Abstract
The initiation of cell division integrates a large number of intra- and extracellular inputs. D-type cyclins (hereafter, cyclin D) couple these inputs to the initiation of DNA replication1. Increased levels of cyclin D promote cell division by activating cyclin-dependent kinases 4 and 6 (hereafter, CDK4/6), which in turn phosphorylate and inactivate the retinoblastoma tumour suppressor. Accordingly, increased levels and activity of cyclin D–CDK4/6 complexes are strongly linked to unchecked cell proliferation and cancer2,3. However, the mechanisms that regulate levels of cyclin D are incompletely understood4,5. Here we show that autophagy and beclin 1 regulator 1 (AMBRA1) is the main regulator of the degradation of cyclin D. We identified AMBRA1 in a genome-wide screen to investigate the genetic basis of the response to CDK4/6 inhibition. Loss of AMBRA1 results in high levels of cyclin D in cells and in mice, which promotes proliferation and decreases sensitivity to CDK4/6 inhibition. Mechanistically, AMBRA1 mediates ubiquitylation and proteasomal degradation of cyclin D as a substrate receptor for the cullin 4 E3 ligase complex. Loss of AMBRA1 enhances the growth of lung adenocarcinoma in a mouse model, and low levels of AMBRA1 correlate with worse survival in patients with lung adenocarcinoma. Thus, AMBRA1 regulates cellular levels of cyclin D, and contributes to cancer development and the response of cancer cells to CDK4/6 inhibitors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequencing data from Tuba-seq experiments and RNA-sequencing data from AMBRA1-knockout U2OS cells are available from the Gene Expression Omnibus under accession numbers GSE146303 and GSE159920, respectively. Mass spectrometry data from shotgun proteomics experiments and analysis of ubiquitylated proteins are available through the ProteomeXchange Consortium, with dataset identifiers PXD021789 and PXD022111, respectively. Public nonprotected RNA-sequencing, copy number alteration, exome sequencing and reverse-phase protein array lung adenocarcinoma datasets from the TCGA were downloaded from https://gdc.cancer.gov/. Clinical data were obtained from a previous publication36 (PMID: 29625055). Gene dependency data from the Cancer Dependency Map are publicly available at www.depmap.org. Protein sequences for mass spectrometry analysis were obtained from the NCBI Homo sapiens protein database (ftp://ftp.ncbi.nlm.nih.gov/refseq/release/release-notes/archive/RefSeq-release86.txt, downloaded 05/11/2018) (shotgun mass spectrometry) and from Uniprot (https://www.uniprot.org/uniprot/?query=proteome:UP000005640%20reviewed:yes, downloaded 02/28/2020) (ubiquitin remnant profiling). All other data are available in the Article and Supplementary Information, or from the corresponding author upon reasonable request. Source data are provided with this paper.
References
Deshpande, A., Sicinski, P. & Hinds, P. W. Cyclins and cdks in development and cancer: a perspective. Oncogene 24, 2909–2915 (2005).
Sherr, C. J. Cancer cell cycles. Science 274, 1672–1677 (1996).
Malumbres, M. & Barbacid, M. Cell cycle, CDKs and cancer: a changing paradigm. Nat. Rev. Cancer 9, 153–166 (2009).
Kanie, T. et al. Genetic reevaluation of the role of F-box proteins in cyclin D1 degradation. Mol. Cell. Biol. 32, 590–605 (2012).
Qie, S. & Diehl, J. A. Cyclin D degradation by E3 ligases in cancer progression and treatment. Semin. Cancer Biol. 67, 159–170 (2020).
Sherr, C. J., Beach, D. & Shapiro, G. I. Targeting CDK4 and CDK6: from discovery to therapy. Cancer Discov. 6, 353–367 (2016).
Walter, D. M. et al. RB constrains lineage fidelity and multiple stages of tumour progression and metastasis. Nature 569, 423–427 (2019).
Wander, S. A. et al. The Genomic landscape of intrinsic and acquired resistance to cyclin-dependent kinase 4/6 inhibitors in patients with hormone receptor-positive metastatic breast cancer. Cancer Discov. 10, 1174–1193 (2020).
Schoninger, S. F. & Blain, S. W. The ongoing search for biomarkers of CDK4/6 inhibitor responsiveness in breast cancer. Mol. Cancer Ther. 19, 3–12 (2020).
