Abstract
Centenarians have a decreased susceptibility to ageing-associated illnesses, chronic inflammation and infectious diseases1,2,3. Here we show that centenarians have a distinct gut microbiome that is enriched in microorganisms that are capable of generating unique secondary bile acids, including various isoforms of lithocholic acid (LCA): iso-, 3-oxo-, allo-, 3-oxoallo- and isoallolithocholic acid. Among these bile acids, the biosynthetic pathway for isoalloLCA had not been described previously. By screening 68 bacterial isolates from the faecal microbiota of a centenarian, we identified Odoribacteraceae strains as effective producers of isoalloLCA both in vitro and in vivo. Furthermore, we found that the enzymes 5α-reductase (5AR) and 3β-hydroxysteroid dehydrogenase (3β-HSDH) were responsible for the production of isoalloLCA. IsoalloLCA exerted potent antimicrobial effects against Gram-positive (but not Gram-negative) multidrug-resistant pathogens, including Clostridioides difficile and Enterococcus faecium. These findings suggest that the metabolism of specific bile acids may be involved in reducing the risk of infection with pathobionts, thereby potentially contributing to the maintenance of intestinal homeostasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Shotgun sequencing data are deposited in NCBI under BioProject PRJNA675598. Genome sequences of the 68 strains isolated from a centenarian and 16S rRNA amplicon sequence data are deposited in the DNA Data Bank of Japan under BioProject PRJDB11902 and PRJDB11894, respectively. LC–MS/MS data are deposited in Metabolomics Workbench (https://www.metabolomicsworkbench.org/) under project ID PR001168 with study ID ST001851 for human faeces data and study ID ST001852 for in vitro data. Source data are provided with this paper.
Code availability
Code for all of the analyses is available on GitHub (https://gitlab.com/xavier-lab-computation/public/centenarianmicrobiome).
References
Hirata, T. et al. Associations of cardiovascular biomarkers and plasma albumin with exceptional survival to the highest ages. Nat. Commun. 11, 3820 (2020).
Sebastiani, P. et al. Meta‐analysis of genetic variants associated with human exceptional longevity. Aging 5, 653–661 (2013).
Wu, L. et al. A cross-sectional study of compositional and functional profiles of gut microbiota in Sardinian centenarians. mSystems 4, e00325-19 (2019).
Franceschi, C., Garagnani, P., Parini, P., Giuliani, C. & Santoro, A. Inflammaging: a new immune-metabolic viewpoint for age-related diseases. Nat. Rev. Endocrinol. 14, 576–590 (2018).
Bárcena, C. et al. Healthspan and lifespan extension by fecal microbiota transplantation into progeroid mice. Nat. Med. 25, 1234–1242 (2019).
Biagi, E. et al. Gut microbiota and extreme longevity. Curr. Biol. 26, 1480–1485 (2016).
Rampelli, S. et al. Shotgun metagenomics of gut microbiota in humans with up to extreme longevity and the increasing role of xenobiotic degradation. mSystems 5, e00124-20 (2020).
Funabashi, M. et al. A metabolic pathway for bile acid dehydroxylation by the gut microbiome. Nature 582, 566–570 (2020).
Ridlon, J. M., Harris, S. C., Bhowmik, S., Kang, D. J. & Hylemon, P. B. Consequences of bile salt biotransformations by intestinal bacteria. Gut Microbes 7, 22–39 (2016).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
Devlin, A. S. & Fischbach, M. A. A biosynthetic pathway for a prominent class of microbiota-derived bile acids. Nat. Chem. Biol. 11, 685–690 (2015).
Nixon, M., Upreti, R. & Andrew, R. 5α-Reduced glucocorticoids: a story of natural selection. J. Endocrinol. 212, 111–127 (2012).
Hang, S. et al. Bile acid metabolites control TH17 and Treg cell differentiation. Nature 576, 143–148 (2019).
Campbell, C. et al. Bacterial metabolism of bile acids promotes generation of peripheral regulatory T cells. Nature 581, 475–479 (2020).
Song, X. et al. Microbial bile acid metabolites modulate gut RORγ+ regulatory T cell homeostasis. Nature 577, 410–415 (2020).
Paik, D. et al. Human gut bacteria produce TH17-modulating bile acid metabolites. Preprint at https://doi.org/10.1101/2021.01.08.425913 (2021).
Li, W. et al. A bacterial bile acid metabolite modulates Treg activity through the nuclear hormone receptor NR4A1. Preprint at https://doi.org/10.1101/2021.01.08.425963 (2021).
Buffie, C. G. et al. Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature 517, 205–208 (2015).
Begley, M., Gahan, C. G. M. & Hill, C. The interaction between bacteria and bile. FEMS Microbiol. Rev. 29, 625–651 (2005).
Thanissery, R., Winston, J. A. & Theriot, C. M. Inhibition of spore germination, growth, and toxin activity of clinically relevant C. difficile strains by gut microbiota derived secondary bile acids. Anaerobe 45, 86–100 (2017).
