Abstract
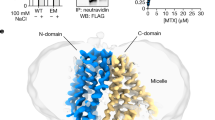
Folates (also known as vitamin B9) have a critical role in cellular metabolism as the starting point in the synthesis of nucleic acids, amino acids and the universal methylating agent S-adenylsmethionine1,2. Folate deficiency is associated with a number of developmental, immune and neurological disorders3,4,5. Mammals cannot synthesize folates de novo; several systems have therefore evolved to take up folates from the diet and distribute them within the body3,6. The proton-coupled folate transporter (PCFT) (also known as SLC46A1) mediates folate uptake across the intestinal brush border membrane and the choroid plexus4,7, and is an important route for the delivery of antifolate drugs in cancer chemotherapy8,9,10. How PCFT recognizes folates or antifolate agents is currently unclear. Here we present cryo-electron microscopy structures of PCFT in a substrate-free state and in complex with a new-generation antifolate drug (pemetrexed). Our results provide a structural basis for understanding antifolate recognition and provide insights into the pH-regulated mechanism of folate transport mediated by PCFT.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The plasmid encoding the chicken PCFT transporter cloned into the pDDGFP-LEU2d expression vector is available from Addgene (165414). The plasmid encoding the nanobody used in this study is available from Addgene (165415). Coordinates for the structures have been deposited in the Protein Data Bank under accession codes PDB 7BC6 (PCFT–nanobody) and 7BC7 (PCFT–nanobody + pemetrexed). The electron microscopy volumes have been deposited in the Electron Microscopy Data Bank under accession codes EMD-12140 (PCFT–nanobody) and EMD-12141 (PCFT–nanobody + pemetrexed). Any other relevant data are available from the corresponding authors upon reasonable request.
References
Zheng, Y. & Cantley, L. C. Toward a better understanding of folate metabolism in health and disease. J. Exp. Med. 216, 253–266 (2019).
Ducker, G. S. & Rabinowitz, J. D. One-carbon metabolism in health and disease. Cell Metab. 25, 27–42 (2017).
Zhao, R., Matherly, L. H. & Goldman, I. D. Membrane transporters and folate homeostasis: intestinal absorption and transport into systemic compartments and tissues. Expert Rev. Mol. Med. 11, e4 (2009).
Zhao, R., Aluri, S. & Goldman, I. D. The proton-coupled folate transporter (PCFT-SLC46A1) and the syndrome of systemic and cerebral folate deficiency of infancy: Hereditary folate malabsorption. Mol. Aspects Med. 53, 57–72 (2017).
Lucock, M. Folic acid: nutritional biochemistry, molecular biology, and role in disease processes. Mol. Genet. Metab. 71, 121–138 (2000).
Zhao, R. & Goldman, I. D. Folate and thiamine transporters mediated by facilitative carriers (SLC19A1-3 and SLC46A1) and folate receptors. Mol. Aspects Med. 34, 373–385 (2013).
Qiu, A. et al. Identification of an intestinal folate transporter and the molecular basis for hereditary folate malabsorption. Cell 127, 917–928 (2006).
Goldman, I. D., Chattopadhyay, S., Zhao, R. & Moran, R. The antifolates: evolution, new agents in the clinic, and how targeting delivery via specific membrane transporters is driving the development of a next generation of folate analogs. Curr. Opin. Investig. Drugs 11, 1409–1423 (2010).
Matherly, L. H., Hou, Z. & Gangjee, A. The promise and challenges of exploiting the proton-coupled folate transporter for selective therapeutic targeting of cancer. Cancer Chemother. Pharmacol. 81, 1–15 (2018).
Zhao, R. et al. The proton-coupled folate transporter: impact on pemetrexed transport and on antifolates activities compared with the reduced folate carrier. Mol. Pharmacol. 74, 854–862 (2008).
Bailey, L. B. Folate in Health and Disease, 2nd edn (Taylor & Francis, 2010).
Crider, K. S., Bailey, L. B. & Berry, R. J. Folic acid food fortification-its history, effect, concerns, and future directions. Nutrients 3, 370–384 (2011).
Visentin, M., Diop-Bove, N., Zhao, R. & Goldman, I. D. The intestinal absorption of folates. Annu. Rev. Physiol. 76, 251–274 (2014).
