Abstract
Changing food preferences brought about by westernization that have deleterious health effects1,2—combined with myriad forces that are contributing to increased food insecurity—are catalysing efforts to identify more nutritious and affordable foods3. Consumption of dietary fibre can help to prevent cardiovascular disease, type 2 diabetes and obesity4,5,6. A substantial number of reports have explored the effects of dietary fibre on the gut microbial community7,8,9. However, the microbiome is complex, dynamic and exhibits considerable intra- and interpersonal variation in its composition and functions. The large number of potential interactions between the components of the microbiome makes it challenging to define the mechanisms by which food ingredients affect community properties. Here we address the question of how foods containing different fibre preparations can be designed to alter functions associated with specific components of the microbiome. Because a marked increase in snack consumption is associated with westernization, we formulated snack prototypes using plant fibres from different sustainable sources that targeted distinct features of the gut microbiomes of individuals with obesity when transplanted into gnotobiotic mice. We used these snacks to supplement controlled diets that were consumed by adult individuals with obesity or who were overweight. Fibre-specific changes in their microbiomes were linked to changes in their plasma proteomes indicative of an altered physiological state.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Data availability
V4-16S rRNA sequences in raw format before post-processing and data analysis, plus shotgun sequencing datasets generated from faecal DNA, have been deposited at the European Nucleotide Archive under study accession PRJEB38148. Raw proteomic datasets generated from the aptamer-based 1.3K SomaLogic platform have been deposited in the European Genome-Phenome Archive (EGA) under accession IDs EGAD00010002133 (pea fibre study) and EGAD00010002132 (fibre blends study). Any other relevant data are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
Code for HOSVD (CP-ALS plus randomization code) and CC-SVD is available via Zenodo (https://doi.org/10.5281/zenodo.4767887).
References
NCD Risk Factor Collaboration (NCD-RisC). Trends in adult body-mass index in 200 countries from 1975 to 2014: a pooled analysis of 1698 population-based measurement studies with 19·2 million participants. Lancet 387, 1377–1396 (2016).
GBD 2017 Diet Collaborators. Health effects of dietary risks in 195 countries, 1990-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet 393, 1958–1972 (2019).
Willett, W. et al. Food in the Anthropocene: the EAT-Lancet Commission on healthy diets from sustainable food systems. Lancet 393, 447–492 (2019).
Hauner, H. et al. Evidence-based guideline of the German Nutrition Society: carbohydrate intake and prevention of nutrition-related diseases. Ann. Nutr. Metab. 60 (Suppl 1), 1–58 (2012).
Reynolds, A. et al. Carbohydrate quality and human health: a series of systematic reviews and meta-analyses. Lancet 393, 434–445 (2019).
Zhao, L. et al. Gut bacteria selectively promoted by dietary fibers alleviate type 2 diabetes. Science 359, 1151–1156 (2018).
Asnicar, F. et al. Microbiome connections with host metabolism and habitual diet from 1,098 deeply phenotyped individuals. Nat. Med. 27, 321–332 (2021).
Kovatcheva-Datchary, P. et al. Dietary fiber-induced improvement in glucose metabolism is associated with increased abundance of Prevotella. Cell Metab. 22, 971–982 (2015).
Sonnenburg, E. D. et al. Specificity of polysaccharide use in intestinal Bacteroides species determines diet-induced microbiota alterations. Cell 141, 1241–1252 (2010).
Ridaura, V. K. et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 341, 1241214 (2013).
Patnode, M. L. et al. Interspecies competition impacts targeted manipulation of human gut bacteria by fiber-derived glycans. Cell 179, 59–73.e13 (2019).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Overbeek, R. et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 42, D206–D214 (2014).
Martino, C. et al. Context-aware dimensionality reduction deconvolutes gut microbial community dynamics. Nat. Biotechnol. 39, 165–168 (2021).
Wesener, D. A. et al. Microbiota functional activity biosensors for characterizing nutrient metabolism in vivo. eLife 10, e64478 (2021).
Temple, M. J. et al. A Bacteroidetes locus dedicated to fungal 1,6-β-glucan degradation: Unique substrate conformation drives specificity of the key endo-1,6-β-glucanase. J. Biol. Chem. 292, 10639–10650 (2017).