Fimia, G. M. et al. Ambra1 regulates autophagy and development of the nervous system. Nature 447, 1121–1125 (2007).
Antonioli, M. et al. AMBRA1 interplay with cullin E3 ubiquitin ligases regulates autophagy dynamics. Dev. Cell 31, 734–746 (2014).
Cianfanelli, V. et al. AMBRA1 links autophagy to cell proliferation and tumorigenesis by promoting c-Myc dephosphorylation and degradation. Nat. Cell Biol. 17, 706 (2015).
Montaudon, E. et al. PLK1 inhibition exhibits strong anti-tumoral activity in CCND1-driven breast cancer metastases with acquired palbociclib resistance. Nat. Commun. 11, 4053 (2020).
Alt, J. R., Cleveland, J. L., Hannink, M. & Diehl, J. A. Phosphorylation-dependent regulation of cyclin D1 nuclear export and cyclin D1-dependent cellular transformation. Genes Dev. 14, 3102–3114 (2000).
Lin, D. I. et al. Phosphorylation-dependent ubiquitination of cyclin D1 by the SCFFBX4-αB crystallin complex. Mol. Cell 24, 355–366 (2006).
Ewen, M. E. et al. Functional interactions of the retinoblastoma protein with mammalian D-type cyclins. Cell 73, 487–497 (1993).
Jahn, S. C. et al. Assembly, activation, and substrate specificity of cyclin D1/Cdk2 complexes. Biochemistry 52, 3489–3501 (2013).
Chytil, A. et al. Construction of a cyclin D1–Cdk2 fusion protein to model the biological functions of cyclin D1–Cdk2 complexes. J. Biol. Chem. 279, 47688–47698 (2004).
Jansen, V. M. et al. Kinome-wide RNA interference screen reveals a role for PDK1 in acquired resistance to CDK4/6 inhibition in ER-positive breast cancer. Cancer Res. 77, 2488–2499 (2017).
Herrera-Abreu, M. T. et al. Early adaptation and acquired resistance to CDK4/6 inhibition in estrogen receptor-positive breast cancer. Cancer Res. 76, 2301–2313 (2016).
James, M. K., Ray, A., Leznova, D. & Blain, S. W. Differential modification of p27Kip1 controls its cyclin D-cdk4 inhibitory activity. Mol. Cell. Biol. 28, 498–510 (2008).
Guiley, K. Z. et al. p27 allosterically activates cyclin-dependent kinase 4 and antagonizes palbociclib inhibition. Science 366, eaaw2106 (2019).
Pagano, M., Theodoras, A. M., Tam, S. W. & Draetta, G. F. Cyclin D1-mediated inhibition of repair and replicative DNA synthesis in human fibroblasts. Genes Dev. 8, 1627–1639 (1994).
Chen, S. H. et al. CRL4AMBRA1 targets elongin C for ubiquitination and degradation to modulate CRL5 signaling. EMBO J. 37, e97508 (2018).
Xia, P. et al. WASH inhibits autophagy through suppression of beclin 1 ubiquitination. EMBO J. 32, 2685–2696 (2013).
Rogers, Z. N. et al. Mapping the in vivo fitness landscape of lung adenocarcinoma tumor suppression in mice. Nat. Genet. 50, 483–486 (2018).
Simoneschi, D. & Pagano, D. CRL4AMBRA1 is a regulator of D-type cyclins. Nature (2021).
Maiani, E., Milletti, G., Bartek, J. & Cecconi, F. AMBRA1 regulates cyclin D to guard S-phase entry and genomic integrity. Nature (2021).
Santra, M. K., Wajapeyee, N. & Green, M. R. F-box protein FBXO31 mediates cyclin D1 degradation to induce G1 arrest after DNA damage. Nature 459, 722–725 (2009).
Burkhart, D. L. & Sage, J. Cellular mechanisms of tumour suppression by the retinoblastoma gene. Nat. Rev. Cancer 8, 671–682 (2008).
Finn, R. S. et al. PD 0332991, a selective cyclin D kinase 4/6 inhibitor, preferentially inhibits proliferation of luminal estrogen receptor-positive human breast cancer cell lines in vitro. Breast Cancer Res. 11, R77 (2009).