Honda, A. et al. Regulation of bile acid metabolism in mouse models with hydrophobic bile acid composition. J. Lipid Res. 61, 54–69 (2020).
Holmes, I., Harris, K. & Quince, C. Dirichlet multinomial mixtures: generative models for microbial metagenomics. PLoS ONE 7, e30126 (2012).
Atarashi, K. et al. Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science 358, 359–365 (2017).
Li, D., Liu, C. M., Luo, R., Sadakane, K. & Lam, T. W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Plaza Oñate, F. et al. MSPminer: abundance-based reconstitution of microbial pan-genomes from shotgun metagenomic data. Bioinformatics 35, 1544–1552 (2019).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841 (2014).
Segata, N., Börnigen, D., Morgan, X. C. & Huttenhower, C. PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat. Commun. 4, 2304 (2013).
Khelaifia, S., Raoult, D. & Drancourt, M. A versatile medium for cultivating methanogenic archaea. PLoS ONE 8, e61563 (2013).
Aziz, R. K. et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9, 75 (2008).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
García-Bayona, L. & Comstock, L. E. Streamlined genetic manipulation of diverse Bacteroides and Parabacteroides isolates from the human gut microbiota. mBio 10, e01762-19 (2019).
Ferrières, L. et al. Silent mischief: bacteriophage Mu insertions contaminate products of Escherichia coli random mutagenesis performed using suicidal transposon delivery plasmids mobilized by broad-host-range RP4 conjugative machinery. J. Bacteriol. 192, 6418–6427 (2010).
Atarashi, K. et al. Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500, 232–236 (2013).
Quinn, R. A. et al. Global chemical effects of the microbiome include new bile-acid conjugations. Nature 579, 123–129 (2020).
McDonald, J. A. K. et al. Evaluation of microbial community reproducibility, stability and composition in a human distal gut chemostat model. J. Microbiol. Methods 95, 167–174 (2013).
Claesson, M. J. et al. Gut microbiota composition correlates with diet and health in the elderly. Nature 488, 178–184 (2012).
Acknowledgements
We thank L. Besse for project management and making data available through the SRA and the Broad Institute Genomics Platform and Microbial ’Omics Core for sample processing and sequencing data generation; A. Minowa, M. Asakawa, K. Sugita and members of the JSR-Keio University Medical and Chemical Innovation Center for their technical support; N. Hasegawa, K. Fukunaga, T. Kanai, K. Masaki, S. Azekawa, K. Sasahara, S. Hosomi, M. Shimura and Y. Abe for their assistance in collecting clinical samples; S. Atsushi and members of the Honda laboratory for their suggestions during the course of this studies. K.H. was funded through AMED LEAP under grant number JP20gm0010003, Grant-in-Aid for Specially Promoted Research from JSPS (no. 20H05627), Public/Private R&D Investment Strategic Expansion Program (PRISM) from Cabinet Office of the Government of Japan, the Naito Foundation and the Takeda Science Foundation. The centenarian study was funded by the Japan Ministry of Agriculture, Forestry and Fisheries (M.S.) and Keio Global Research Institute (Y.A. and N.H.). The older-participant recruitment study was funded by a Grant-in-Aid for Scientific Research (no. 18H03055) from the Japan Society for the Promotion of Science and JST Research Complex Program (JP15667051). D.R.P. and R.J.X. were funded by the Center for the Study of Inflammatory Bowel Disease (DK043351), Center for Microbiome informatics and Therapeutics (CMIT) and AT009708. Y. Sato. was supported by the Terumo Life Science Foundation.
Author information
Authors and Affiliations
Contributions
K.H., Y. Sato and K.A. planned experiments, analysed data and wrote the paper together with D.R.P., R.J.X., A.N.S. and S.M.K.; Y.A., K.T. and N.H. collected clinical samples; Y. Sato and K.A. performed bacterial experiments; D.R.P., H.V. and R.J.X. performed metagenome analyses; K.A., S.M.K., Y.O., W.S. and M.H. performed whole-genome and meta-16S rRNA gene sequencing and analyses; S. Sasajima, Y. Sato, K.A., H.T., H.N., S.N., Y. Sugiura and M.S. performed metabolomic analysis; T.S., S.O., S. Sasajima and T.M. synthesized chemical compounds; N.M. and S. Shibata. performed electron microscopy imaging; A.H., Y.U., T.I., Y.L., T.T., J.I., H.I. and K.M. provided essential materials; D.R.L. and M.A.F. supervised bacterial experiments.
Corresponding authors
Ethics declarations
Competing interests
K.H. is a scientific advisory board member of Vedanta Biosciences and 4BIO CAPITAL.