Larsbrink, J. et al. A discrete genetic locus confers xyloglucan metabolism in select human gut Bacteroidetes. Nature 506, 498–502 (2014).
Schröder, C. et al. Characterization of a theme C glycoside hydrolase family 9 endo-beta-glucanase from a biogas reactor metagenome. Protein J. 37, 454–460 (2018).
Shimizu, H. et al. Characterization and structural analysis of a novel exo-type enzyme acting on β-1,2-glucooligosaccharides from Parabacteroides distasonis. Biochemistry 57, 3849–3860 (2018).
Li, W. et al. PspAG97A: a halophilic α-glucoside hydrolase with wide substrate specificity from glycoside hydrolase family 97. J. Microbiol. Biotechnol. 26, 1933–1942 (2016).
Gloster, T. M., Turkenburg, J. P., Potts, J. R., Henrissat, B. & Davies, G. J. Divergence of catalytic mechanism within a glycosidase family provides insight into evolution of carbohydrate metabolism by human gut flora. Chem. Biol. 15, 1058–1067 (2008).
Helbert, W. et al. Discovery of novel carbohydrate-active enzymes through the rational exploration of the protein sequences space. Proc. Natl Acad. Sci. USA 116, 6063–6068 (2019).
Ndeh, D. et al. Complex pectin metabolism by gut bacteria reveals novel catalytic functions. Nature 544, 65–70 (2017).
Hashimoto, W., Miyake, O., Ochiai, A. & Murata, K. Molecular identification of Sphingomonas sp. A1 alginate lyase (A1-IV′) as a member of novel polysaccharide lyase family 15 and implications in alginate lyase evolution. J. Biosci. Bioeng. 99, 48–54 (2005).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Derecka, M. et al. Tyk2 and Stat3 regulate brown adipose tissue differentiation and obesity. Cell Metab. 16, 814–824 (2012).
Pedersen, L., Olsen, C. H., Pedersen, B. K. & Hojman, P. Muscle-derived expression of the chemokine CXCL1 attenuates diet-induced obesity and improves fatty acid oxidation in the muscle. Am. J. Physiol. Endocrinol. Metab. 302, E831–E840 (2012).
Kraja, A. T. et al. Genetic analysis of 16 NMR-lipoprotein fractions in humans, the GOLDN study. Lipids 48, 155–165 (2013).
ZhuGe, D. L., Javaid, H. M. A., Sahar, N. E., Zhao, Y. Z. & Huh, J. Y. Fibroblast growth factor 2 exacerbates inflammation in adipocytes through NLRP3 inflammasome activation. Arch. Pharm. Res. 43, 1311–1324 (2020).
Michalak, L. et al. Microbiota-directed fibre activates both targeted and secondary metabolic shifts in the distal gut. Nat. Commun. 11, 5773 (2020).
Bucholz, K. K., Heath, A. C. & Madden, P. A. Transitions in drinking in adolescent females: evidence from the Missouri adolescent female twin study. Alcohol. Clin. Exp. Res. 24, 914–923 (2000).
Mifflin, M. D. et al. A new predictive equation for resting energy expenditure in healthy individuals. Am. J. Clin. Nutr. 51, 241–247 (1990).
Subar, A. F. et al. Comparative validation of the Block, Willett, and National Cancer Institute food frequency questionnaires: the Eating at America’s Table Study. Am. J. Epidemiol. 154, 1089–1099 (2001).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl Acad. Sci. USA 108 (Suppl 1), 4516–4522 (2011).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2017).
Di Luccia, B. et al. Combined prebiotic and microbial intervention improves oral cholera vaccination responses in a mouse model of childhood undernutrition. Cell Host Microbe 27, 899–908.e5 (2020).
Baym, M. et al. Inexpensive multiplexed library preparation for megabase-sized genomes. PLoS ONE 10, e0128036 (2015).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnetJ. 17, 10–12 (2011
Joshi, N. A. & Fass, J. N. Sickle: A Sliding-Window, Adaptive, Quality-based Trimming tool for FastQ Files (Version 1.33) Software (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Peng, Y., Leung, H. C. M., Yiu, S. M. & Chin, F. Y. L. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012).