Gong, X. et al. Genomic aberrations that activate D-type cyclins are associated with enhanced sensitivity to the CDK4 and CDK6 inhibitor abemaciclib. Cancer Cell 32, 761–776.e6 (2017).
Finn, R. S. et al. Biomarker analyses of response to cyclin-dependent kinase 4/6 inhibition and endocrine therapy in women with treatment-naïve metastatic breast cancer. Clin. Cancer Res. 26, 110–121 (2020).
DeMichele, A. et al. CDK 4/6 inhibitor palbociclib (PD0332991) in Rb+ advanced breast cancer: phase II activity, safety, and predictive biomarker assessment. Clin. Cancer Res. 21, 995–1001 (2015).
Xue, Y. et al. CDK4/6 inhibitors target SMARCA4-determined cyclin D1 deficiency in hypercalcemic small cell carcinoma of the ovary. Nat. Commun. 10, 558 (2019).
Liu, J. et al. An integrated TCA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400–416 (2018).
Acknowledgements
We thank S. Rubin and J. Skotheim for critical reading of the manuscript; A. Koff, V. Dulic and K. Keyomarsi for helpful discussions; and all of the members of the laboratory of J.S., and especially G. Coles, for their help and support throughout this study. Research reported in this Article was supported by the NIH (J.S., R01CA228413 and 1R35CA231997; A.C.C., 1F99CA245471-01; E.E.J., 2T32CA009302; R.C., 5T32GM007276; M.M.W. and D.A.P., R01CA2344349; and N.J.K., P50AI150476 and U54CA209891), the California TRDRP (J.S., 28IR-0046) and the NSF (GRFP, to A.C.C.). E.E.J. and M.C.L. were supported by a Stanford Graduate Fellowship. S.L. was supported by a Boehringer Ingelheim Fonds MD Fellowship. C.L. is the Connie and Bob Lurie Fellow of the Damon Runyon Cancer Research Foundation (DRG-2331). J.S. is the Harriet and Mary Zelencik Scientist in Children’s Cancer and Blood Diseases and the Elaine and John Chambers Professor in Pediatric Cancer.
Author information
Authors and Affiliations
Contributions
A.C.C. and J.S. designed most of the experiments and interpreted the results. A.C.C. and E.E.J. performed and analysed the CRISPR–Cas9 screen under the supervision of M.C.B.; A.C.C., M.C.L. and C.W.M. performed the Tuba-seq experiments under the supervision of M.M.W.; C.L. performed the computational analysis of the Tuba-seq experiments under the supervision of M.M.W. and D.P.; A.C.C. and A.H. performed the xenograft experiments. Y.T.S., S.Q.H. and A.H. performed immunostaining; Y.T.S. dissected mouse embryos; C.K. performed the histopathological analysis. J.A.S. and P.S. performed the analysis of human lung cancer data under the supervision of C.C.; S.L., E.M. and C.P. performed experiments related to cell cycle phenotypes under the guidance of A.C.C.; A.P.D. analysed the RNA-sequencing data; A.Y. and J.A.D. helped to prepare and design the protein stability and ubiquitylation experiments; A.C.C, R.C., J.D. and P.K.J. performed and analysed the shotgun mass-spectrometry experiments; D.L.S., S.-H.C., B.W.N., J.R.J. and N.J.K. performed and analysed the ubiquitylation mass-spectrometry experiments; A.C.C. and J.S. wrote the manuscript, with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
J.S. has received research funding from Stemcentrx/Abbvie, Pfizer and Revolution Medicines. M.M.W. and D.P. have equity in, and are advisors for, D2GOncology. C.C. is a scientific advisor to GRAIL and reports stock options as well as consulting for GRAIL and Genentech. N.J.K. has received research support from Vir Biotechnology and F. Hoffmann-La Roche. The authors declare no other competing interests.