Kamen, B. A. & Smith, A. K. A review of folate receptor alpha cycling and 5-methyltetrahydrofolate accumulation with an emphasis on cell models in vitro. Adv. Drug Deliv. Rev. 56, 1085–1097 (2004).
Zhao, R. et al. A role for the proton-coupled folate transporter (PCFT-SLC46A1) in folate receptor-mediated endocytosis. J. Biol. Chem. 284, 4267–4274 (2009).
Matherly, L. H., Wilson, M. R. & Hou, Z. The major facilitative folate transporters solute carrier 19A1 and solute carrier 46A1: biology and role in antifolate chemotherapy of cancer. Drug Metab. Dispos. 42, 632–649 (2014).
Qiu, A. et al. Rodent intestinal folate transporters (SLC46A1): secondary structure, functional properties, and response to dietary folate restriction. Am. J. Physiol. Cell Physiol. 293, C1669–C1678 (2007).
Kronn, D. & Goldman, I. D. in GeneReviews (eds Adam, M. P. et al.) https://www.ncbi.nlm.nih.gov/books/NBK1673/ (1993).
Furst, D. E. The rational use of methotrexate in rheumatoid arthritis and other rheumatic diseases. Br. J. Rheumatol. 36, 1196–1204 (1997).
Giovannetti, E. et al. Role of proton-coupled folate transporter in pemetrexed resistance of mesothelioma: clinical evidence and new pharmacological tools. Ann. Oncol. 28, 2725–2732 (2017).
Chattopadhyay, S., Moran, R. G. & Goldman, I. D. Pemetrexed: biochemical and cellular pharmacology, mechanisms, and clinical applications. Mol. Cancer Ther. 6, 404–417 (2007).
Pascale, R. M., Calvisi, D. F., Simile, M. M., Feo, C. F. & Feo, F. The Warburg effect 97 years after its discovery. Cancers 12, E2819 (2020).
Desmoulin, S. K., Hou, Z., Gangjee, A. & Matherly, L. H. The human proton-coupled folate transporter: biology and therapeutic applications to cancer. Cancer Biol. Ther. 13, 1355–1373 (2012).
Drew, D. & Boudker, O. Shared molecular mechanisms of membrane transporters. Annu. Rev. Biochem. 85, 543–572 (2016).
Huang, Y., Lemieux, M. J., Song, J., Auer, M. & Wang, D.-N. Structure and mechanism of the glycerol-3-phosphate transporter from Escherichia coli. Science 301, 616–620 (2003).
Shin, D. S., Zhao, R., Fiser, A. & Goldman, I. D. Role of the fourth transmembrane domain in proton-coupled folate transporter function as assessed by the substituted cysteine accessibility method. Am. J. Physiol. Cell Physiol. 304, C1159–C1167 (2013).
Chen, L. Q. & Pagel, M. D. Evaluating pH in the extracellular tumor microenvironment using CEST MRI and other imaging methods. Adv. Radiol. 2015, 206405 (2015).
Mahadeo, K. et al. Properties of the Arg376 residue of the proton-coupled folate transporter (PCFT–SLC46A1) and a glutamine mutant causing hereditary folate malabsorption. Am, J. Physiol. Cell Physiol. 299, C1153–C1161 (2010).
Shin, D. S. et al. Functional roles of aspartate residues of the proton-coupled folate transporter (PCFT-SLC46A1); a D156Y mutation causing hereditary folate malabsorption. Blood 116, 5162–5169 (2010).
Unal, E. S., Zhao, R. & Goldman, I. D. Role of the glutamate 185 residue in proton translocation mediated by the proton-coupled folate transporter SLC46A1. Am. J. Physiol. Cell Physiol. 297, C66–C74 (2009).
Unal, E. S. et al. The functional roles of the His247 and His281 residues in folate and proton translocation mediated by the human proton-coupled folate transporter SLC46A1. J. Biol. Chem. 284, 17846–17857 (2009).
Desmoulin, S. K. et al. Targeting the proton-coupled folate transporter for selective delivery of 6-substituted pyrrolo[2,3-d]pyrimidine antifolate inhibitors of de novo purine biosynthesis in the chemotherapy of solid tumors. Mol. Pharmacol. 78, 577–587 (2010).
Wang, L. et al. Synthesis and antitumor activity of a novel series of 6-substituted pyrrolo[2,3-d]pyrimidine thienoyl antifolate inhibitors of purine biosynthesis with selectivity for high affinity folate receptors and the proton-coupled folate transporter over the reduced folate carrier for cellular entry. J. Med. Chem. 53, 1306–1318 (2010).
