Abstract
Haematopoiesis in the bone marrow (BM) maintains blood and immune cell production throughout postnatal life. Haematopoiesis first emerges in human BM at 11–12 weeks after conception1,2, yet almost nothing is known about how fetal BM (FBM) evolves to meet the highly specialized needs of the fetus and newborn. Here we detail the development of FBM, including stroma, using multi-omic assessment of mRNA and multiplexed protein epitope expression. We find that the full blood and immune cell repertoire is established in FBM in a short time window of 6–7 weeks early in the second trimester. FBM promotes rapid and extensive diversification of myeloid cells, with granulocytes, eosinophils and dendritic cell subsets emerging for the first time. The substantial expansion of B lymphocytes in FBM contrasts with fetal liver at the same gestational age. Haematopoietic progenitors from fetal liver, FBM and cord blood exhibit transcriptional and functional differences that contribute to tissue-specific identity and cellular diversification. Endothelial cell types form distinct vascular structures that we show are regionally compartmentalized within FBM. Finally, we reveal selective disruption of B lymphocyte, erythroid and myeloid development owing to a cell-intrinsic differentiation bias as well as extrinsic regulation through an altered microenvironment in Down syndrome (trisomy 21).
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
There are no restrictions on data availability for new data presented in this study. FASTQ and raw count matrices for droplet-based scRNA-seq data for FBM from fetuses with Down syndrome and fetuses without Down syndrome have been deposited at EMBL-EBI ArrayExpress and at the European Nucleotide Archive (ENA), with accession codes as follows: E-MTAB-9389 (FBM from fetuses with Down syndrome and FBM from fetuses without Down syndrome), E-MTAB-10042 (FBM from fetuses with Down syndrome) and ERP125305 (FBM from fetuses without Down syndrome). FASTQ and raw count matrices for all other new data in this study have been deposited at EMBL-EBI ArrayExpress and at the Gene Expression Omnibus (GEO) with accession codes E-MTAB-9801 (FBM Smart-seq2 scRNA-seq); E-MTAB-9389 (BCR- or TCR-enriched VDJ FBM scRNA-seq; FASTQs only); GSE166895 (CD34+ FBM, fetal liver and cord blood CITE-seq) and GSE166895 (FBM total CITE-seq). The following data are also available to download as Scanpy h5ad objects with transformed counts through our interactive webportal: https://fbm.cellatlas.io/: (i) FBM scRNA-seq (fetuses with Down syndrome); (ii) FBM scRNA-seq (fetuses without Down syndrome); (iii) CD34+ FBM, fetal liver and cord blood CITE-seq; and (iv) FBM total CITE-seq. All source data are available in the accompanying source data file, unless manuscript or figure legend refers to a Supplementary Table. External datasets incorporated into this study include: (i) human fetal liver and yolk sac scRNA-seq data4 (E-MTAB-7407); (ii) human blood monocyte–DC scRNA-seq data10 (GSE94820); (iii) mouse BM scRNA-seq data19 (GSE122467); (iv) fetal and paediatric thymus scRNA-seq data3 (E-MTAB-8581); and (v) adult BM and cord blood scRNA-seq data from the Human Cell Atlas Data Coordination Portal ‘Census of Immune Cells’ project (https://data.humancellatlas.org/explore/projects/cc95ff89-2e68-4a08-a234-480eca21ce79). At the time of submission, there are no known accessibility restrictions on these external datasets. Source data are provided with this paper.
Code availability
Single-cell sequencing data were processed and analysed using publicly available software packages. Python and R code and notebooks for reproducing single-cell analyses are available at https://github.com/haniffalab/FCA_bone_marrow.
References
O’Byrne, S. et al. Discovery of a CD10-negative B-progenitor in human fetal life identifies unique ontogeny-related developmental programs. Blood 134, 1059–1071 (2019).
Charbord, P., Tavian, M., Humeau, L. & Péault, B. Early ontogeny of the human marrow from long bones: an immunohistochemical study of hematopoiesis and its microenvironment. Blood 87, 4109–4119 (1996).
Park, J.-E. et al. A cell atlas of human thymic development defines T cell repertoire formation. Science 367, eaay3224 (2020).
Popescu, D.-M. et al. Decoding human fetal liver haematopoiesis. Nature 574, 365–371 (2019).
Wiemels, J. L. et al. Prenatal origin of acute lymphoblastic leukaemia in children. Lancet 354, 1499–1503 (1999).
Muntean, A. G., Ge, Y., Taub, J. W. & Crispino, J. D. Transcription factor GATA-1 and Down syndrome leukemogenesis. Leuk. Lymphoma 47, 986–997 (2006).