Additional information
Peer review information Nature thanks Marianne Bronner, Piotr Sicinski and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification of AMBRA1 and other factors involved in the response of cells to CDK4/6 inhibitors.
a, Proliferation of U937 cells in the presence of 0.5 μM palbociclib (palbo) over 6 d, determined by cell counting (n = 1 experiment). b, Immunoassay of total RB and RB phosphorylated at S807 and S811 (p-RB S807/811) in U937 cells over 36 h of palbociclib treatment. c, Quantification of phosphorylated RB relative to total RB from b (n = 1 experiment). d, Schematic of the CRISPR–Cas9 screen in U937 cells. e, Protein–protein interaction map of screen results, generated using Metascape. Coloured nodes represent densely connected gene neighbourhoods. Legend indicates the gene ontology term that is most significantly enriched within each neighbourhood. Node size indicates the degree of connectedness. Gene names can be found in Supplementary Table 3. f, Schematic of the screen results among RB-pathway genes expressed in U937 cells. g, Number of control and knockout U937 cells treated with 0.5 μM palbociclib or DMSO control for 48 h. Each symbol is an isogenic clone (n = 3 biological replicates per clone). h, Left, schematic of the competition assay between GFP-negative parental U937 cells and GFP-positive knockout cell populations. Right, example of flow cytometry analysis for one experiment with AMBRA1-knockout cells. i, Percentage of GFP-positive control or knockout populations in competition assays as in h (n = 3 biological replicates). j, Representative flow cytometry plots of annexin V and propidium iodide (PI) staining in U937 cells treated with 0.5 μM palbociclib for 24 h. k, Percentage of apoptotic (annexinV+PI+) U937 cells after a 24-h palbociclib treatment (n = 3 biological replicates per clone). Palbociclib does not induce apoptosis in any genotype. l, Representative flow cytometry plots of BrdU and PI staining in U937 cells treated with 0.5 μM palbociclib for 24 h. m, Percentage of S-phase cells by BrdU and PI staining in U937 cells treated with 1 μM abemaciclib for 24 h (n = 3 biological replicates per clone). n, Immunoassay for AMBRA1 and RB in control and knockout cancer cell lines generated by CRISPR–Cas9. For U2OS (osteosarcoma), NCI-H1792 (lung adenocarcinoma) and NCI-H460 (large cell lung cancer), each lane is an isogenic clone. MCF7 cells (breast cancer) are populations. o, Percentage of cycling S-phase cells from n after a 24-h treatment with palbociclib (0.5 μM for all cell lines except for MCF7 cells, 0.04 μM). U2OS, NCI-H1792 and NCI-H460 cells were analysed by BrdU and PI staining, and each symbol is an isogenic clone (n = 3 biological replicates per clone). MCF7 cells were analysed by PI staining (n = 3 biological replicates). p, Quantification of RB phosphorylated at S795 (p-RB S795) over total RB in U937 cells treated with increasing doses of palbociclib for 24 h, measured by immunoassay (n = 4 biological replicates). q, Fold-change in mRNA levels of E2F target genes in U937 cells treated with 0.5 μM palbociclib for 24 h, measured by quantitative PCR with reverse transcription (RT–qPCR) (n = 3 biological replicates). All data are mean ± s.d. P values calculated by two-sided unpaired t-test (g, k, m, o) and two-sided paired t-test (i, p, q). Tubulin, HSP90 and actin are loading controls.
Extended Data Fig. 2 AMBRA1 loss regulates cyclin D post-transcriptionally and dependency on AMBRA1 correlates with cyclin D signalling networks.
a, b, RT–qPCR analysis of the genes encoding D-type cyclins (CCND genes) and CDK4 in U937 cells (a) (n = 3 biological replicates per clone) or expressed D-type cyclins in other cancer cell lines (b). For U2OS, NCI-H1792 and NCI-H460 cells, each symbol is an isogenic clone (n = 2 biological replicates per clone). MCF7 cells are populations (n = 3 biological replicates). P values evaluate differences between knockout cells and controls for each gene. c, Immunoassay of D-type cyclins in cancer cell lines in b. d, Immunoassay of AMBRA1, cyclin D1 and CDK4 in U2OS cells after 48 h of AMBRA1 knockdown by siRNA pools. e, Quantification of cyclin D1 and CDK4 protein levels in d (n = 3 biological replicates). f, Correlation of gene dependency scores between AMBRA1, RB pathway genes and additional cancer drivers, according to DepMap. Red lines mark the top and bottom 5% of genes. g, The 20 most significantly enriched gene ontology terms among the top 100 genes, the loss of which best correlate with loss of AMBRA1 in DepMap. h, Principal component (PC) analysis of RNA-sequencing data from U2OS cells, three biological replicates per cell line. i, Volcano plot of RNA-sequencing results comparing control and AMBRA1-knockout U2OS cells. Significantly differentially expressed genes (P < 0.01) are in red. All data shown as mean ± s.d. P values calculated by two-sided unpaired t-test (a, b), two-sided paired t-test (e), hypergeometric test (g) and Wald test (i).