Parker, J. L. & Newstead, S. Method to increase the yield of eukaryotic membrane protein expression in Saccharomyces cerevisiae for structural and functional studies. Protein Sci. 23, 1309–1314 (2014).
Diop-Bove, N. K., Wu, J., Zhao, R., Locker, J. & Goldman, I. D. Hypermethylation of the human proton-coupled folate transporter (SLC46A1) minimal transcriptional regulatory region in an antifolate-resistant HeLa cell line. Mol. Cancer Ther. 8, 2424–2431 (2009).
Zhao, R., Gao, F., Hanscom, M. & Goldman, I. D. A prominent low-pH methotrexate transport activity in human solid tumors: contribution to the preservation of methotrexate pharmacologic activity in HeLa cells lacking the reduced folate carrier. Clin. Cancer Res. 10, 718–727 (2004).
Huo, J. et al. Neutralizing nanobodies bind SARS-CoV-2 spike RBD and block interaction with ACE2. Nat. Struct. Mol. Biol. 27, 846–854 (2020).
Pardon, E. et al. A general protocol for the generation of nanobodies for structural biology. Nat. Protocols 9, 674–693 (2014).
Caesar, J. et al. SIMPLE 3.0. Stream single-particle cryo-EM analysis in real time. J. Struct. Biol. X 4, 100040 (2020).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Asarnow, D., Palovcak, E. & Cheng, Y. UCSF pyem v.0.5., https://doi.org/10.5281/zenodo.3576630 (2019).
Tan, Y. Z. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat. Methods 14, 793–796 (2017).
Brown, A. et al. Tools for macromolecular model building and refinement into electron cryo-microscopy reconstructions. Acta Crystallogr. D 71, 136–153 (2015).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Emsley, P. Tools for ligand validation in Coot. Acta Crystallogr. D 73, 203–210 (2017).
Moriarty, N. W., Grosse-Kunstleve, R. W. & Adams, P. D. Electronic Ligand Builder and Optimization Workbench (eLBOW): a tool for ligand coordinate and restraint generation. Acta Crystallogr. D 65, 1074–1080 (2009).
Prisant, M. G., Williams, C. J., Chen, V. B., Richardson, J. S. & Richardson, D. C. New tools in MolProbity validation: CaBLAM for cryoEM backbone, UnDowser to rethink “waters,” and NGL viewer to recapture online 3D graphics. Protein Sci. 29, 315–329 (2020).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Marrink, S. J., Risselada, H. J., Yefimov, S., Tieleman, D. P. & de Vries, A. H. The MARTINI force field: coarse grained model for biomolecular simulations. J. Phys. Chem. B 111, 7812–7824 (2007).
Wu, Z., Alibay, I., Newstead, S. & Biggin, P. C. Proton control of transitions in an amino acid transporter. Biophys. J. 117, 1342–1351 (2019).
Vickery, O. & Corey, R. owenvickery/cg2at: CG2AT2 a fragment based conversion version 0.2, https://doi.org/10.5281/zenodo.3994618 (2020).
Maier, J. A. et al. ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Jämbeck, J. P. M. & Lyubartsev, A. P. An extension and further validation of an all-atomistic force field for biological membranes. J. Chem. Theory Comput. 8, 2938–2948 (2012).
Petrova, J. et al. Molecular simulation of the structure of folate and antifolates at physiological conditions. J. Mol. Graph. Model. 87, 172–184 (2019).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Bayly, C. I., Cieplak, P., Cornell, W. & Kollman, P. A. A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the RESP model. J. Phys. Chem. 97, 10269–10280 (1993).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Berendsen, H. J., Postma, J. P. M., van Gunsteren, W. F., DiNola, A. & Haak, J. R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Gapsys, V., Michielssens, S., Seeliger, D. & de Groot, B. L. pmx: automated protein structure and topology generation for alchemical perturbations. J. Comput. Chem. 36, 348–354 (2015).
Goga, N., Rzepiela, A. J., de Vries, A. H., Marrink, S. J. & Berendsen, H. J. Efficient algorithms for Langevin and DPD dynamics. J. Chem. Theory Comput. 8, 3637–3649 (2012).