Roy, A. et al. Perturbation of fetal liver hematopoietic stem and progenitor cell development by trisomy 21. Proc. Natl Acad. Sci. USA 109, 17579–17584 (2012).
Olsson, A. et al. Single-cell analysis of mixed-lineage states leading to a binary cell fate choice. Exp. Hematol. 44, S24 (2016).
Dahl, R. et al. Regulation of macrophage and neutrophil cell fates by the PU.1:C/EBPα ratio and granulocyte colony-stimulating factor. Nat. Immunol. 4, 1029–1036 (2003).
Villani, A.-C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356, eaah4573 (2017).
Mullighan, C. G. et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 446, 758–764 (2007).
Summers, Y. J., Heyworth, C. M., de Wynter, E. A., Chang, J. & Testa, N. G. Cord blood G0 CD34+ cells have a thousand-fold higher capacity for generating progenitors in vitro than G1 CD34+ cells. Stem Cells 19, 505–513 (2001).
Pimanda, J. E. et al. Gata2, Fli1, and Scl form a recursively wired gene-regulatory circuit during early hematopoietic development. Proc. Natl Acad. Sci. USA 104, 17692–17697 (2007).
Iwasaki, H. et al. Distinctive and indispensable roles of PU.1 in maintenance of hematopoietic stem cells and their differentiation. Blood 106, 1590–1600 (2005).
Palii, C. G. et al. Single-cell proteomics reveal that quantitative changes in co-expressed lineage-specific transcription factors determine cell fate. Cell Stem Cell 24, 812–820 (2019).
Muskens, I. S. et al. The genome-wide impact of trisomy 21 on DNA methylation and its implications for hematopoiesis. Nat. Commun. 12, 821 (2021).
Yu, S. et al. GABP controls a critical transcription regulatory module that is essential for maintenance and differentiation of hematopoietic stem/progenitor cells. Blood 117, 2166–2178 (2011).
Sullivan, K. D. et al. Trisomy 21 causes changes in the circulating proteome indicative of chronic autoinflammation. Sci. Rep. 7, 14818 (2017).
Baccin, C. et al. Combined single-cell and spatial transcriptomics reveal the molecular, cellular and spatial bone marrow niche organization. Nat. Cell Biol. 22, 38–48 (2020).
Suchting, S. et al. The Notch ligand Delta-like 4 negatively regulates endothelial tip cell formation and vessel branching. Proc. Natl Acad. Sci. USA 104, 3225–3230 (2007).
Kusumbe, A. P., Ramasamy, S. K. & Adams, R. H. Coupling of angiogenesis and osteogenesis by a specific vessel subtype in bone. Nature 507, 323–328 (2014).
Itkin, T. et al. Distinct bone marrow blood vessels differentially regulate haematopoiesis. Nature 532, 323–328 (2016).
Long, M. W., Briddell, R., Walter, A. W., Bruno, E. & Hoffman, R. Human hematopoietic stem cell adherence to cytokines and matrix molecules. J. Clin. Invest. 90, 251–255 (1992).
Lane, W. J. et al. Stromal-derived factor 1–induced megakaryocyte migration and platelet production is dependent on matrix metalloproteinases. Blood 96, 4152–4159 (2000).
Schulz-Knappe, P. et al. HCC-1, a novel chemokine from human plasma. J. Exp. Med. 183, 295–299 (1996).
Butko, E., Pouget, C. & Traver, D. Complex regulation of HSC emergence by the Notch signaling pathway. Dev. Biol. 409, 129–138 (2016).
Mulè, M. P., Martins, A. J. & Tsang, J. S. Normalizing and denoising protein expression data from droplet-based single cell profiling. Preprint at https://doi.org/10.1101/2020.02.24.96360 (2021).