Extended Data Fig. 3 AMBRA1 deletion in mouse embryos results in increased cyclin D levels.
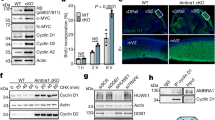
a, sgRNA design to knockout Ambra1 in mouse zygotes by microinjection of sgRNAs and Cas9. Controls were injected with a non-targeting sgRNA. b, Representative bright-field images of control (n = 5) and mutant (n = 3) embryos at embryonic day (E)13.5. Similar to previous reports10, the Ambra1-mutant embryos generated here have neural tube defects with midbrain and hindbrain exencephaly and/or spina bifida (arrows). Scale bar, 2 mm. c, Representative cyclin D immunofluorescence (red signal, the antibody recognizes cyclin D1 and cyclin D2) in control and Ambra1-mutant E13.5 embryos (from n = 3 embryos per sgRNA). DAPI shows DNA. The liver is autofluorescent. Scale bar, 1 mm. d, High-magnification view of the developing brain from one control and one Ambra1-mutant embryo (asterisks in c). v, ventricle, cp, choroid plexus. Scale bar, 500 μm. Representative of three embryos per sgRNA.
Extended Data Fig. 4 Pathways previously associated with AMBRA1 do not explain tolerance to CDK4/6 inhibitors.
a, Immunoblot analysis of autophagy flux by LC3 conversion (LC3-I to LC3-II, which occurs during autophagosome formation) and RB phosphorylation (p-RB S795) in U937 cells treated with 0.5 μM palbociclib for 24 h and acutely treated with 25 μM chloroquine (CQ) (an autophagy inhibitor) for the final 4 h. b, Quantification of LC3-II levels with 4 h of chloroquine treatment, indicating autophagy flux, from cells in a (n = 3 biological replicates). No significant differences were identified by two-way ANOVA (Pcell line = 0.44, Ptreatment = 0.38, Pinteraction = 0.92). c, Immunoblot of total and phosphorylated RB and LC3 conversion in wild-type U937 cells treated with 0.5 μM palbociclib, 25 μM chloroquine or both for 24 h. Representative of three independent experiments. d, Representative flow cytometry plots of BrdU and PI staining in cells from c. e, Quantification of S-phase cells from d (n = 3 biological replicates). Autophagy inhibition does not alter palbociclib response. f, Immunoassay of the mTORC1 target phosphorylation sites (S2448 of mTOR, and T37 and T46 of 4EBP1) in U937 cells following amino acid starvation. Representative of two independent experiments. g, Immunoassay of MYC in U937 clones. h, Quantification of MYC from g. Each symbol is an isogenic clone (n = 3 biological replicates per clone). i, Immunoassay of PLK1 and AURKA and immunoblot of AURKB in control and AMBRA1-knockout U2OS cells. Each lane is a biological replicate. j, Quantification of i (n = 3 biological replicates). All data are mean ± s.d. P values calculated by two-way ANOVA (b), two-sided paired t-test (e, j), and two-sided unpaired t-test (h). HSP90, tubulin and actin are loading controls.
Extended Data Fig. 5 Cyclin D mediates the phenotypes of AMBRA1-mutant cells.
a, Immunoassay of cyclin D1, D2, and D3 in wild-type U2OS cells overexpressing all three D-type cyclins from the same lentiviral vector or RFP as a control. b, Representative flow cytometry plots of BrdU and PI staining in cells from a treated with increasing doses of palbociclib for 24 h. c, Percentage of cycling S-phase cells from b (n = 3 biological replicates). Data are mean ± s.d. P values calculated by two-way ANOVA (Pcell line < 0.0001) with post hoc Sidak test. d, Representative flow cytometry plots of BrdU and PI staining in U2OS cells overexpressing stabilized cyclin D1(T286A)–HA or RFP control, treated with increasing doses of palbociclib for 24 h. e, Representative flow cytometry plots of BrdU and PI staining in control and AMBRA1-knockout U2OS clones after 48 h of cyclin D1 (CCND1) knockdown with siRNA pools. f, Co-immunoprecipitation of p27 in control, knockout and cyclin-D1(T286A)-overexpressing U2OS cells, and immunoassay of relevant protein complexes (n = 2 biological replicates). HSP90 is a loading control.