Klimovich, P. V., Shirts, M. R. & Mobley, D. L. Guidelines for the analysis of free energy calculations. J. Comput. Aided Mol. Des. 29, 397–411 (2015).
Wu, Z. Alchemicalitp: a Gromacs parser for alchemical transformation version 0.1, https://github.com/xiki-tempula/alchemicalitp (2020).
The PLUMED consortium. Promoting transparency and reproducibility in enhanced molecular simulations. Nat. Methods 16, 670–673 (2019).
Grossfield, A. WHAM: the weighted histogram analysis method version 2.0.10.2., http://membrane.urmc.rochester.edu/wordpress/?page_id=126 (accessed 2020).
Acknowledgements
The Central Oxford Structural Microscopy and Imaging Centre is support by the Wellcome Trust (201536), the EPA Cephalosporin Trust and a Royal Society/Wolfson Foundation Laboratory Refurbishment Grant (WL160052). Computing was supported via the Advanced Research Computing facility (Oxford), the ARCHER UK National Supercomputing Service and JADE (EP/P020275/1) granted via the High-End Computing Consortium for Biomolecular Simulation, (HECBioSim (http://www.hecbiosim.ac.uk)), supported by EPSRC (EP/L000253/1). This research was supported by Wellcome awards to S.M.L. (209194;100298), P.C.B. (219531) and S.N. (215519;219531) and through MRC grants to S.M.L. (MR/M011984/1) and J.L.P. (MR/S021043/1). Z.W. is a Wellcome Trust PhD student (203741). Figures were created using BioRender.com, PyMol and ChimeraX.
Author information
Authors and Affiliations
Contributions
J.L.P. and S.N. conceived the project. G.K. maintained cell stocks, and undertook large-scale expression and tissue culture. J.L.P. and S.N. performed all protein preparation. J.H. and R.J.O. screened the Abcore library. J.C.D. and S.M.L. performed all cryo-EM sample processing, data collection and image analysis. J.C.D., S.M.L. and S.N. constructed the atomic models. I.D.G. contributed the HeLa-derived cell line HeLa R1-11, and provided insights into human mutations and physiology. J.L.P. conducted all transport and biochemical assays. Z.W. and P.C.B. performed all molecular dynamics analysis. J.L.P. and S.N. wrote the manuscript and prepared figures, with contributions and discussions from Z.W., I.D.G., P.C.B., J.C.D. and S.M.L. G.K. and Z.W. contributed equally to this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Larry H. Matherley, D. J. Slotboom and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Comparison of human and chicken PCFT homologues.
a, Sequence alignment of PCFT from human (Hs) (Uniprot Q96NT5) and chicken (Gg) (E6Y8U5) coloured via amino acid chemistry. Human and chicken PCFT homologues share an overall 58% identity and 87% similarity. Functionally relevant residues are highlighted with yellow stars, and mutations found to cause hereditary folate malabsorption disorder are in red. b, Cell-based uptake assay comparing the transport of 3H folic acid via human and chicken PCFT at both pH 5.5 and 7.5. The human and chicken homologues transport similar amounts of folic acid to each other and do so only at acidic pH (5.5). n = 4 independent experiments, mean and s.d. are shown. c, Pemetrexed competition of 3H folic acid uptake into cells, overexpressing either human or chicken PCFT. The calculated mean (from 4 independent experiments) half-maximal inhibitory concentration values are indicated ± s.d. d, Calculated KM for folic acid uptake at pH 5.5 by chicken PCFT in a liposome-based uptake assay. n = 3 independent experiments, calculated mean and s.d. values are shown e, Effect of pH on PCFT uptake in a liposome-based assay. n = 3 independent experiments, with the mean and s.d. shown. f, Membrane potential induced through potassium diffusion gradient (plus valinomycin) does not affect transport, an observation that has also been seen for human PCFT in cells34, suggesting that transport is thermodynamically coupled to two protons (assuming a −2 charge on folic acid). n = 4 independent experiments, data are mean and s.d.