Acknowledgements
We acknowledge funding from the Wellcome Human Cell Atlas Strategic Science Support (WT211276/Z/18/Z), MRC Human Cell Atlas award and Wellcome Human Developmental Biology Initiative. M.H. is funded by Wellcome (WT107931/Z/15/Z), The Lister Institute for Preventive Medicine and NIHR and Newcastle Biomedical Research Centre; S.A.T. is funded by Wellcome (WT206194), the ERC Consolidator Grant ThDEFINE and EU FET-OPEN MRG-GRAMMAR awards; relevant research in the B.G. group was funded by Wellcome (206328/Z/17/Z) and the MRC (MR/M008975/1 and MR/S036113/1); I.R. is funded by Blood Cancer UK and by the NIHR Oxford Biomedical Centre Research Fund; A.R. is funded by a Wellcome Trust Clinical Research Career Development Fellowship (216632/Z/19/Z) and supported by the NIHR Oxford Biomedical Centre Research Fund; L.J. is funded by an NIHR Academic Clinical Lectureship; S.W. is funded by a Barbour Foundation PhD studentship; M.M. is funded by an Action Medical Research Clinical Fellowship (GN2779); and E.L. is funded by a Sir Henry Dale fellowship from Wellcome and the Royal Society (107630/Z/15/Z), BBSRC (BB/P002293/1) and core support grants to Wellcome and MRC to the Wellcome–MRC Cambridge Stem Cell Institute (203151/Z/16/Z). This research was funded in part by the Wellcome Trust (see above for grant numbers). We thank the Newcastle University Flow Cytometry Core Facility, Bioimaging Core Facility, NovoPath, Genomics Facility and NUIT for technical assistance, the School of Computing for access to the High-Performance Computing Cluster, CellGenIT and A. Farnworth for clinical liaison. We thank the CRUK Cambridge Institute Genomics core for processing all Cambridge libraries amd sequencing, and K. Nazor, B. Yeung and T. S. Huang (BioLegend) for discussions to optimize the TotalSeq panel and protocol. The human embryonic and fetal material was provided by the joint MRC–Wellcome (MR/R006237/1) HDBR (https://www.hdbr.org/). This publication is part of the Human Cell Atlas (https://www.humancellatlas.org/publications/).
Author information
Authors and Affiliations
Contributions
M.H., S.A.T., I.R. and B.G. conceived and directed the study. M.H., S.A.T., B.G., and N.K.W. designed scRNA-seq and CITEseq experiments and data analysis approaches. L.J. and E.L. designed HSC/MPP culture experiments. I.R. and A. Roy designed experiments on FBM from fetuses with Down syndrome. Samples were isolated by S.L., R.A.B., I.G., J.E., P.B., K.A., S.O. and N.E., libraries were prepared by E.P. and E.S. and sequencing was performed by J.C., R.Q., R.H. and the Wellcome Sanger Institute core facility. Flow cytometry and FACS experiments were performed by R.A.B., L.J. and D. Maunder, supported by D. McDonald and A.F. Cytospins were performed by L.J. and D. Dixon, and in vitro culture differentiation experiments were performed by L.J., C.M. and D. Maunder. Immunofluorescence microscopy was performed by C.J., T.N., R.C., C.D.C., C.S. and M.A., with analysis performed by M.M., B.O., C.S., E.P. and I.G. M.S.K., B.L., O.A., M.T., D. Dionne, T.L.T., M.S., O.R.-R. and A. Regev generated adult and cord blood scRNA-seq datasets. CITE-seq datasets were generated by E.S., N.M. and N.K.W. Computational analysis was performed by S.W., I.G., M.Q.L., G.R., E.D., I.K., M.M., J.B., M.S.J. and M.E., and web portals were constructed by I.G., D.H. and J.M., with disease information assembled by K.P. and T.C. M.H., L.J., S.W., I.G., G.R., B.O., H.K., K.B.M., T.C., N.M., N.K.W., K.B.M., D.H., D.-M.P., S.B., A. Roy, E.L., B.G., I.R., I.G. and S.A.T. interpreted the data. M.H., L.J., S.W., I.G., G.R., B.G., I.R. and S.A.T. wrote the manuscript, with input from M.L.R.H and J.E.L. All authors read and accepted the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.O. is now an employee of Becton, Dickinson and Company (BD); the contributions of this author to the work were made before the commencement of employment at BD. O.R.-R. is an employee of Genentech. O.R.-R. is a co-inventor on patent applications filed at the Broad Institute related to single-cell genomics. All other authors declare no competing interests.
Additional information
Peer review information Nature thanks Ana Cumano, James Palis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 A single-cell atlas of human FBM.