Extended Data Fig. 6 AMBRA1 regulates the ubiquitylation of D-type cyclins.
a, Immunoblot analysis of cyclin D3 in wild-type U937 cells (left) or cyclin D1 in wild-type U2OS cells (right) treated with 0.5 μM palbociclib, 25 μM chloroquine or both for 24 h. LC3 and HSP90 blots for U937 cells are the same as in Extended Data Fig. 4c, as the experiments were performed simultaneously. Untreated AMBRA1-knockout cells serve as a control for increased cyclin D expression. Asterisk, unspecific band. n = 3 (U937) or n = 1 (U2OS) biological replicates. b, c, Immunoassay quantification of cyclin D2 (b) and cyclin D3 (c) in U2OS cells treated with 1 μM bortezomib for 4 h (n = 4 biological replicates). d, Quantification of ubiquitylated cyclin D1 relative to total cyclin D1 isolated from U2OS clones pretreated with 1 μM bortezomib for 4 h using TUBEs. Each symbol is an isogenic clone (n = 3 (sgCtrl) or n = 5 (sgAMBRA1)). e, f, Immunoassay of ubiquitylated cyclin D1 isolated using TUBEs following AMBRA1 knockdown in U2OS cells (e) or in populations of control and AMBRA1-knockout MCF7 cells (f). g, h, Quantification of ubiquitylated cyclin D1 relative to total cyclin D1 in AMBRA1-knockdown U2OS cells (g) (n = 2 biological replicates) or AMBRA1-knockout MCF7 cells (h) (n = 2 (sgCtrl) or n = 3 (sgAMBRA1) biological replicates) as shown in e, f, respectively. For all TUBE experiments, only quantification of samples with similar levels of ubiquitin pull down are shown. See Supplementary Table 9 for all data. i, Immunoblot analysis of AMBRA1 in 293T cells expressing control or AMBRA1-targeting shRNAs, pretreated with 10 μM MG132 for 4 h. (n = 1 experiment). j, Principal component analysis of mass spectrometry data from cells in i (two replicates each of shNT no. 1 and shAMBRA1 no. 1 and no. 2) after enriching for ubiquitylated peptides. k, Volcano plot of mass-spectrometry data comparing ubiquitylated peptides in control and AMBRA1 knockdown 293T cells. Each dot is a peptide. Red symbols, significantly upregulated peptides; blue symbols, significantly downregulated peptides, with the indicated cut-offs. All other data are mean ± s.d. P values calculated by two-sided paired t-test (b, c), two-sided unpaired t-test (d) and two-sided unpaired t-test followed by Benjamini–Hochberg correction (k). HSP90 and GAPDH are loading controls.
Extended Data Fig. 7 AMBRA1 binding to CUL4 is required for regulating cyclin D.
a, Co-immunoprecipitation of transfected AMBRA1–Myc–Flag and cyclin D–HA (D1, D2 or D3) in 293T cells, analysed by immunoassay. b, Co-immunoprecipitation of transfected Myc-tagged cullin proteins with endogenous AMBRA1 in U2OS cells, analysed by immunoassay. c, RT–qPCR analysis of CCND1 mRNA expression in U2OS cells following knockdown of AMBRA1 or various cullin genes by siRNA pools (n = 3 biological replicates). d, Co-immunoprecipitation of transfected wild-type (WT) AMBRA1 and AMBRA1(ΔH) with endogenous CUL4A and CUL4B in 293T cells. e, Immunoassay of AMBRA1 in control and AMBRA1-knockout U2OS cells with doxycycline-inducible expression of wild-type AMBRA1, AMBRA1(ΔH) or GFP control, after treatment with 500 ng ml−1 doxycycline (+Dox) or DMSO (−Dox) for 2 d. f, Immunoassay of cyclin D1 ubiquitylation in 293T cells with overexpression of wild-type AMBRA1 or AMBRA1(ΔH). Cells were pretreated with 1 μM bortezomib for 3 h and lysed in denaturing conditions before immunoprecipitation of cyclin D1. Representative of two independent experiments. g, Immunoassay of cyclin D1–HA in U2OS cells expressing wild-type cyclin D1 or phosphomutant cyclin D1 (cyclin D1(T286A)) treated with 10 μg ml−1 cycloheximide for up to 2 h. Cells were transfected with control or AMBRA1-targeted siRNA pools 3 d previously. h, Quantification of cyclin D1–HA protein levels in U2OS cells from g with best-fit curves for one-phase decay (n = 3 biological replicates). i, Co-immunoprecipitation of cyclin D1–HA (wild-type or T286A) and endogenous AMBRA1 in U2OS cells. CDK4 serves as a positive control for cyclin D1 binding. Representative of two independent experiments. All data are mean ± s.d. P values calculated by two-sided paired t-test (c) and two-way ANOVA (h). HSP90 and actin are loading controls.