Additional information
Peer review information Nature thanks Pieter Dorrestein, Andrew Patterson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Clinical characteristics of centenarians.
a, b, Activities of daily living (ADL) scores assessed via the Barthel index (a) and mini-mental state exam (MMSE) scores (b) of centenarian (top, ADL, n = 96; MMSE, n = 67) and older (bottom, ADL, n = 111; MMSE, n = 111) groups. c, Blood tests from centenarian (CE, n = 146-150, orange), older (n = 111, blue), and young (n = 15-39, grey) participants; red blood cell (RBC) count; albumin; C-reactive protein (CRP); white blood cell (WBC) count; total protein; blood urea nitrogen (BUN); fasting blood glucose; uric acid; and creatinine. d, Faecal lipocalin level of centenarian (n = 122), older (n = 67), and young (n = 19) individuals quantified by ELISA. e, Body-mass-index (BMI) values of centenarian (n = 89), older (n = 111), and young (n = 39) individuals. f, Percentages of individuals with medical histories of diabetes mellitus, hypertension, or cancer. Black colour represents affected individuals. Medical history was surveyed from 155-156 centenarian, 110-111 older, and 39 young individuals. In c-e, data are mean ± s.d. ***P < 0.001; **P < 0.01; *P < 0.05; Kruskal–Wallis with Dunn’s test. ns, not significant. Grey areas within graphs are ranges of normal values.
Extended Data Fig. 2 Gut microbiome signatures in centenarian, older and young Japanese participants based on whole-metagenome shotgun sequencing and de novo assembly analysis.
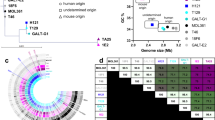
a, Principal coordinate analysis based on species-level Bray–Curtis dissimilarity from the assembled faecal metagenomes of centenarian [CE, n = 176, orange (154 individuals: 3 participants undergoing antibiotic treatment and 3 with insufficient bacterial DNA yield were excluded from the total 160 participants; analysis also included samples collected twice from 20 individuals and three times from 1 individual with an intervening one or two year interval)], older [n = 110, blue (110 individuals: 1 participant undergoing antibiotics treatment and 1 participant with insufficient bacterial DNA yield were excluded from 112 participants)], and young [n = 44, grey (44 individuals: 3 participants with insufficient bacterial DNA yield were excluded from 47 participants]. The microbiota composition of centenarians was significantly different from that of both control groups (PERMANOVA false-discovery rate (FDR)-adjusted P < 0.05). b, c, Relative abundance across phyla. d, Shannon diversity index (*P < 0.05, linear model). e-g, Changes in the relative abundance (RelAb) of gut metagenome species (MSPs) between centenarian, older, and young participants grouped according to the following signatures of differential abundance: 1) ageing signature, 2) rejuvenation signature, and 3) centenarian signature. Each signature is accompanied by models depicting microbial relative abundance patterns in centenarian (C), older (O), and young (Y) groups that would fall into the given signature. Colour scale represents the coefficient from the linear model and indicates enrichment (red) or depletion (blue) of a species in the respective comparisons: centenarian compared to older, centenarian compared to young, and older compared to young; in each case, the latter group is used as a reference in the model. Differentially abundant species that are significant at FDR P < 0.05 are indicated with asterisks. The first signature (‘ageing signature’) included taxa whose abundance was increased or decreased with age (e). For example, Eubacterium siraeum and undefined Firmicutes species (msp_161, 213) were most abundant in centenarians, followed by the older and then the young controls, whereas Blautia wexlerae displayed the opposite trend, being most abundant in young controls, followed by the older participants, and finally the centenarians. These findings are in alignment with previous studies that suggest the relative abundances of these taxa reflect adaptation to ageing, and may be related to physical activity and diet3,40. The second signature (‘rejuvenation signature’) included taxa whose abundance was similar in centenarians and young controls, but distinct from the older participants (f). These species might reflect the maintenance of youth or possess reverse-ageing effects. Notably, R. gnavus and E. lenta were part of this signature, as they were comparably abundant in both centenarians and young controls. Notably, these species have been implicated in bile acid metabolism, particularly the biosynthesis of iso-bile acids11. The third signature (‘centenarian signature’) included centenarian-specific taxa whose abundance was significantly different between centenarians and both the older and young control groups, but not between these two control groups (g). In the third signature, Alistipes, Parabacteroides, Bacteroides and Clostridium species, as well as Methanobrevibacter, a predominant archaeon in the human gut, were specifically enriched in centenarians compared with the other groups. One of the most abundant species in centenarians was C. scindens. By contrast, key butyrate producers such as F. prausnitzii and E. rectale were selectively depleted in centenarians. h, i, Abundance of genes homologous to the C. scindens bai operon in Japanese (h) and Sardinian (i) centenarian, older, and young age groups. Sardinian centenarians (n = 19), older controls (n = 23), and young controls (n = 17) from the European Nucleotide Archive (accession number PRJEB25514). c, d, h, i, Horizontal lines indicate the median; box boundaries indicate the interquartile range; whiskers represent values within 1.5 × the interquartile range of the first and third quartiles. Each circle represents one sample. In c, e-i, *FDR-adjusted P < 0.05; Wilcoxon rank-sum test. ns, not significant.