Extended Data Fig. 2 Characterization of high-affinity nanobody to PCFT.
a, The nanobody identified from a naive llama library has a KD of about 8 nM for PCFT binding. n = 3 independent experiments, calculated mean ± s.d. shown. b, The nanobody blocks uptake of folic acid into liposomes containing PCFT, whereas a non-specific nanobody has no effect. n = 3 independent experiments, mean and s.d. shown. c, SDS–PAGE analysis of the PCFT–nanobody complex after size exclusion. Experiment was performed four times with similar results. d, Representative gel filtration trace of chicken PCFT in DDM:CHS detergent at pH 6.5. The protein elutes as a monomer of about 50 kDa. Insets show Coomassie-stained SDS–PAGE gel of the purified PCFT protein and circular dichroism analysis. Experiment was repeated eight times with similar results. e, Analysis of the thermostability of PCFT under different pH conditions (7.5 and 5.5) indicates that acidic pH stabilizes the protein. The presence of both folic acid (FA) or pemetrexed (PMX) further stabilize the protein, but only at acidic pH. Experiments were repeated three times with similar results.
Extended Data Fig. 3 Cryo-EM processing workflow, showing local and global map quality for the PCFT–nanobody complex.
a, Image processing workflow for PCFT–nanobody. b, Local-resolution estimation of reconstructed map as determined within RELION. Detergent density omitted for clarity. c, Gold-standard FSC curves used for global-resolution estimates within (i) cryoSPARC, (ii) RELION, or (iii) 3DFSC. d, Close-up view of map and side-chain density for transmembrane helices and lateral helix. Volume contoured at threshold level of 0.805.
Extended Data Fig. 4 Analysis of PCFT–pemetrexed complex.
a, Cryo-EM density of PCFT in complex with pemetrexed and nanobody. b, In molecular dynamics simulations of the apo structure (grey), the TM1–TM2 loop is positioned such that the entrance to the binding pocket is accessible. In simulations with pemetrexed bound (orange), the loop closes quickly (within 250 ns) as measured by the relative position of Ser68 on the TM1–TM2 loop and Ala308 on the TM7–TM8 loop. c, Time course of the distance between the Cα atoms of Ser68 and Ala308 for both apo (grey) and pemetrexed-bound (orange) simulations. The closure event observed with pemetrexed bound occurs around 250 ns, and this closed conformation remains stable for the remainder of the simulation. On removal of pemetrexed from the end of the simulation shown in this panel (orange), the TM1–TM2 moves away from the TM7–TM8 loop (blue). The movement is shown in b (cyan). d, The presence of 2 mM DTT does not alter the uptake of folic acid into liposomes by PCFT, consistent with previous studies on the human transporter35. e, The presence of 2 mM DTT does lead to a destabilization of the protein as determined by differential scanning fluorimetry. f, View of pemetrexed within the binding cavity, with surface charge highlighted. g, Schematic of the binding pose observed for pemetrexed. Hydrogen-bond donors and acceptors are highlighted by directional arrows, the sole charge–charge interaction by a solid line and the π–π interaction in orange. Values indicate distance in Å.
Extended Data Fig. 5 Structural relationship between the observed water pocket and the gating helices in PCFT.
a, Cartoon of PCFT, showing the open and closed states of the extracellular and intracellular gates, respectively. b, Molecular dynamic simulations demonstrate the water pocket is both stable and accessible to bulk solvent via the substrate-binding cavity. The space occupied by the polar pocket results in fewer interactions between TM7 and TM10, which is likely to facilitate the movement of the helices against one another during transport.
Extended Data Fig. 6 Cryo-EM processing workflow, showing local and global map quality for the PCFT–nanobody complex bound to pemetrexed.
a, Image processing workflow for pemetrexed-bound PCFT–nanobody. b, Local-resolution estimation of reconstructed map as determined within RELION. Detergent density omitted for clarity. Top, full map; bottom, central slab through map. c, Gold-standard FSC curves used for global resolution estimates within (i) cryoSPARC, (ii) RELION or (iii) 3DFSC. d, Close-up of side-chain density for all transmembrane helices. Volume contoured at threshold level of 0.3. e, Density for pemetrexed (PMX) and side chains surrounding PCFT. Volume contoured at threshold level of 0.4.