Extended Data Fig. 8 AMBRA1 ubiquitylates cyclin D.
a, Coomassie-blue-stained gel with protein extracts from insect Sf9 cells (–) or Sf9 cells expressing cyclin D1 and CDK4 (arrows). b, Immunoblot for cyclin D1, cyclin D1 phosphorylated on T286 (P-T286) and CDK4 in protein extracts, similar to a. c, Immunoassay of Flag and Myc tag in untransfected 293T cells (−) or 293T cells transfected with AMBRA1–3×Flag or Myc3–CUL4B. Actin is a loading control. n = 1 experiment.
Extended Data Fig. 9 AMBRA1 is a tumour suppressor in KRAS-mutant mouse lung adenocarcinoma.
a, Lollipop plot for RB1 and AMBRA1 mutations in 10,953 patients (10,967 samples) in 32 studies from TCGA (data downloaded from https://cbioportal.org in September 2020). b, Immunoassay of AMBRA1 and cyclin D1 in AMBRA1-knockout U2OS cells upon stable expression of GFP, wild-type AMBRA1 (WT) or two mutant forms of AMBRA1 from a (stop codons at the position indicated by an asterisk). HSP90 is a loading control. Expression of 217* was not detected, suggesting an unstable protein. c, Quantification of cyclin D1 in b (n = 3 biological replicates). Data are mean ± s.d. P values calculated by two-sided paired t-test. d, e, Relative tumour sizes for each sgRNA in KT mice (lacking Cas9) (d) (n = 4 mice) and KPTC mice (e) (n = 5 mice). Tumour sizes were calculated from merged data from all tumours in all mice and normalized to inert sgRNAs 15 (d) or 14 (e) weeks after cancer initiation. f, g, Tumour number for each sgRNA in KTC mice (f) (n = 9 mice) and KPTC mice (g) (n = 5 mice). Data from all tumours in all mice were merged and normalized to the average tumour number across inert sgRNAs. For d–g, error bars denote 95% confidence intervals determined by bootstrap sampling. h, Representative H&E staining of tumours from KC mice infected with lentiviral vectors encoding Cre recombinase and either a control or Ambra1-targeted sgRNA. Scale bar, 100 μm. n = 6 (Neo no. 1) or n = 5 (Ambra1 no. 1) mice). i, Representative immunofluorescence for cyclin D in control and Ambra1-knockout KC tumours. The cyclin D antibody used recognizes cyclin D1 and D2. Scale bars, 100 μm. From n = 2 mice per sgRNA).
Extended Data Fig. 10 AMBRA1 is a tumour suppressor in KRAS-mutant human lung adenocarcinoma.