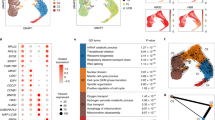
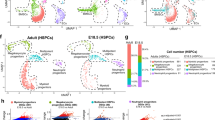
1a) Summary of FBM scRNA-seq dataset and reference scRNA-seq datasets used in this study, including published yolk sac and fetal liver data4 and publicly available cord blood/ABM data from the Human Cell Atlas Data Coordination Portal. 1b) UMAP of FBM scRNA-seq data (as per Fig. 1a) pre- and post- Harmony batch correction. Sequencing type and sample is represented by colour. 1c) Logistic regression for intersecting cell states annotated in FBM, ABM, cord blood, and FBM with Down syndrome scRNA-seq datasets. Prediction probability indicated by colour scale. 1d) Dotplots for expression of selected cell-state defining genes (left) in FBM CITE-seq data where corresponding protein was available in the ADT panel (right). Genes were selected from DE analysis (two-sided Wilcoxon rank-sum statistical test with Benjamini-Hochberg procedure for multiple testing correction; Supplementary Table 12). Markers used for FACS isolation are shown in bold type. Log-transformed, normalized and scaled gene expression values (upper limit 3) and DSB-normalized protein expression values (upper limit 15) are represented by the colour of the dots. Percentage of cells in each cell type expressing the marker is shown by the size of the dot. To contextualize dotplots shown in this panel, DE proteins were independently calculated for the FBM CITE-seq data (Supplementary Table 13), with method as described above for genes. 1e) FACS strategy used to isolate cell types for validation based on cell-state defining markers from scRNA-seq data. Representative plots from n = 2 samples (17 PCW) are shown. Gating strategy is described in Methods. 1f) UMAP of FBM Smart-seq2 scRNA-seq data (n = 2, k = 486, 17 PCW; Supplementary Table 15) with a 50-cell per subset sample of predicted 10x scRNA-seq counterparts (n = 9, k = 600). 10x scRNA-seq data represented by coloured areas and Smart-seq2 data represented by dots of equivalent colour.
Extended Data Fig. 2 A single-cell atlas of human FBM.
2a) Continuous decision tree constructed using the Rpart package to distinguish between the lineage-committed immune cell types in FBM (total) CITE-seq data using the 198 epitopes detected (see Supplementary Methods). Node splits were labelled ‘lo’ or ‘hi’ for visualization purposes. The branch rules of the tree operate on continuous protein expression values. 2b) Confusion matrix showing the actual cell type labels and the predicted cell type labels for the decision tree (in panel a) when run on the test data (see Methods). Sensitivity and specificity are illustrated in this heat map, with complete statistics provided in Supplementary Table 14. Overall accuracy computed with 95% CI using a binomial test and checked with a one-sided test (see caret package documentation for confusionMatrix function).
Extended Data Fig. 3 Diversification of innate myeloid and lymphoid cells.
3a) UMAP of monocyte, DC, neutrophil and macrophage subsets (k=34,318) in the FBM scRNA-seq dataset. 3b) FDG of DC, myeloid progenitor and HSC/MPP subsets (k=5,702) in the FBM scRNA-seq dataset. 3c) Left: Illustration displaying role of SPI1 and CEBPA dosage in influencing monocyte and neutrophil differentiation from GMPs. Right: log-transformed, normalized and scaled expression of CEBPA and SPI1 in GMPs from fetal liver and FBM scRNA-seq datasets. 3d) Heat map showing gene expression (GEX) for early monocyte and neutrophil commitment markers (two-sided Wilcoxon rank-sum statistical test with Benjamini-Hochberg procedure for multiple testing correction; Supplementary Table 30) in FBM scRNA-seq progenitors. Hierarchical clustering of each cell type shown (see Methods). GEX values are log-transformed, normalized and scaled (upper limit of 3). Sig = signature. 3e) Heat map showing expression of genes implicated in severe congenital neutropenia (Supplementary Table 31) across FBM scRNA-seq Monocle 3-inferred neutrophil pseudotime (DEGs across pseudotime marked with asterisk; one-sided Moran’s I statistical test; Supplementary Tables 32, 33). GEX values are log-transformed, normalized and scaled (upper limit of 1.5). 3f) Heat map showing gene enrichment (see Methods) of blood DC/monocyte signatures10 in developing and mature haematopoietic tissues (YS, fetal liver4, FBM, ABM). 3g) FDG of NK/ILCs (k=915) in the FBM scRNA-seq dataset. Grey ellipse highlights proliferating cells. 3h) Heat map showing NK cytotoxicity gene enrichment in NK cell states in YS, fetal liver and FBM (See Methods). Relative enrichment is indicated by colour scale. 3i) Heat map of predicted TF activity across inferred FBM DC pseudotime (FBM scRNA-seq DC-lineage cell states as input). TF activity inferred using iRegulon and pseudotime calculated using the Scanpy sc.tl.dpt function. GEX normalized to between 0-1 prior to plotting. 3j) Dotplot showing cell state-defining genes for NK and ILCs in the FBM scRNA-seq dataset. Methods/interpretation as in Fig. 1b. 3k) Dotplot showing expression of cell state-defining genes for DC and monocytes in the FBM scRNA-seq dataset. Methods/interpretation as in Fig. 1b. 3l) Dotplot showing expression of cell state-defining genes for myeloid precursors and neutrophils in the FBM scRNA-seq dataset. Methods/interpretation as in Fig. 1b.
Extended Data Fig. 4 Expanded B-lymphopoiesis in FBM.