Extended Data Fig. 3 The microbiota compositions of centenarians, their lineal descendants and patients with IBD.
Faecal meta 16S rRNA gene sequencing of centenarians (CE, n = 157), older participants (n = 111), young participants (n = 40), lineal relatives of centenarians (CE-L, n = 22, 48-95 years old, average 74.7 years old), and patients with IBD (Crohn’s disease; n = 12, ulcerative colitis; n = 91, 15-78 years old, average 49.0 years old) (all Japanese). a, PCoA plots based on unweighted UniFrac distance among faecal microbiome of centenarian (orange), older (blue), young (grey), CE-L (yellow), and IBD (red) groups. Note that the microbiota composition of centenarians was distinct from that of patients with IBD. b-e, Relative abundance of ASV (amplicon sequence variants) that are significantly enriched in centenarians (b); depleted in centenarians (c); commonly enriched in both centenarians and their lineal relatives (d); and commonly depleted or enriched in both centenarians and young participants (e). The closest species of ASVs were assigned using the National Center for Biotechnology Information Reference Sequence (NCBI RefSeq) database. Data are mean ± s.d. ***P < 0.001; **P < 0.01; *P < 0.05; Wilcoxon rank-sum test. ns, not significant. Each dot represents an individual.
Extended Data Fig. 4 Faecal short-chain fatty acids and pH in centenarians.
a, GC-MS-based quantification of faecal short-chain fatty acids (SCFAs) from centenarian (CE, n = 47, orange), older (n = 31, blue), and young (n = 23, grey) individuals. Faecal SCFAs are shown in μmol/g wet weight faeces. b, c, Faecal ammonia/ammonium (b) and faecal pH levels (c) from individuals within each group. d, Correlation between the levels of secondary bile acids (LCA, 3-oxoLCA, isoLCA, alloLCA, 3-oxoalloLCA, and isoalloLCA) and pH. Each circle represents an individual. Spearman’s coefficient (r) and significance (P) were calculated separately for each group. There is significant positive correlation between pH and faecal alloLCA (P = 0.0156), 3-oxoalloLCA (P = 0.0237), and isoalloLCA (P = 0.0034) in centenarians. In a-c, data are mean ± s.d. ***P < 0.001; **P < 0.01; *P < 0.05; one-way ANOVA with Tukey’s test. ns, not significant.
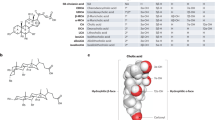
Extended Data Fig. 5 Quantification of faecal bile acids by age group.
Faecal bile acid composition of centenarian (CE, n = 125, orange), older (n = 107, blue), young (n = 47, grey), and lineal relatives of centenarians (CE-L, n = 18, yellow) were profiled and quantified by LC-MS/MS (μmol/g wet weight faeces). In pilot studies, we found that 94 of 137 examined bile acids were minor components of centenarians’ faeces (see Supplementary Table 3). We thus selected the remaining 43 bile acid compounds for follow-up quantitative analysis (see also Fig. 1a). a, Multi-dimensional scaling plot using Spearman’s correlation highlights differences among the four groups’ bile acid profiles. Each circle represents an individual participant from the indicated age group. P = 4.27 × 10−9 for CE versus older; P = 2.72 × 10−12 for CE versus young; P = 4.18 × 10−6 for CE versus CE-L; P = 0.00123 for older versus young; Wilcoxon rank-sum test. b, c, Average ratio of total primary versus secondary (b) and CA- versus CDCA-derived bile acids (c). d, Sum of conjugated and unconjugated bile acids. e, Concentration of each individual bile acid. In d, e, each circle represents an individual. Data are mean ± s.d. ***P < 0.001; **P < 0.01; *P < 0.05; one-way ANOVA with Tukey’s test. ns, not significant. Faecal bile acids are shown in μmol/g wet weight faeces. f, Distribution of participants’ faecal isoalloLCA concentrations. The median faecal isoalloLCA concentration in centenarians was 19.5 μM, meaning that 63 centenarian samples among 125 (50.4%) had >19.5 μM isoalloLCA. In contrast, only 17 older (15.9%) and 3 young (7.7%) participants were found to have >19.5 μM isoalloLCA.
