX1 (e) shows expression in testes post-birth. GFRA2 (f) expression in brain may make this target more amenable to targeting by antibody-based therapies that do not cross blood-brain barrier.
Extended Data Fig. 7 Neuroblastoma antigen processing, presentation, and immunogenicity.
a. Schematic of immunogenicity experiment. HLA-A2 neuroblastoma cells were either infected with H1N5 or pulsed with synthetic CEF1 peptide and co-cultured with M1 antigen-specific T cell hybridoma line. T cell activation was evaluated by IL-2 release. b. Viral titration of neuroblastoma cells using H1N5 influenza virus measured by FACS staining of viral nucleoprotein expression at 0-200 heamagglutination (HAU). c. Experimental schematic of assay: 1) HLA-A2 neuroblastoma cells are pulsed with 5 μM CEF1 peptide and co-cultured with M1 antigen-specific hybridoma and IL-2 release is measured by ELISA; 2) Neuroblastoma cells are infected with 50 HAU of H1N5, co-cultured with M1 antigen-specific hybridoma, measured by IL-2 secretion. Four of seven tested HLA-A*02:01 elicit T hybridoma response when pulsed with CEF1 peptide; three of seven lines induce a response when infected with H1N5 virus. n = 3 independent experiments, each performed in triplicate; plots show mean +/- SD. d-e. T hybridoma activation is not associated with HLA expression, but activation is lower in MYCN amplified tumors (MNA) as compared to non-amplified tumors (MNN). Created with BioRender.com.
Extended Data Fig. 8 Detection of tumor self-antigen specific CD8 T cells in normal donors.
a. Gating used to select CD8 T cells. Similar gating strategy was used to select live singlets for Jurkat and primary cells transduced with CAR constructs (without selection for CD3 and exclusion of CD14, CD19, and CD4). b. Four normal donors stained with IGFBPL1 dextramer on x-axis and GFRA2 on y-axis shows rare population of antigen-specific cells varying by donor and antigen (left panel showing unstained and right panel showing tetramer stained). c. Frequencies and MFIs of antigen specific cells across donors. d. Top constructs generated from normal donor-derived antigen-specific TCRs found by single-cell sequencing show range of antigen binding PHOX2B, TH, and IGFBPL1 as compared to DMF5 receptor targeting MART-1. Screening for antigen specific T cells in 3 donors reveals that PHOX2B TCRs have minimal target binding, suggesting that PHOX2B self-antigen is immunogenically silent and warrants targeting using synthetic scFv receptors.
Extended Data Fig. 9 Development of antigen-specific CARs for neuroblastoma antigens.
a. ELISA of PHOX2B scFv A7 using PHOX2B p/MHC and decoy peptide on HLA-A*24:02. b. Schematic of second-generation CAR constructs. c. A7 CAR transduced into primary CD8 cells binds PHOX2B dextramer but not HLA-A*02:01 dextramer. d. A7 CAR preferentially binds PHOX2B dextramer but cross-reacts with mismatched peptides PBK and CHRNA3 on HLA-A*24:02 at high affinity. e. A7 CAR potently kills HLA-A*24:02 neuroblastoma lines, but also kills HLA+/antigen− tumor cells. f. Tetramer and dextramer gating strategy for pMHC staining.
Extended Data Fig. 10 PHOX2B A7 CAR.
a. SKNAS tumor cells co-cultured with A7 CAR shows potent killing. SKNAS cells plated on day 0 (top), non-targeting CAR (left) and A7 CAR (right) added after 18 h; measuring tumor confluence (green) and cleaved caspase (red) on day 2 (bottom) shows tumor outgrowth with non-specific CAR and killing of all tumor cells with A7 CAR. Representative images shown from experiment repeated three times using n = 3 replicates in each experiment. b. PHOX2B is expressed in neuroblastoma cell lines (SKNAS, SKNFI, and NBSD), and not in HLA-matched controls (SW620, KATO III, and HEPG2). PHOX2B is expressed in SW620 cells transduced with full-length PHOX2B. For gel source data from single experiment, see Supplementary Fig. 1.