Extended Data Fig. 7 The effect of Glu407Asn on apo and pemetrexed-bound PCFT and hereditary folate malabsorption mutations in the context of the chicken PCFT structure.
a–e, Alchemical transformations show that a Glu407Asn mutation would stabilize the protein by forming hydrogen bonds with Asn166, while remaining capable of preserving the interaction with the pyrrole amine of pemetrexed. a, In the wild-type apo state Glu407 does not interact with Asn166. The Glu407Asn variant, however, can readily hydrogen-bond with Asn166 (b), resulting in the Glu407Asn variant being 1.5 kcal mol−1 more stable (e) (blue). In the pemetrexed-bound state, Glu407 makes a hydrogen bond with the pyrrole amine group of pemetrexed (PMX) (c), which is also preserved in the Glu407Asn variant (d). The coordinated hydrogen-bond network among Asn166, Glu407Asn and pemetrexed further stabilizes the protein-bound state by 2 kcal mol−1 compared to the apo state (e) (orange). n = 3 independent repeats with mean and s.d. plotted. f, Cartoon of chicken PCFT structure with residues involved in hereditary folate malabsorption shown as spheres. Asp164 (Asp156) and Arg384 (Arg376) are highlighted as these residues have an important role in the transport mechanism. The table shows the corresponding human residue number and the associated phenotype. PubMed identifiers for the respective studies that describe mutations associated with hereditary folate malabsorption are included.
Extended Data Fig. 8 Analysis of residues involved in proton binding in PCFT.
a, Analysis of wild type and variants of chicken PCFT studied in liposome assays. SDS–PAGE gel showing pre- and post-solubilization of proteoliposomes with 1% DDM:CHS. A proportion of purified PCFT is not folded correctly in the liposome and runs as aggregates on the pre-solubilized samples in the SDS–PAGE gel. Experiments were repeated three times with similar results. b, Protonation of D164 results in easier breakage of the salt bridge interaction to R156. There is a free energy barrier of 2 kcal mol−1 associated with the breakage of the R156–D164 salt bridge in the apo state (i and ii). The protonation of the D164 (DH164) would lower this free energy barrier to a level lower than 0.5 kcal mol−1 (iii and iv) and thus bring it under the level of thermal fluctuation. The presence of pemetrexed does not affect this process. Rather, it controls the likelihood that D164 is protonated in the first place and thus effects the salt bridge stability indirectly. c, Convergence of the PMF profile shown in b. The PMF profile is computed for the first 20% (blue), 40% (orange), 60% (green), 80% (red) and 100% (purple) of the data. The columns are three repeats of the calculations. d, Convergence plots of the protonation free energy. The forward plot (orange lines) is the protonation free energy computed from the first 10% up to 100% of data, in 10% increments. The backward plot (blue lines) is the protonation free energy computed from the final 10% up to 100% of the data, in 10% increments. The columns are three repeats of the calculations. The green line is the final estimate of the free energy for each simulation and the width shows the uncertainty in the multistate Bennet acceptance ratio calculation. e, Table showing calculated pKa and free energy values (mean ± s.d.). f, Western blot analysis of the cell-based assay using an anti-Flag antibody for the PCFT variants and a loading control of anti-β-actin. Experiments were repeated twice with similar results.
Extended Data Fig. 9 Model for ligand-induced closure of the extracellular gate.
a, Structure of PCFT highlighting the extracellular gate helices, TM7 and TM8 (red) and their relationship to the bound pemetrexed molecule (yellow). The arrow indicates the movement required to seal the binding site from the extracellular side of the membrane. The interaction of Phe290 with the benzyl group of pemetrexed is likely to have an important role in triggering gate closure. b, Structural comparison between the apo and pemetrexed-bound states reveals repositioning of His289, resulting in the breakage of its interaction with Asn350 and facilitating the movement of TM7. The water pocket substantially enlarges in the pemetrexed-bound state, consistent with greater flexibility in the C-terminal bundle under acidic conditions.
Supplementary information
Supplementary Figure
Original SDS PAGE gels used to generate Extended Data Figures 2c, 2d and 8a.
Rights and permissions
About this article
Cite this article
Parker, J.L., Deme, J.C., Kuteyi, G. et al. Structural basis of antifolate recognition and transport by PCFT. Nature 595, 130–134 (2021). https://doi.org/10.1038/s41586-021-03579-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03579-z
This article is cited by
-
Ion and lipid orchestration of secondary active transport
Nature (2024)
-
Future opportunities in solute carrier structural biology
Nature Structural & Molecular Biology (2024)
-
Structural basis of promiscuous substrate transport by Organic Cation Transporter 1
Nature Communications (2023)
-
cGAMP-activated cGAS–STING signaling: its bacterial origins and evolutionary adaptation by metazoans
Nature Structural & Molecular Biology (2023)
-
Molecular basis for substrate recognition and transport of human GABA transporter GAT1
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.