Extended Data Fig. 6 Predicted bile acid biosynthesis by gut microorganisms.
a, Biosynthetic pathway of secondary bile acids metabolized by the gut microbiota from primary bile acid chenodeoxycholic acid (CDCA). Responsible enzymes are indicated within boxes. The glycine or taurine conjugated primary bile acids are deconjugated (not depicted) and biotransformed into a variety of secondary bile acids by the gut microbiota. The predominant biotransformation is 7α-dehydroxylation of CDCA by bai operon genes, thereby converting it into lithocholic acid (LCA). In addition, bile acids can undergo oxidation and epimerization to generate oxo- (keto-), iso- (3β-hydroxy-), allo- (5α-H-), as well as cis- (indicated in a blue box) and trans-forms (indicated in a pink box). Brackets indicate predicted pathways to allo-form LCA production. Chemical structures are simplified by depicting only A and B steroid rings. b, 3-Oxo-Δ4-LCA (also termed 3-oxo-4,5-dehydro-LCA) and 3-oxoalloLCA are structurally similar to testosterone and 5α-dihydrotestosterone (DHT), respectively. Both DHT and 3-oxoalloLCA have A and B steroid rings in a planar (trans) conformation (indicated in a pink box). We predicted that alloLCA and isoalloLCA might be generated from 3-oxo-Δ4-LCA by the sequential action of a 5α-reductase (5AR) homologue and 3α-HSDH (for alloLCA) or 3β-HSDH (for isoalloLCA), through a 3-oxoalloLCA intermediate (see a), analogous to the 5AR-mediated conversion of testosterone into DHT by hydrogenating across the C4-C5 double bond, thereby forcing the A and B steroid rings into a planar conformation. c, Biosynthetic pathway of DCA and related bile acids by the gut microbiota based on ref. 11. 3-OxoDCA can be generated from 3-oxo-Δ4-DCA by hydrogenation across the C4-C5 double bond such that the C5 hydrogen is in the β position. This reaction is mediated by a 3-oxo-5β-steroid 4-dehydrogenase (also termed 5β-reductase, 5BR) encoded by the BaiCD gene. We predicted that LCA and isoLCA might be generated from 3-oxo-Δ4-LCA by the sequential action of a 5BR homologue and 3α-HSDH (for LCA) or 3β-HSDH (for isoLCA), through a 3-oxoLCA intermediate (see panel a), mirroring the previously characterized conversion of 3-oxoDCA to DCA or isoDCA. In a–c, dashed-wedged lines indicate α-positions of -H and -OH groups, while bold-wedged lines indicate β-positions.
Extended Data Fig. 7 Identification of bile acid-metabolizing bacterial strains isolated from the microbiota of a centenarian.
In vitro bile acid metabolism by 68 CE91-derived isolates using 50 μM of CDCA (a), LCA (b), and 3-oxo-Δ4-LCA (c) as starting substrates in pH 9-adjusted media. Data was obtained by LC-MS/MS analysis of 48 h culture supernatants. A list of the 68 isolated strains is shown in the left panel. The closest species to each isolate was identified based on 16S rRNA sequence similarity to the National Center for Biotechnology Information Reference Sequence (NCBI RefSeq) database. Graphs with red backgrounds indicate trans-bile acids and blue backgrounds indicate cis-bile acids. Bile acid profiles after culturing in pH 7-adjusted media are shown in Supplementary Fig. 1. a, Incubation with CDCA did not result in production of target bile acids in any of the cultures, though C. scindens strains 59-60 (St59-60) and C. hylemonae St63 were able to produce LCA, albeit at low levels. b, When cultured with LCA, Gordonibacter pamelaeae St32 and E. lenta St33-35 were found to produce 3-oxoLCA and isoLCA, implying carriage of 3α-HSDH and 3β-HSDH as predicted in a previous study11. In addition, Raoulibacter timonensis St30-31 and Lachnospiraceae spp. St57 were also capable of transforming LCA into 3-oxoLCA, suggesting their possession of 3α-HSDH, similar to E. lenta. c, When 3-oxo-Δ4-LCA was used as a substrate, 3-oxoLCA accumulated to high levels in the supernatants of H. hathewayi St54-55 and Lachnospiraceae spp. St62 cultures, suggesting their carriage of 5BR. Similarly, isoLCA was generated from 3-oxo-Δ4-LCA at high levels in C. innocuum St51 and Lachnospiraceae spp. St58 cultures, suggesting carriage of 5BR and 3β-HSDH. It is noteworthy that P. distasonis St4-5 converted LCA to 3-oxoLCA and further to 3-oxo-Δ4-LCA, as well as 3-oxo-Δ4-LCA to isoLCA and LCA, suggesting that these strains possess 3α-HSDH, 3β-HSDH, and 5BR. Data are mean + s.d. of duplicate samples and representative of two independent experiments.