Li, D., Liu, C.-M., Luo, R., Sadakane, K. & Lam, T.-W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Mistry, J., Finn, R. D., Eddy, S. R., Bateman, A. & Punta, M. Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Res. 41, e121 (2013).
Chen, R. Y. et al. Duodenal microbiota in stunted undernourished children with enteropathy. N. Engl. J. Med. 383, 321–333 (2020).
Plerou, V. et al. Random matrix approach to cross correlations in financial data. Phys. Rev. E 65, 066126 (2002).
Winkler, E. S. et al. Human neutralizing antibodies against SARS-CoV-2 require intact Fc effector functions for optimal therapeutic protection. Cell 184, 1804–1820.e16 (2021).
Zou, W. et al. Ablation of fat cells in adult mice induces massive bone gain. Cell Metab. 32, 801–813.e6 (2020).
Adamo, L. et al. Proteomic signatures of heart failure in relation to left ventricular ejection fraction. J. Am. Coll. Cardiol. 76, 1982–1994 (2020).
Tsingas, M. et al. Sox9 deletion causes severe intervertebral disc degeneration characterized by apoptosis, matrix remodeling, and compartment-specific transcriptomic changes. Matrix Biol. 94, 110–133 (2020).
Joly, J. H., Lowry, W. E. & Graham, N. A. Differential gene set enrichment analysis: a statistical approach to quantify the relative enrichment of two gene sets. Bioinformatics 36, 5247–5254 (2020).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
Blakeney, A. B., Harris, P. J., Henry, R. J. & Stone, B. A. A simple and rapid preparation of alditol acetates for monosaccharide analysis. Carbohydr. Res. 113, 291–299 (1983).
Englyst, H. N. & Cummings, J. H. Improved method for measurement of dietary fiber as non-starch polysaccharides in plant foods. J. Assoc. Off. Anal. Chem. 71, 808–814 (1988).
Blumenkrantz, N. & Asboe-Hansen, G. New method for quantitative determination of uronic acids. Anal. Biochem. 54, 484–489 (1973).
Thibault, J.-F. Automatisation du dosage des substances pectiques par la méthode au métahydroxydiphényle. Lebensm. Wiss. Technol. 12, 247–251 (1979).
Filisetti-Cozzi, T. M. C. C. & Carpita, N. C. Measurement of uronic acids without interference from neutral sugars. Anal. Biochem. 197, 157–162 (1991).
Levigne, S., Thomas, M., Ralet, M.-C., Quemener, B. & Thibault, J.-F. Determination of the degrees of methylation and acetylation of pectins using a C18 column and internal standards. Food Hydrocoll. 16, 547–550 (2002).
Pettolino, F. A., Walsh, C., Fincher, G. B. & Bacic, A. Determining the polysaccharide composition of plant cell walls. Nat. Protoc. 7, 1590–1607 (2012).
Buffetto, F. et al. The deconstruction of pectic rhamnogalacturonan I unmasks the occurrence of a novel arabinogalactan oligosaccharide epitope. Plant Cell Physiol. 56, 2181–2196 (2015).
Amicucci, M. J. et al. A rapid-throughput adaptable method for determining the monosaccharide composition of polysaccharides. Int. J. Mass Spectrom. 438, 22–28 (2019).
Xu, G., Amicucci, M. J., Cheng, Z., Galermo, A. G. & Lebrilla, C. B. Revisiting monosaccharide analysis - quantitation of a comprehensive set of monosaccharides using dynamic multiple reaction monitoring. Analyst 143, 200–207 (2018).
Galermo, A. G. et al. Liquid chromatography-tandem mass spectrometry approach for determining glycosidic linkages. Anal. Chem. 90, 13073–13080 (2018).
Galermo, A. G., Nandita, E., Castillo, J. J., Amicucci, M. J. & Lebrilla, C. B. Development of an extensive linkage library for characterization of carbohydrates. Anal. Chem. 91, 13022–13031 (2019).