4a) Dotplot of cell state-defining genes for FBM scRNA-seq B-lineage. Methods/interpretation as in Fig. 1b. Abbreviations as in Fig. 2b. 4b) Barplot for mean proportions of B-lineage cell states in fetal liver (n=14), FBM (n=9) and ABM (n=4) scRNA-seq datasets (n=biologically independent samples; B lineage absent in n=3 YS). P-values resulting from quasibinomial regression model (subject to one-sided ANOVA; with correction for sort gates; computed at 95% CI and adjusted for multiple testing using Bonferroni correction) are shown in parentheses; **p < 0.01, ****p < 0.0001; Supplementary Tables 19, 23). 4c) Violin-plot of DEGs (computed using two-sided Wilcoxon rank-sum statistical test with Benjamini-Hochberg procedure for multiple testing correction; ****=p < 0.0001; Supplementary Table 34) across FBM scRNA-seq Pre-B progenitor paths as in Fig. 2b; see Methods). GEX are log-transformed, normalized and scaled. 4d) Barplot showing mean proportion of productive heavy/light chains in FBM B-lineage cells present in both mRNA/BCR-enriched scRNA-seq (n=2, k=5,052). Pie-charts show the proportion of cycling cells (see Methods) per cell-type. 4e) Heat map of shared clonotypes between FBM B-lineage cell types, as defined by CellRanger. 4f) FDG of B-cell development (k=30,066) in the FBM scRNA-seq dataset. Colour indicates state (left) and Monocle 3 pseudotime value (right). Paths as in Fig. 2b. 4g) FDG of B-lineage (k=28,583) in the FBM scRNA-seq dataset. Colour indicates apoptotic gene enrichment score (see Methods/legend). 4h) Heat map of B-ALL-implicated genes across Monocle 3-inferred FBM/ABM B cell development pseudotime (see panel F; Supplementary Tables 32, 35–37; DEGs across pseudotime marked with asterisk; one-sided Moran’s I statistical test). Log-transformed, normalized and scaled GEX (upper limit of 2). 4i) Dotplot comparing expression of characteristic T-cell genes in cell states from thymus and FBM scRNA-seq datasets (see Methods). Interpretation as in Fig. 1b. Abbreviations: ETP = early thymocyte precursor; DN = double negative; DP = double positive. 4j) Barplot of productively rearranged TRA/B/G/D chains by T-cell state in n=4 biologically independent FBM samples at 14-17 PCW, k=194 cells. Chain rearrangements were as defined by CellRanger. Bars show mean and error bars SD. Mean±SD TCR productivity was 97±7%, 90±14% and 96±8% for CD4-T, CD8-T and Treg.
Extended Data Fig. 5 Tissue-specific properties of HSC/MPPs.
5a) Violin-plots showing GEX for MEM-, myeloid- and lymphoid- lineage genes in FBM scRNA-seq progenitors. GEX are log-transformed, normalized and scaled. Abbreviations: HSC/MPP = haematopoietic stem cell/ multipotent progenitor; CMP = common myeloid progenitor; eo/baso/mast pre. = eosinophil/basophil/mast cell precursor. 5b) FDG visualization of CD34+ FBM/fetal liver/cord blood CITE-seq cells on gene expression landscape (total k=35,273; FBM n=3, k=8,829, 14-17 PCW; FL n=4, k=18,904, 14-17 PCW; CB n=4, k=7,540, 40-42 PCW). Cell type is represented by colour, as shown in legend. HSC/MPP groups refer to unsupervised sub-clusters of the most immature compartment rather than functional MPP subpopulations. Abbreviations: MEP = megakaryocyte erythroid progenitor; MkP = megakaryocyte progenitor; EryP = erythroid progenitor; EoBasoMC = eosinophil/basophil/mast cell progenitor; MyP = myeloid progenitor; LyP = lymphoid progenitor. 5c) Logistic Regression for intersecting cell states in CD34+ CITE-seq data and FBM scRNA-seq data (see Methods). Prediction probability is indicated by colour scale. Cell type abbreviations as shown in panel a and b legend. 5d) Heat map showing cell-cycle gene enrichment in CD34+ FBM/fetal liver/cord blood CITE-seq progenitors. Colour indicates relative enrichment. 5e) Dotplot showing expression of genes used for progenitor characterization in the CD34+ CITE-seq data. Methods/interpretation as in Fig. 1b. 5f) Dotplot showing expression of proteins used for progenitor characterization in CD34+ CITE-seq data. Methods/interpretation as in Extended Data Fig. 1d (protein expression upper limit of 4). 5g) Bar-graph showing proportion of progenitor subsets out of total progenitors in fetal liver (n=4), FBM (n=3) and cord blood (n=4) CD34+ CITE-seq data (n= biologically independent samples). Proportions are normalized across donors. Bars indicate mean and error bars SD. Cell-type proportions across tissue were tested using a quasibinomial regression model (subject to one-sided ANOVA; with correction for sort gates; computed at 95% CI and adjusted for multiple testing using Bonferroni correction); *=p < 0.05 (Supplementary Table 38). 5h) Dotplot showing expression of protein markers significantly differentially expressed between HSC/MPP across tissues (fetal liver, FBM, cord blood) in the CD34+ CITE-seq dataset. Top differentially expressed proteins by log(fold change) are shown for each tissue (Supplementary Table 39; two-sided Wilcoxon rank-sum statistical test with Benjamini-Hochberg procedure for multiple testing correction). Methods/interpretation as in Extended Data Fig. 1d.