Extended Data Fig. 8 Elucidated biosynthetic pathway of secondary bile acids.
a, Elucidated biosynthetic pathway of secondary bile acids metabolized by CE91-derived bacterial strains. Colour-coded lines indicate in vitro identified metabolic capabilities of each strain. Responsible enzymes are indicated within boxes. Chemical structures are simplified by depicting only A and B steroid rings. b, The genome sequences of 68 isolates from a centenarian (CE91) were determined using PacBio Sequel and Illumina MiSeq sequencers. The gene prediction and annotation of the generated contigs were performed using the Rapid Annotations using Subsystem Technology (RAST) server. Gene clusters containing 5AR (magenta), 5BR (blue), 3β-HSDH-I (green), and 3β-HSDH-II (purple) homologues identified in 23 Bacteroidales strains are shown. Note that the gene clusters containing 5AR, 5BR, 3β-HSDH-I and 3β-HSDH-II homologues are present in close proximity to genes annotated with tricarboxylic acid (TCA) cycle-related functions (beige) and transporter/efflux system membrane fusion proteins (yellow). Gene homologues are defined as similarity of <1 × 10−12 E-value, >30% sequence identity, and >60% query coverage. Arrows represent coding sequences and annotated functions are colour-coded accordingly. Scale bar is 2kb. Note that 3α-HSDH homologue is not present in any of the 5AR clusters.
Extended Data Fig. 9 Metabolism of LCA-related compounds by Bacteroidales strains and cooperative isoalloLCA production.
a-c, Bacteroidales strains were cultured with 3-oxoalloLCA (a), 3-oxoLCA (b), or isoLCA (c) at a final concentration of 50 μM in pH 9-adjusted WCA medium. There was substantial substrate specificity and strain-to-strain variation in transformation efficiency. For instance, P. merdae St3, P. distasonis St4–5, and Odoribacteraceae St21 exhibited strong 3β-HSDH activity, reflected by both high isoalloLCA production from 3-oxoalloLCA (a) and isoLCA production from 3-oxoLCA (b), whereas other strains showed less efficient biotransformation or substrate specificity despite carriage of putative 3β-HSDH genes. The strength of 5BR activity also differed among the strains: P. merdae St3, P. distasonis St4–5, B. dorei St6–7, and B. uniformis St10-13 effectively transformed 3-oxoLCA to 3-oxo-Δ4-LCA (and further to 3-oxoalloLCA by 5AR), while other strains displayed moderate to weak activity (b). Porphyromonas somerae St14 lacked a putative 3β-HSDH gene but was able to generate isoalloLCA from 3-oxoalloLCA nonetheless (a), suggesting that it carries a strain-specific gene with 3β-HSDH activity. d, Co-culture of P. merdae St3 or Odoribacteraceae St21 (5BR, 5AR, and 3β-HSDH encoders) with E. lenta St34 (top, green, a 3α-HSDH and 3β-HSDH encoder), or P. distasonis St4 (bottom, yellow, a 3α-HSDH, 3β-HSDH, and 5BR encoder) supplemented with 50 μM LCA in pH 9-adjusted WCA medium. Notably, co-culture of P. merdae St3 and E. lenta St34 with LCA resulted in generation of alloLCA in addition to isoalloLCA. e, Different combinations of P. merdae St3, Odoribacteraceae St21, or C. scindens St59 (a bai operon encoder capable of converting CDCA to LCA) co-cultured with E. lenta St34 (top) or P. distasonis St4 (bottom) in the presence of 50 μM CDCA. In a–e, the presence of genes homologous to 5AR, 5BR, 3β-HSDH-I, 3β-HSDH-II, 3α-HSDH, or the bai operon in corresponding combinations of strains are indicated as grey boxes within the adjacent charts. Note that the bai operon in C. scindens contains 5BR and 3α-HSDH genes. Culture supernatants were collected after 48 h of anaerobic incubation at 37 °C for LC-MS/MS quantification. Data are mean + s.d. of duplicate samples and representative of two independent experiments.
Extended Data Fig. 10 IsoalloLCA inhibits Gram-positive pathogens.
a, b, Gram-positive (upper) and -negative (lower) pathogens were incubated with varying concentrations of the indicated bile acids under anaerobic conditions at 37 °C in WCA medium until each strain reached stationary growth phase in the control medium (5-75 h). Bacterial growth was determined by OD600 measurement. Maximum growth densities (a) and growth curves (b) of pathogens in varying concentrations of bile acids are shown. Data are mean ± s.d. (error bars shown with fill area). c, MIC90 value (minimal inhibitory concentration required to prevent 90% growth) of isoalloLCA for each pathogenic strain was determined by serially diluting the compound and incubating each dilution with the pathogen. Shaded area indicates isoalloLCA concentrations with growth inhibitory effects, and MIC90 is reported inside the shaded box. Data are mean ± s.d. of duplicate samples and are representative of two independent experiments. d, In vitro growth inhibition of C. difficile 630 and vancomycin-resistant E. faecium (VRE) by co-culturing with CE91-derived isolates in the presence or absence of 12.5 μM 3-oxo-Δ4-LCA in WCA medium. Average CFU of overnight cultures are shown (n = 6). Data are mean ± s.d. and are representative of two independent experiments. ***P < 0.001; Mann–Whitney test (two-tailed) with Welch’s correction. ns, not significant.