Extended Data Fig. 11 Saturation mutagenesis of A7 construct resulted in single-antigen-specific population.
a-b. Cross-reactive binding to mismatched HLA-A24 peptides. Flow cytometry of Jurkat cells transduced with A7 CAR stained with PHOX2B dextramer on x-axis and mismatched PBK peptide on HLA-A*24:02 on y-axis. c. Saturation mutagenesis was performed on CDR loops 1-3 of the heavy chain. Each pool of mutants was stained with target pMHC and counter-stained with mismatched pMHC. Contribution of each CDR binding loop (mutagens labeled CDR loop # - position #) to binding HLA-A*24:02 is shown (green: no contribution to binding; red: significant HLA binding at specific amino acid). Contribution of each position to HLA binding was calculated as follows: (MFItarget(mut)/MFImismatch(mut))/((MFItarget(WT)/MFImismatch(WT). d. Mutations of A7 CAR at CDR3 positions 2 and 3 result in 12.4x and 4.7x shifts towards single specificity, respectively. Double mutation of D3A and R2A resulted in modified A7 CAR with no MHC cross-reactivity and reduced binding to target. Created with BioRender.com.
Extended Data Fig. 12 sCRAP cross-reactivity algorithm.
a. Cross-reactivity algorithm was developed to identify peptides presented on normal tissue with similar biophysical properties to tumor antigens such as to pre-emptively predict cross-reactivities and screen for specificity. b. Illustration of peptide scoring system described in methods. c. Schematic of algorithm workflow describing how tumor peptides are scored against each peptide predicted to be presented from the normal proteome (totaling 92.4x106 potential MHC peptides). Binding affinity is predicted for each normal peptide and maximum gene expression of parent gene are factored into the overall score of each peptide. Peptides are referenced against a normal tissue immunopeptidomics database. Created with BioRender.com.
Extended Data Fig. 13 pMHC Cross-Reactivity Algorithm sCRAP Predicts MAGE-A3 toxicity through TITIN.
a. Table of top predicted cross-reactive peptides to MAGE-A3 peptide EVDPIGHLY reveals cross-reactivity with Titin peptide ESDPIVAQY ranks 4th out of 1,143,861 potential peptides presented on HLA-A*01:01. b. TITIN is highly expressed in GTEx RNA-sequencing of heart (n = 108) and muscle (n = 138) tissues.
Extended Data Fig 14 Cross-HLA recognition of PHOX2B peptide.
a. Schematic of hypothesis proposing that PHOX2B peptide QYNPIRTTF detected by immunopeptidomics can be presented by additional HLA alleles after undergoing a common antigen processing pathway. b. Population-scale presentation across the length of the PHOX2B protein (all potential 9mers on x-axis) generated by ShinyNAP8 predicts PHOX2B peptide QYNPIRTTF to be presented by an additional 8 HLA alleles in addition to HLA-A*24:02. Additionally, QYNPIRTTF was found to bind additional common HLA alleles HLA-C*07:01, HLA-C*06:02, HLA-A*29:02, and HLA-A*32:01 using NetMHCpan 4.165 and also predicted by HLAthena66. c. Size exclusion chromatography of PHOX2B peptide QYNPIRTTF refolded with HLA-A*23:01, HLA-B*14:0, and HLA-C*07:02 shows formation of stable pMHC complex with each allotype. d. PC-CAR 10LH binds PHOX2B on HLA-A*23:01 demonstrates higher binding than 302LH, in concordance with observed in vivo activity (Fig. 4g). e. 10LH CAR kills HLA-A*23:01/PHOX2B− WM873 cells when pulsed with PHOX2B peptide but not with CHRNA3 peptide n = 2 technical replicates; reported as mean +/- SD. f. Matched peptide search identifies unfragmented peaks in additional neuroblastoma tumors (NBSD shown). Unfragmented NBSD peaks are within 0.006 Da m/z of peaks in which PHOX2B peptide QYNPIRTTF was identified by MS/MS in other samples and eluted within one minute of fragmented peaks. While validated MS/MS peaks were found in 2/8 PDX tumors and 2/8 primary tumors, peaks with m/z and retention times matched to validated QYNPIRTTF peaks were identified in 6/8 PDX tumors and 7/8 primary tumors. Created with BioRender.com.