Extended Data Fig. 6 Tissue-specific properties of HSC/MPPs.
6a) Direction of Transition (DoT)-scores computed between CD34+ CITE-seq HSC/MPP1 across tissue (using cross-tissue HSC/MPP1 DEGs as input - see Supplementary Table 40). ABM scRNA-seq was used as reference and the origin point was defined as HSC/MPP (see Methods). Red-coloured cells indicate a shift towards their state (blue colour vice versa). 6b) Sort gates for HSC culture experiments. HSC/MPP, LMPP/MLP and CD34+CD38mid cells were index-sorted for single-cell culture on an MS5 stromal layer, as described in Methods. LMPP/MLP and CD34+CD38mid cells were analysed as ‘committed progenitors’. Abbreviations: LMPP/MLP = lymphoid-primed multipotent progenitor/ multipotent lymphoid progenitor. Figure was created using BioRender.com. 6c) Examples of single-cell HSC culture outputs showing outputs for: i) MK (CD41+), erythroid (CD235a+) and myeloid (CD14+ monocyte and CD15+ neutrophil), ii) NK (CD56+) and myeloid (CD14+ monocyte and CD15+ neutrophil). 6d) Outputs from single-cell culture on MS5 stromal layer for paired fetal liver and FBM HSC/MPPs. Proportion of culture wells producing colonies by cell-type/tissue (assessed by light microscopy under 4x magnification, analysed per plate- k=7 from n=3 biologically independent samples per tissue; lines display mean and error bars SEM). *p = 0.011 by 2-sided Mann Whitney test of BM HSC/MPP vs. fetal liver HSC/MPP and ***p = 0.0006 BM committed progenitor vs. fetal liver committed progenitor. 6e) Well contents analysed by flow cytometry and number of lineage outputs per well compared between HSC/MPP and committed progenitors from fetal liver vs. FBM. U=colony present but lineage undefinable by this assay. Statistical comparison (binomial test) for unipotential vs. multipotential colonies: HSC/MPP fetal liver vs. FBM *** p = 0.0008, 2-sided; committed progenitor fetal liver vs. BM ‘ns’ p = 0.27, 2-sided. 6f) Proportion of fetal liver and FBM HSC/MPPs producing myeloid-containing colonies in single-cell culture on MS5 stromal layer (paired FBM and fetal liver from n=2 biologically independent samples). Statistical comparison is of ‘myeloid-only’ vs. ‘myeloid plus other’ in k=77 wells producing myeloid colonies; *** p = 0.0001, 2-sided by binomial test. Abbreviations as follows: M = myeloid; E = erythroid; MK = megakaryocyte; NK = natural killer.
Extended Data Fig. 7 Perturbed haematopoiesis in Down syndrome.