Cowardin, C. A. et al. Mechanisms by which sialylated milk oligosaccharides impact bone biology in a gnotobiotic mouse model of infant undernutrition. Proc. Natl Acad. Sci. USA 116, 11988–11996 (2019).
Acknowledgements
We thank M. Karlsson and D. O’Donnell for their assistance with gnotobiotic mouse husbandry; S. Marion for her role in collecting clinical metadata and faecal samples from members of the MOAFTs study cohort with obesity, and the two human studies; S. Waller for design of the controlled diet study menus and oversight of the metabolic kitchen; S. Torbitzky for her assistance with coordination and planning of the human studies; M. Patnode for his input about the selection of lead fibres; S. Le Gall, L. Saulnier and B. Laillet for carbohydrate analysis of pea, orange and barley bran fibre preparations; G. Cesbron Lavau and M. Okoniewska for formulation and scale-up production of snack prototypes for human studies; S. Abdel-Hamid for production of the HiSF–LoFV diets given to gnotobiotic mice; L. Dimartino for analysis of the fibre ingredients in these diets; and L. Kyro for assistance with figure illustrations. Technical support was provided by S. Deng, J. Serugo, J. Lelwala-Guruge, K. Ahsan, S. Bale, J. Veitinger, J. Forman and S. Karlsson (archiving and processing mouse and human biospecimens), M. Meier, J. Hoisington-López and M. Crosby (bacterial V4-16S rDNA amplicon and faecal microbiome shotgun sequencing), T. Juehne, A. Lutz and J. Yu (generating SOMAscan datasets), and R. Head and C. Storer (assistance with CompBio analyses). This work was funded by grants from the NIH (DK078669, DK70977, UL1-TR002345) and from Mondelēz Global LLC. Controlled diet studies of human participants were overseen by members of the Clinical Science Research Core of the Nutrition Obesity Research Center (NORC), which is supported by NIH grant P30 DK056341. Plasma proteomic data sets were generated by the Genome Technology Access Center at Washington University School of Medicine, which is supported in part by NIH Grants P30 CA91842 and UL1TR002345. O.D.-B. received support from NIH R25GM103757, T32GM007067 and T32HL130357 as a pre-doctoral trainee. J.I.G. is the recipient of a Thought Leader Award from Agilent Technologies.
Author information
Authors and Affiliations
Contributions
O.D.-B. and J.I.G. designed the gnotobiotic mouse studies. A.C.H. oversaw collection of faecal samples from human donors with obesity used to colonize germ-free mice. O.D.-B. and N.H. performed mouse studies. M.J.B., S.K., O.D.-B. and J.I.G. designed the human studies together with D.K.H., A.M. and S.V., who oversaw the design, manufacture and quality-control analysis of the fibre snack prototypes used in the two human studies. A.M. and S.V. organized carbohydrate and glycosidic linkage composition analysis of fibre preparations. Controlled diet studies of human participants were overseen by S.K. together with K.K. and T.W. J.J.C., G.C. and C.B.L. conducted mass spectrometric assays of mouse diets and faecal samples. J.C. performed LC–QTOF–MS analyses of human faecal samples collected from participants consuming the two- and four-fibre-containing snacks. O.D.-B. oversaw the archiving and processing of mouse and human biospecimens and generated the 16S rDNA and shotgun sequencing datasets from these samples. M.C.H. and C.D. implemented the metagenomic assembly and annotation pipeline. D.A.R., S.A.L. and A.O. performed mcSEED pathway reconstructions of faecal microbiomes, and V.L. and B.H. provided CAZyme annotations. A.S.R. developed the HOSVD and R.Y.C. the CC-SVD analytic platforms that were applied to datasets generated from mice and humans. O.D.-B. and R.A.B. performed CompBio analyses of plasma proteome datasets generated from human studies. O.D.-B., C.D., M.J.B. and J.I.G. analysed the data. O.D.-B. and J.I.G. wrote the paper with assistance provided by the co-authors.