Extended Data Fig. 15 a. Tetramerized 10LH binds to PHOX2B pMHC.
Pulsing SW620 HLA-A*24:02 with 50μM PHOX2B peptide results in 10LH tetrabody binding. b. CARs predicted to be cross-reactive show killing in HLA-matched cells. CARs 320LH and 280LH predicted to cross-react with peptides presented on normal tissue demonstrate significant cross-reactivity in HLA-matched SW620 cells. Representative images shown from n = 3 technical replicates.
Extended Data Fig. 16 Immunohistochemistry of tumors collected from mice exceeding tumor burden.
COG-564x tumor-bearing mice treated with 10LH and 302LH PC-CARs show T cell infiltration as measured by CD3, co-localized with loss of PHOX2B target expression and tissue necrosis. No evidence of T cell infiltration observed in mice treated with untransduced T cells. Tumors were collected from one of each mouse in treatment arms reaching tumor burden 11 days after receiving T cells. All other mice in treatment arms went on to achieve complete responses. Images shown from tumors collected from lone mice reaching tumor burden each study arm in single experiment.
Extended Data Fig. 17 PC-CARs result in tumor ablation in mice engrafted with patient-derived xenografts.
a. Tumor regression in individual mice in vivo treated with PC-CARs (n = 6; source data provided in Supplementary Table 3). b. COG-564x tumors collected from 302LH PC-CAR-treated mice collected at when mice reached tumor burden on day 11 (left) and endpoint of study on day 38 (right). Though tumors are detectable at endpoint, H&E and PHOX2B staining of 302LH PC-CAR-treated tumors reveals entirely necrotic tissue by endpoint of the study. Single available tumor collected from mouse reaching tumor burden for D11 and single endpoint tumor in one experiment collected for immunohistochemistry shown.
Supplementary information
Supplementary Figure 1
The original source images for all data obtained by immunoblotting that show the uncropped form of the gels shown in Extended Data Fig. 10. Gels are labelled according to loading control (Ku80) or experimental samples (PHOX2B).
Supplementary Table 1
Immunopeptidomics data for neuroblastoma PDX/CDX and primary patient tumours. LC/MS/MS proteomics for HLA class I peptides eluted from eight PDX/CDX and eight primary patient tumours using 1% FDR. Peptides are annotated by source protein/gene, tumour line, best predicted HLA binder based on patient HLA class I allotype and predicted binding affinity/rank.
Supplementary Table 2
Annotation of raw sample files uploaded on PRIDE (https://www.ebi.ac.uk/pride/archive/projects/PXD027182).
Supplementary Table 3
Source data for mouse tumour measurements shown in Fig. 4g and Extended Data Fig. 17.
Supplementary Video 1
10LH CAR induces specific killing in SKNAS HLA-A*24:02 PHOX2B+ neuroblastoma cells at 5:1 E:T ratio. Cytotoxicity visualized by T cell clustering and cleaved caspase.
Supplementary Video 2
10LH CAR induces no killing in SW620 HLA-A*24:02 PHOX2B− cells expressing at 5:1 E:T ratio. Cytotoxicity visualized by T cell clustering and cleaved caspase.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yarmarkovich, M., Marshall, Q.F., Warrington, J.M. et al. RETRACTED ARTICLE: Cross-HLA targeting of intracellular oncoproteins with peptide-centric CARs. Nature 599, 477–484 (2021). https://doi.org/10.1038/s41586-021-04061-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04061-6
This article is cited by
-
Discovery of immunotherapy targets for pediatric solid and brain tumors by exon-level expression
Nature Communications (2024)
-
MHC-I-presented non-canonical antigens expand the cancer immunotherapy targets in acute myeloid leukemia
Scientific Data (2024)
-
Antigen presentation in cancer — mechanisms and clinical implications for immunotherapy
Nature Reviews Clinical Oncology (2023)
-
HLA3DB: comprehensive annotation of peptide/HLA complexes enables blind structure prediction of T cell epitopes
Nature Communications (2023)
-
CAR immune cells: design principles, resistance and the next generation
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.