7a) UMAP of Down syndrome FBM scRNA-seq (n=4, k=16,743, 12-13 PCW) (Supplementary Table 20). Abbreviations as in Fig. 1a. 7b) Top 30 PySCENIC-inferred differentially active TFs in Down syndrome vs. non-Down syndrome FBM scRNA-seq HSC/MPPs, MEMPs and MKs (Supplementary Tables 7, 20). TFs (red) described in text. 7c) Proportions of erythroid lineage cell states in Down syndrome (n=4) and age-matched non-Down syndrome FBM scRNA-seq data (n=2, where n=biologically independent samples). **** p < 10−15, 2-sided, by chi-squared test. Abbreviations: eryth = erythroid. 7d) Heat map showing cell-cycle gene enrichment in Down syndrome and age-matched non-Down syndrome FBM erythroid lineage cell states. Colour indicates relative enrichment. 7e) Representative images from single-cell HSC/MPP methylcellulose cultures, showing relative erythrocyte colony size/structure in Down syndrome (top; n=2 biological independent samples; PCW=17, 19; k=246) and non-Down syndrome (bottom; n=3 biologically independent samples; PCW=17, 19, 21; k=365) FBM; scale bar; 400µm. 7f) Heat map of erythropoiesis-implicated genes across Monocle 3-inferred erythroid development pseudotime in Down syndrome and non-Down syndrome FBM scRNA-seq datasets (all genes shown are DEGs across both pseudotime trajectories; one-sided Moran’s I statistical test; Supplementary Tables 32, 41, 42). Log-transformed, normalized and scaled GEX (upper limit of 3). 7g) Dotplot showing chromosome 21 TFs differentially expressed in Down syndrome and non-Down syndrome FBM scRNA-seq datasets (two-sided Wilcoxon rank-sum statistical test with Benjamini-Hochberg procedure for multiple testing correction; adjusted-p-value=<0.05; Supplementary Table 21). Dot-size = average log2 fold-change in Down syndrome expression. Colour = log(log(adjusted-p-value)). 7h) Dotplot showing TFs for differentially active regulons in Down syndrome vs. non-Down syndrome FBM scRNA-seq (see panel b). 7i) Top: Heat map showing TNF expression in Down syndrome and non-Down syndrome FBM scRNA-seq cell states. Bottom: TNFα-signalling pathway enrichment (see Methods, Supplementary Table 43). Dot-size = normalized enrichment score (NES) for TNFα-signalling pathway. Line = ±log10(0.25). 7j) Sankey-plot of putative TNF superfamily interactions in Down syndrome FBM scRNA-seq (see Methods; Supplementary Table 44). Fold-change expression in Down syndrome relative to non-Down syndrome (red scale). Combined expression in Down syndrome/non-Down syndrome (blue scale).
Extended Data Fig. 8 Stromal cell heterogeneity in FBM.
8a) UMAP of FBM scRNA-seq stromal cells (k=6,287). Dotted lines indicate broad lineages. Abbreviations: mac = macrophage; Fb = fibroblast; pre. = precursor. 8b) Dotplot showing cell state-defining genes for osteoclasts and macrophages in the FBM scRNA-seq dataset. Methods/interpretation as in Fig. 1b. 8c) Dotplot showing cell state-defining genes for osteochondral-lineage cells in the FBM scRNA-seq dataset. Methods/interpretation as in Fig. 1b. 8d) Barplot showing frequency of stromal cell states in FBM scRNA-seq samples. Samples are grouped into 4 developmental stages to facilitate statistical comparison over gestational stage. P-values resulting from quasibinomial regression model (subject to one-sided ANOVA; with correction for sort gates; computed at 95% CI and adjusted for multiple testing using Bonferroni correction) are shown in parentheses; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; Supplementary Tables 19, 45). 8e) Heat map showing gene enrichment (see Methods) of postnatal mouse BM stromal cell19 signatures in human FBM scRNA-seq stromal cells (coloured as in panel a). 8f) Dotplot showing cell state-defining genes for endothelial cells in the FBM scRNA-seq dataset. Methods/interpretation as in Fig. 1b. 8g) Left panel: Violin-plot showing expression of genes with documented role in sinusoidal endothelial cell function4 in sinusoidal endothelial cells from FBM and fetal liver scRNA-seq datasets. Right panel: Equivalent protein dotplot where marker was present in CD34+ CITE-seq ADT panel. Asterisks indicate DEGs (two-sided Wilcoxon rank-sum statistical test with Benjamini-Hochberg procedure for multiple testing correction; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; Supplementary Tables 46, 47). Methods/interpretation as in Fig. 3a, c (upper limit of 5.5 for protein dotplot). 8h) Heat map showing expression of mouse BM endothelial cell DEGs from Baryawno et al (see Supplementary Methods) in FBM scRNA-seq endothelial cells. GEX values are log-transformed, normalized and scaled (upper limit of 3). Hierarchical clustering for each cell type is shown (see Methods). 8i) Heat map of differentially enriched inflammatory and cytokine production pathways in Down syndrome vs. non-Down syndrome FBM scRNA-seq stroma defined by a two-sided Wilcoxon rank-sum test with Benjamini-Hochberg procedure for multiple testing correction; Supplementary Tables 20, 43).
Extended Data Fig. 9 FBM stromal cells in situ.