Corresponding author
Ethics declarations
Competing interests
J.I.G. is a co-founder of Matatu, Inc., a company characterizing the role of diet-by-microbiota interactions in animal health. A.O. and D.R. are co-founders of Phenobiome Inc., a company pursuing development of computational tools for predictive phenotype profiling of microbial communities. C.B.L. is a co-founder of Evolve Biosystems, interVenn Bio and BCD Bioscience, companies involved in the characterization of glycans and developing carbohydrate applications for human health. D.K.H., A.M. and S.V. are employees of Mondelēz Global LLC, a multinational company engaged in production of snack foods. The remaining authors declare that they have no competing financial interests. A patent application related to the fibre-snack formulations described in this Article has been filed and published (WO 2021/016129).
Additional information
Peer review information Nature thanks David Bolam, Eran Elinav and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Monosaccharide content and glycosidic linkages present in the fibre preparations, and in the unsupplemented and fibre-supplemented HiSF–LoFV diets fed to gnotobiotic mice.
a, b, Monosaccharides and linkages, some represented by their methylated monosaccharide derivatives, in the fibre preparations. Stacked bars represent the mean for technical replicates (n = 3) for each glycosyl linkage determination. c, Monosaccharides in the unsupplemented and fibre-supplemented HiSF–LoFV diets. Bars represent the mean ± s.d. for technical replicates (n = 3). *P < 0.05, **P < 0.01, ****P < 0.0001 compared to the unsupplemented diet phase (one-way ANOVA with Holm–Šídák multiple comparison correction). d, Linkages, represented by their methylated monosaccharide derivatives in the unsupplemented and fibre-supplemented HiSF–LoFV diets. Stacked bars represent the mean for technical replicates (n = 3) for each glycosidic linkage determination. Glc, glucose; Gal, galactose; GalA, galacturonic acid; GlcA, glucuronic acid; Ara, arabinose; Xyl, xylose; Man, mannose; Fru, fructose; Fuc, fucose; Rha, rhamnose; Rib, ribose; Hex, hexose; dHex, deoxyhexose; T, terminal; f, furanose; p, pyranose; X, undefined linkage.
Extended Data Fig. 2 The effects of dietary fibres in gnotobiotic mice fed a HiSF-LoFV diet and colonized with faecal microbial communities of nine human donors with obesity.
a, Experimental design. b, HOSVD. Three-dimensional matrices are termed ‘tensors’. A tensor (O) with dimensions n, m and p, in which n represents subjects (mice or humans), m represents features (CAZymes, mcSEED metabolic pathways and ASVs) and p represents time, can be analysed by HOSVD in which a ‘core tensor’ (G) is created—a tensor in which the only non-zero values are along the diagonal (boxes shown in G). Each box represents a ‘tensor component’ (TC). Each tensor component relates the variation between each axis of the O tensor. Additionally, three new matrices are created that are related to each other through each tensor component; for example, ‘Projection of variables onto TC1’ indicates that variation defined by tensor component 1 is defined by variation across the first row of n, the first column of m and the first column of p. c–e, HOSVD applied to CAZymes in faecal microbiomes of mice colonized with microbial communities of nine human donors with obesity (n = 348 faecal samples analysed) during each of the three dietary fibre interventions in the diet oscillation experiment. f, Heat map of discriminatory CAZymes with log2-tranformed fold changes in abundance that were defined as statistically significant during at least one dietary intervention. The grand mean of the data is shown for mice containing the nine different human donor microbiomes sampled at the indicated time points and normalized to day 14 values (n = 6–10 mice per group; n = 232 faecal samples analysed). The order of CAZymes from left to right is based on their function (rows below) and magnitude of their change within and across fibre treatments. ‡q value < 0.10; *q value < 0.05 (linear mixed-effects model, FDR-corrected).
Extended Data Fig. 3 Responses of CAZymes and mcSEED metabolic pathways identified by HOSVD analysis as discriminatory for fibre snack consumption in gnotobiotic mice colonized with the faecal communities of nine human donors with obesity.
a, b, Heat map of discriminatory CAZymes and mcSEED metabolic pathways with changes in abundance that were statistically significant during at least one dietary intervention. Data are averaged for mice containing a given human donor microbiota (n = 6 to 10 mice per group; n = 232 faecal samples analysed) sampled at the indicated time points and normalized to day-14 values. The order of CAZymes from left to right of the heat map in a follows the same order as in Extended Data Fig. 2f, and the order of mcSEED pathways from top to bottom in b follows the same order as shown in Fig. 1d. Hierarchical clustering (Euclidean distances) of CAZyme and mcSEED metabolic pathway profiles was used to group donor microbiomes.