Extended Data Fig. 11 In vivo suppression of faecal C. difficile shedding in Cyp2a12−/−Cyp2c70−/− mice.
a, b, Faecal and plasma concentrations of isoalloLCA (a) and plasma analytes (b) from SPF C57BL/6N mice after 14 days on an 0.01%, 0.05%, or 0.1% (w/w) isoalloLCA-containing diet. Plasma analytes from each animal: albumin; aspartate aminotransferase (AST); alanine transaminase (ALT); lactate dehydrogenase (LDH); total bilirubin; cholinesterase; total cholesterol; and total protein. Data are mean ± s.d.; n = 3 each; ***P < 0.001; **P < 0.01; *P < 0.05; one-way ANOVA with Tukey’s test. ns, not significant. c, SPF C57BL/6N mice were pretreated with cefoperazone through the drinking water during the period from 7 to 2 days before inoculation and then infected with C. difficile 630 by oral gavage on day 0. The mice were subsequently placed on an 0.1% isoalloLCA-containing diet from day 1 to day 3 after infection (p.i.) (n = 17-18 animals each). Faecal C. difficile CFUs and isoalloLCA levels throughout the course of infection were determined. Data are mean ± s.d. ***P < 0.001; Mann–Whitney test (two-tailed) with Welch’s correction. d, Simplified pathway of bile acid metabolism in rodents. In rodents’ livers, CDCA is immediately hydroxylated at the 6β-position to generate muricholic acids (α- and β-muricholic acids (MCAs)) by CYP2C70, and as such we employed a Cyp2c70−/− mouse model. As resulting excessive CDCA causes hepatotoxicity in Cyp2c70−/− mice, we cross-bred Cyp2c70−/− mice with Cyp2a12−/− mice lacking 7α-rehydroxylation capacity to minimize liver injury. e-g, Faecal bile acids of SPF wild-type C57BL/6N mice (WT) and Cyp2a12−/−Cyp2c70−/− double-knockout (CypDKO) mice (n = 3 each) were quantified by LC-MS/MS (μmol/g wet weight faeces). The percent of each bile acid among total faecal bile acids from each mouse was calculated (e). Note that Cyp2a12−/−Cyp2c70−/− double-knockout mice showed elevated levels of LCA and related bile acids. Amount of total faecal bile acids (f) and each bile acid (g) from each mouse. Data are mean ± s.d. ***P < 0.001; **P < 0.01; *P < 0.05; one-way ANOVA with Tukey’s test. ns, not significant. Each dot represents an individual.
Extended Data Fig. 12 MIC90 of isoalloLCA against gut commensals in WCA and BHI media.
a, Growth inhibitory effect of isoalloLCA on gut commensals in WCA medium. Gram-positive and Gram-negative commensal isolates were selected from our culture collection and incubated anaerobically at 37 °C with the indicated concentrations of isoalloLCA. Bacterial growth was determined by OD600 measurement. Each strain was cultured until it reached stationary growth phase in the control medium (7-48 h). Shaded area indicates isoalloLCA concentrations with growth inhibitory effects. MIC90 is shown inside the shaded box. b, MIC90 of isoalloLCA against Gram-positive and Gram-negative commensals in WCA and BHI media. c, Representative scanning electron microscopy images of C. difficile 630, C. sporogenes, C. indolis and Clostridium HGF2 (innocuum) grown in control or 2.5 μM isoalloLCA-containing WCA medium for 5 h. The right panels show the corresponding high-resolution images. Scale bars are 5.0 μm and 1.0 μm. Arrows indicate morphological alterations after isoalloLCA treatment. d, The differences in growth inhibitory effect of isoalloLCA between WCA and BHI medium on C. difficile 630, E. faecium (VRE), S. dysgalactiae subsp. equisimilis (SDSE), C. symbiosum, C. scindens and Clostridium HGF2 (innocuum). Data in a, d are mean ± s.d. from duplicate samples and are representative of two independent experiments.
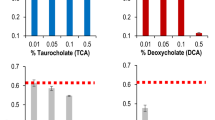
Extended Data Fig. 13 The effects of isoalloLCA on the complex gut microbiota.
a, b, f–h, Human faecal samples from healthy young donors (a, b) and patients with C. difficile infection (CDI) (f–h) were incubated for 48 h in modified WCA medium supplemented with 3-oxoLCA, LCA, or isoalloLCA (50 μM). Shannon index of diversity (a and c) and a compositional shift in the microbiome at the genus level (b and g) of faecal cultures after secondary bile acid treatment. Each dot in a and f represents a single donor’s faecal culture. ns, not significant by Kruskal–Wallis with Dunn’s test. Although α-diversity was not significantly affected, isoalloLCA induced broad changes in microbial community structure, with a significant alteration in the ratio of Gram-negative to Gram-positive bacteria. C. difficile CFUs in each CDI microbiota pre- or post- 48 h incubation with medium or secondary bile acids were determined (h). Data are mean ± s.d. of duplicate samples. **P < 0.01; *P < 0.05; Unpaired t-test. ns, not significant. c–e, Correlation between the levels of faecal isoalloLCA and gut MSPs in centenarian (CE, n = 119, orange), older (n = 107, blue), and young (n = 39, grey) participants. Spearman’s coefficient (r) and significance (P) for msp_103 A. putredinis and msp_095 O. laneus (both 5AR and 3β-HSDH encoders) (c), combined Streptococcus signal (d), and Streptococcus species (e). Each dot represents an individual. Several Streptococcus spp. showed a negative association with isoalloLCA concentration in our cohort.