9a) Expression of IF microscopy markers in the FBM (total) CITE-seq dataset; genes (left) and their protein equivalent (right). Methods/interpretation as in Extended Data Fig. 1d (upper limits of 3/15 applied for gene/protein expression). 9b) Left: Scatterplot of CD34 and CD117 co-expression in FBM leukocytes by flow cytometry and expression of CLEC9A in gated fractions (representative from n=2). Right: Scatterplots of CD34, CD38 and CD117 protein expression in the FBM (total) CITE-seq dataset (Supplementary Table 48). Values are ln(DSB-normalized) and scaled to a lower limit of 0. Quadrants denote manual ‘gating’ thresholds (see Methods). Kernel density gradient is overlaid. 9c) Longitudinal section of fetal bone stained with hematoxylin and eosin. Left: location of haematopoietic tissue (10x magnification), scale bar; 1mm. Right top: haematopoietic architecture (20x magnification), scale bar; 200µm. Right bottom: identification of morphologically distinct cell types (50x magnification), scale bar; 20µm. Abbreviations: Eryth. = erythroid Neut. = neutrophil lineage; Eo. = eosinophil. Representative from n=4 samples (14-15 PCW). 9d) Longitudinal section of fetal femur with multiplex IF staining. Yellow boxes = regions of interest. Representative image from n=4 samples (14-15 PCW) at 4x magnification. Scale bar; 3mm. 9e) Sections of fetal femur, each stained with a single primary antibody from the multiplex and co-stained with DAPI. Representative fields of view from n=2 samples (14-15 PCW) at 20x magnification. Scale bars; 100µm. 9f) Identification of HSC/MPP and progenitors as cells co-expressing CD34 and CD117 (white arrow): left = all multiplex channels, middle = CD117 only, right = CD34 only. Representative image at 100x magnification. Replicates as per panel g. Scale bars; 50µm. 9g) Scatterplot showing proportions of CD34+ CD117+ HSC/MPP and progenitors per nucleated cells in metaphyseal (M) versus diaphyseal (D) regions of interest. Bars display mean and error bars SD of 522 HSC/MPP locations across 127 regions of interest in n=4 biologically independent FBM samples (14-15 PCW). Difference in frequency of CD34+CD117+ HSC/MPP and progenitors relative to cellular density was assessed by Wald test (p= 0.431) (Supplementary Table 28).
Extended Data Fig. 10 Predicted stromal cell interactions.
10a) Summary of receptor-ligand interactions predicted by CellPhoneDB (see Methods) between FBM stromal ligands and HSC/MPP receptors (Supplementary Table 49). Significant putative receptor-ligands across FBM neighbourhoods are indicated in Venn diagram overlapping regions. Figure was created using BioRender.com. 10b) GEX dotplot for FBM scRNA-seq stromal ligands and HSC/MPP receptors with role in CellPhoneDB-predicted receptor-ligand interactions shown in panel a. Methods/interpretation as shown in Fig. 1b (upper limit of 2 for both dotplots). Colours represent grouping of stromal cell types, as in panel a. 10c) Protein dotplot for CD34+ CITE-seq HSC/MPPs receptors with role in CellPhoneDB-predicted receptor-ligand interactions as per panels a-b. Methods/interpretation as in Extended Data Fig. 1f (upper limit of 1.5). 10d) Summary of receptor-ligand interactions predicted by CellPhoneDB between FBM HSC/MPP ligands and stromal receptors (Supplementary Table 49). Interpretation as detailed in panel a, and cell-type groupings detailed in Methods. 10e) GEX dotplot for FBM scRNA-seq stromal receptors and HSC/MPP ligands with role in CellPhoneDB-predicted receptor-ligand interactions shown in panel d. Methods/interpretation as shown in Fig. 1b (upper limit of 20% was placed on the HSC/MPP dotplot and upper limit of 2 was placed on both dotplots).
Supplementary information
Supplementary Information
This file contains Supplementary Methods (additional methodological information for both wet-lab experimental protocols and bioinformatic analysis) and the legends for Supplementary Tables 1–53 (see separate file for the tables).
Supplementary Tables
This file contains Supplementary Tables 1–53 (see Supplementary Information PDF for table legends).
Rights and permissions
About this article
Cite this article
Jardine, L., Webb, S., Goh, I. et al. Blood and immune development in human fetal bone marrow and Down syndrome. Nature 598, 327–331 (2021). https://doi.org/10.1038/s41586-021-03929-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03929-x
This article is cited by
-
Mapping human hematopoiesis
Nature Immunology (2024)
-
Targeting initial tumour–osteoclast spatiotemporal interaction to prevent bone metastasis
Nature Nanotechnology (2024)
-
InPACT: a computational method for accurate characterization of intronic polyadenylation from RNA sequencing data
Nature Communications (2024)
-
Generating human bone marrow organoids for disease modeling and drug discovery
Nature Protocols (2024)
-
Generation of complex bone marrow organoids from human induced pluripotent stem cells
Nature Methods (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.