Extended Data Fig. 4 HOSVD applied to mcSEED metabolic pathway and ASV datasets generated from the faecal microbial communities of mice containing microbial communities of human donors with obesity during the pea, orange and barley bran fibre phases of the diet oscillation.
a, Microbiome configurations as defined by the representation of mcSEED metabolic pathways on tensor component 1 and tensor component 2 during pea fibre, orange fibre and barley bran phase of diet oscillation. b, Projections of microbiota configuration as defined by the representation of bacterial taxa (ASVs) on tensor component 1 and tensor component 2 during pea fibre, orange fibre and barley bran phases of the diet oscillation protocol (n = 9 human microbiomes; n = 6–10 mouse recipients of each human microbiome; n = 348 faecal samples analysed for data presented in a, b).
Extended Data Fig. 5 Responses of bacterial taxa (ASVs) identified by HOSVD as discriminatory for dietary fibre consumption in gnotobiotic mice colonized with human donor microbiota, and in human participants enrolled in the controlled-diet studies.
a, Bray–Curtis dissimilarity distances calculated from the ASV content of communities sampled at all time points (days 4, 9, 14, 19, 24, 29, 34, 39, 44, 49, 54, 59 and 64 after colonization) from a given group of recipient mice compared to the ASV content of their corresponding human donor community before transplantation (Bray–Curtis distances calculated from ASV abundances in faecal samples collected from each group of mice (n = 6–10 mice; n = 752 faecal samples in total) compared to the abundances of these ASVs in each of their corresponding nine human-donor faecal communities). ****P value < 0.0001 (one-way ANOVA, Šídák’s correction). b, Heat map of statistically significant log2-transformed fold changes in the abundances of discriminatory ASVs in gnotobiotic mice during at least one of the fibre interventions. The heat map on the left shows the grand mean for data obtained from all groups of mice and the heat map on the right shows averaged data for mice containing a given donor microbiota (n = 6 to 10 mice per group; n = 232 faecal samples obtained at the indicated time points with data normalized to day-14 values). Hierarchical clustering (Euclidean distances) of ASV profiles was used to group donor microbiota with similar responses to each fibre supplement. c, ASVs with log2-transformed fold changes in abundance that were statistically significant in human participants after at least one of the fibre snack interventions. Left, mean values for participants enrolled in each study; the three panels to the right show changes in ASV abundances in individual participants after consumption of each of the fibre snacks. Data are normalized to pretreatment time points, that is, day 14 (study 1) and day 11 (study 2) (n = 12 and 14 participants for study 1 and 2, respectively, n = 66 faecal samples analysed). Hierarchical clustering (Euclidean distances) of ASV profiles was used to group participants with similar responses to a given fibre snack. ‡q value < 0.1, *q value < 0.05 (linear mixed-effects model, FDR-corrected).
Extended Data Fig. 6 Identification by HOSVD of fibre-snack-discriminatory CAZymes and mcSEED metabolic pathways in human participants in the controlled-diet studies.
a, b, CAZymes and mcSEED metabolic pathways with log2-transformed fold changes in abundance were statistically significant during at least one of the three fibre snack interventions. Data are shown for each participant after consumption of each fibre snack type and are normalized to pretreatment time points, that is, day 14 (study 1) and day 11 (study 2) (n = 12 and 14 participants for study 1 and 2, respectively, n = 66 faecal samples analysed). Hierarchical clustering (Euclidean distances) of CAZyme and mcSEED metabolic pathway profiles was used to group participants with similar responses to each fibre snack type. CAZymes marked with a + were also fibre-treatment discriminatory in the gnotobiotic mouse studies.