Extended Data Fig. 14 Associations between species encoding cluster V1, V2 or V3, or non-encoders with secondary bile acids.
a, Relative abundance of gut MSPs categorized into clusters V1, V2, and V3 from the assembled gut metagenomes of centenarian (CE, n = 119, orange), older (n = 107, blue), and young (n = 39, grey) groups. b, c, Spearman’s coefficient (r) and significance (P) between faecal bile acid level and summed relative abundance of cluster V1-, V2-, or V3-encoding species (b) and ratio of species encoding clusters V1 and V3 versus V2 from assembled gut metagenomes (c). Each dot represents an individual. d, Spearman correlation coefficient (r) for each bile acid against species encoding cluster V1, V2, or V3, or non-encoders. Non-encoders that correlate strongly with bile acid concentration are indicated within a grey circle, and the species are listed.
Extended Data Fig. 15 Stratification into microbial community types with respective microbiome characteristics.
a, Principal coordinate analysis of three microbial community types (1, 2, and 3) stratified using Dirichlet Multinomial Mixtures from the gut microbiomes of centenarian [CE, n = 176 (153 individuals)], older (n = 110), and young (n = 44) individuals. b, c, Relative abundances across phyla (b) and top differentially abundant species from each phylum (c) from the assembled gut metagenomes in each community type. Community type 1 is characterized by a high relative abundance of Firmicutes (for example, Oscillibacter spp.) and Proteobacteria (for example, Desulfovibrio spp.), whereas community type 2 exhibits relative enrichment of Actinobacteria and depletion of Proteobacteria. Community type 3 is structurally similar to type 1, but exhibits a higher abundance of Verrucomicrobia (for example, Akkermansia spp.) and Euryarchaeota (for example, Methanobrevibacter spp.). d, Species harbouring gene clusters V1, V2, and V3 from the assembled gut metagenomes in each community type. Each dot in c and d represents an individual from the centenarian (orange), older (blue), or young (grey) groups. Horizontal lines indicate the median; box boundaries indicate interquartile range (IQR); whiskers represent values within 1.5 x IQR of the first and third quartiles. Asterisks indicate significantly different abundance in the specified comparison at FDR P < 0.05 based on a Wilcoxon rank-sum test. ns, not significant. e, Longitudinal change in faecal bile acid composition from the same individual over the course of 1-2 years. Upper numbers indicate collection interval (days). Faecal bile acids were quantified by LC-MS/MS (μmol/g wet weight faeces), and the percent of each bile acid among total faecal bile acids in each sample was calculated. f, Stability of microbial community type (1, 2, and 3) in each individual over time.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-4, Supplementary Tables 1, 5, 8, 9, and legends for Supplementary Tables 2, 3, 4, 6 and 7.
Supplementary Table 2
Quantification of 16 faecal fatty acids by GC-MS.
Supplementary Table 3
A list of 137 bile acid compounds used in this study.
Supplementary Table 4
Genome and 16S rRNA gene sequencing data of 68 CE91-derived isolates.
Supplementary Table 6
Detailed LC-MS/MS setting for in vivo samples (48 bile acid compounds) using SCIEX Triple Quad™ 6500+ LC-MS/MS system.
Supplementary Table 7
Detailed LC-MS/MS setting for in vitro samples (14 bile acid compounds) using Shimadzu LCMS-8040 system.
Source data
Rights and permissions
About this article
Cite this article
Sato, Y., Atarashi, K., Plichta, D.R. et al. Novel bile acid biosynthetic pathways are enriched in the microbiome of centenarians. Nature 599, 458–464 (2021). https://doi.org/10.1038/s41586-021-03832-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03832-5
This article is cited by
-
Intestinal metabolites predict treatment resistance of patients with depression and anxiety
Gut Pathogens (2024)
-
The gut-liver axis in hepatobiliary diseases
Inflammation and Regeneration (2024)
-
The changing metabolic landscape of bile acids – keys to metabolism and immune regulation
Nature Reviews Gastroenterology & Hepatology (2024)
-
Effects of bathing in different hot spring types on Japanese gut microbiota
Scientific Reports (2024)
-
Another renaissance for bile acid gastrointestinal microbiology
Nature Reviews Gastroenterology & Hepatology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.