Extended Data Fig. 7 Spearman-rank cross-correlation analysis of the representation of CAZymes, monosaccharides and glycosidic linkages in the faecal communities of participants consuming the pea-fibre snack prototype.
a, b, Correlations between the log2-transformed fold change of statistically significant, HOSVD-defined discriminatory CAZyme gene abundances (matched by time and participant) to the log2-transformed fold change in levels of monosaccharides and glycosidic linkages at days 25 and 35 (fibre snack consumption), and days 45 and 49 (post-intervention phase) normalized to day 14 (pre-intervention phase; n = 60 faecal samples analysed; 12 participants). Green boxes in a highlight a statistically significant positive correlation between GH43_37 (arabinofuranosidase) and arabinose, a prominent monosaccharide component of pea fibre. In b, evidence is provided that participant microbiomes contain CAZymes that cleave multiple branches of pea fibre arabinan, resulting in accumulation of its 1,5-arabinofuranose backbone in faeces. Further details are in Supplementary Results. *P < 0.05; **P < 0.01. Glucose (Glc), galacturonic acid (GalA), arabinose (Ara), xylose (Xyl), galactose (Gal), mannose (Man), rhamnose (Rha), fucose (Fuc), fructose (Fru), glucuronic acid (GlcA), N-acetylglucosamine (GlcNAc), N-acetylgalactosamine (GalNAc), allose (All), ribose (Rib), hexose (Hex), deoxyhexose (dHex); terminal (T), pyranose (p), furanose (f), undefined linkage (X).
Extended Data Fig. 8 Schematic of the analytic pipeline for identifying associations between changes in the plasma proteome and CAZyme responses after fibre snack consumption.
Step 1 shows cross-correlation analysis between plasma proteins and discriminatory CAZymes with changes in abundance that were statistically significant. Step 2 shows SVD analysis of Spearman’s rho values of the cross-correlation matrix. Proteins with projections along SV1 are plotted in a histogram to identify those proteins most correlated with discriminatory CAZymes (those within the 10th and 90th percentile, α < 0.1). Step 3 represents a CompBio-based analysis of groups of proteins with SV1 projections within the 10th and 90th percentiles. Biological themes enriched in proteins binned in the 10th and 90th percentiles are generated (threshold cutoff for enrichment score (log2) > 14.8). Step 4 is a SVD analysis of protein profiles within each biological theme for all participants, followed by cross-correlation between SVD projections (SV1) of themed proteins and discriminatory CAZyme responses to treatment.
Extended Data Fig. 9 CAZyme-associated plasma proteome responses to consumption of the four-fibre snack prototype.
a–c, Contextual language processing literature analysis (CompBio) of proteins with abundances that were significantly correlated with treatment-discriminatory CAZymes in participants consuming the four-fibre snack. The analysis procedure is summarized in Extended Data Fig. 8. Treatment-responsive proteins, identified by CC-SVD as having projections at the extremes of SV1 (10th and 90th percentiles of the distribution), are grouped into biological themes identified by CompBio, on the basis of a conditional probability analysis, as being significantly enriched for contextually associated biological concepts (processes or pathways) over those that occur by random sampling of the literature. Themes with enrichment scores (log2-transformed) > 14.8 in the plasma proteomes of participants who consumed the four-fibre snack are shown in b, c (Supplementary Table 10d–f provides a comprehensive list of themes associated with this and the other fibre snacks). d, e, Biological themes based on proteins positioned in 10th and 90th percentiles (d and e, respectively) are portrayed as spheres. The size of a sphere is related to its enrichment score in the plasma proteome after consumption of the four-fibre snack. The thickness of the blue lines connecting themes signifies the number of proteins shared between them. Component proteins of exemplary themes (orange spheres) are listed in boxes and coloured by their median log2-transformed fold change in response to consumption of the snack (blue, decrease; red, increase).
Extended Data Fig. 10 Connecting host responses defined by plasma proteomic features to microbiome responses defined by CAZyme features in participants consuming the four-fibre snack.
CC-SVD analysis of the plasma proteome with significantly changed discriminatory CAZymes in all participants after consuming the four-fibre snack. Two distinct groups of proteins with significant correlations with CAZymes are shown; one group with SV1 projections situated in the 10th percentile (top) and the other in the 90th percentile (bottom). CompBio analysis revealed biological themes that were significantly correlated with these CAZymes. Themes, their enrichment scores (log2-transformed), the number of proteins comprising each theme and the cross-correlation (Spearman’s rho) values between SV1 projections of themes and discriminatory CAZymes are presented. Each circle represents the correlation between a biological theme and a fibre-responsive CAZyme, with larger and darker circles indicating stronger correlations (positive correlations are coloured in red; negative correlations are coloured in blue).
Extended Data Fig. 11 Individual responses of the plasma proteome of participants consuming the four-fibre snack prototype.
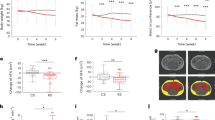
a, b, Heat maps plotting the projections on SV1 of changes in the representation of biological themes during consumption of the four-fibre snack. c, Heat map plotting the log2-transformed fold change in the levels of plasma proteins enriched in the glucose metabolism theme. Data for each participant are shown, normalized to the last day of the pretreatment phase on day 11. The four-fibre snack produced the greatest reduction in HOMA-IR among the three different snacks tested (Supplementary Table 5c). However, this reduction did not achieve statistical significance (P = 0.078, linear mixed-effects model) after the short period of snack consumption in this study. d–f, LC-QTOF-MS analysis of a biomarker of orange fibre consumption present in faecal samples obtained from gnotobiotic mice and humans. d, Comparison of levels of the m/z 274.1442 analyte in colonized and germ-free mice fed the unsupplemented, orange-fibre-supplemented or pea-fibre-supplemented HiSF–LoFV diet for 10 days. The analyte is detectable only when orange fibre is consumed and is not dependent upon on the human donor microbiome for its generation. Bars represent mean values ± s.d. for biological replicates (n = 5 mice per group). d, e, Comparisons of levels of the analyte in faecal samples obtained from participants in human study 2 on days 25 and 49 when they were consuming the maximum dose of the two-fibre (pea and inulin) and four-fibre (pea fibre, inulin, orange fibre plus barley bran) snack food prototypes. The bar graph in e represents mean values ± s.d. for technical replicates (n = 14 participants). The difference documented between participants consuming the 2- versus 4-fibre snacks is statistically significant (P = 0.0004, paired two-tailed t-test). The horizontal dashed line in f denotes a baseline value operationally defined as the highest level of detection of the analyte in participants consuming the two-fibre snack food prototype lacking orange fibre.
Extended Data Fig. 12 Plasma proteins with statistically significant changes in their abundances as a function of fibre treatment type and participant.
Heat map plotting the log2-transformed fold change in the abundances of plasma proteins in participants consuming the indicated fibre snack prototype. Data from the nine participants in study 1 (pea-fibre snack) who were also enrolled in study 2 (two- and four-fibre snacks) are shown. Changes in protein levels are referenced to their abundances on the last day of the pretreatment phase (day 14 and day 11 in study 1 and study 2, respectively) (n = 66 blood plasma samples analysed). *P < 0.05; **P < 0.01; ***P < 0.001 (linear model, limma25).
Supplementary information
Supplementary Information
This file contains the Supplementary Results and approved Study Protocols for NCT04159259 and NCT04101344.
Supplementary Tables
This file contains Supplementary Tables 1-10.
Rights and permissions
About this article
Cite this article
Delannoy-Bruno, O., Desai, C., Raman, A.S. et al. Evaluating microbiome-directed fibre snacks in gnotobiotic mice and humans. Nature 595, 91–95 (2021). https://doi.org/10.1038/s41586-021-03671-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03671-4
This article is cited by
-
Effects of dietary fibre on metabolic health and obesity
Nature Reviews Gastroenterology & Hepatology (2024)
-
Bioactive glycans in a microbiome-directed food for children with malnutrition
Nature (2024)
-
Resistant starch intake facilitates weight loss in humans by reshaping the gut microbiota
Nature Metabolism (2024)
-
Causal relationship between gut microbiota and cancers: a two-sample Mendelian randomisation study
BMC Medicine (2023)
-
Distinct gene expression signatures comparing latent tuberculosis infection with different routes of Bacillus Calmette-Guérin vaccination
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.