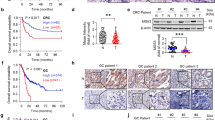
a, Immunoassay of AMBRA1, RB and cyclin D1 in control and knockout human A549 lung adenocarcinoma cells. Actin is a loading control. b, Growth of control and mutant A549 xenografts in NOD-SCID-gamma (NSG) mice (n = 8 tumours per sgRNA). ****Pinteraction < 0.0001 by two-way ANOVA comparing the AMBRA1-knockout curve with control. Tumour volume measurements for RB1-knockout tumours were staggered 1 d behind control and AMBRA1-knockout tumours, which precludes two-way ANOVA. Data are mean ± s.e.m., with best-fit curves for exponential growth. c, Final tumour weights from b. Each symbol is one tumour (n = 8 per sgRNA). Data are mean ± s.d. d, g, j, Cyclin D1 protein levels as measured by reverse phase protein array in relation to the mRNA expression as measured by RNA sequencing (upper quartile of fragments per kilobase of transcript per million mapped reads (FPKM-UQ)) of RB pathway genes that best predict cyclin D1 protein in TCGA KRAS G12-mutant lung adenocarcinoma (d) (n = 90 samples), KRAS wild-type lung adenocarcinoma (g) (n = 257 samples) and EGFR-mutant lung adenocarcinoma (j) (n = 41 samples), using a step-wise regression model. For g, j, AMBRA1 was not selected in the final model but is shown for comparison. Each column is an individual sample, and samples are sorted by cyclin D1 protein levels. e, h, Kaplan–Meier plot of AMBRA1 expression (high, upper third; low, bottom third) in TCGA KRAS wild-type lung adenocarcinoma (e) (n = 361 patients) and EGFR-mutant lung adenocarcinoma (h) (n = 60 patients). f, i, Forest plot of Cox proportional hazard model of TCGA KRAS wild-type lung adenocarcinoma (f) (n = 340 patients) and EGFR-mutant lung adenocarcinoma (i) (n = 60 patients). Model is adjusted by stage, age and gender. P values calculated by two-way ANOVA (b), two-sided unpaired t-test (c), F-test (d, g, j), log-rank test (e, h) and Wald test (f, i).
Extended Data Fig. 11 AMBRA1 regulates cyclin D protein stability and signalling through the RB pathway.
AMBRA1 limits CDK4/6 activity by mediating ubiquitylation and degradation of D-type cyclins as part of the CRL4 E3 ligase complex. Loss of AMBRA1 leads to accumulation of cyclin D protein and decreased sensitivity to CDK4/6 inhibitors, owing to sustained RB phosphorylation and therefore persistent cell cycle progression.
Supplementary information
Supplementary Information
This file contains Methods, References, Supplementary Figure 1: gating strategies for flow cytometry experiments, and Supplementary Methods.
Supplementary Tables 1-3
Analysis of a genome-wide CRISPR/Cas9 screen to identify genetic determinants of palbociclib response in U937 cells. Table 1: casTLE analysis of the screen results. Table 2: Functional enrichment analysis of significant hits by Metascape. Table 3: MCODE clusters identified by Metascape following protein-protein interaction network analysis of significant hits.
Supplementary Table 4
The top 100 genes most highly correlated with AMBRA1 based on dependency score in the CRISPR (Avana) Public 19Q4 dataset from the Cancer Dependency Map (downloaded January 23, 2020).
Supplementary Table 5
Differential gene expression analysis of AMBRA1 knock-out U2OS cells compared to controls, determined by RNA-seq.
Supplementary Table 6-8
Shotgun mass spectrometry analysis of wild-type and AMBRA1 knock-out (KO) U2OS cells, with or without 24 hours of palbociclib treatment. Table 6: Peptide counts. Table 7: Fold-change between wild-type and KO cells. Table 8: Fold-change between untreated and palbociclib-treated conditions.
Supplementary Table 9
Immunoassay quantification of cyclin D1 ubiquitylation upon loss of AMBRA1 in multiple cellular contexts.
Supplementary Table 10
Mass spectrometry analysis of enriched ubiquitylated peptides from wild-type and AMBRA1 knock-down 293T cells.
Source data
Rights and permissions
About this article
Cite this article
Chaikovsky, A.C., Li, C., Jeng, E.E. et al. The AMBRA1 E3 ligase adaptor regulates the stability of cyclin D. Nature 592, 794–798 (2021). https://doi.org/10.1038/s41586-021-03474-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03474-7
This article is cited by
-
Clinically relevant stratification of lung squamous carcinoma patients based on ubiquitinated proteasome genes for 3P medical approach
EPMA Journal (2024)
-
Reciprocal antagonism of PIN1-APC/CCDH1 governs mitotic protein stability and cell cycle entry
Nature Communications (2024)
-
Determinants of response to CDK4/6 inhibitors in the real-world setting
npj Precision Oncology (2023)
-
E3 ligase MG53 suppresses tumor growth by degrading cyclin D1
Signal Transduction and Targeted Therapy (2023)
-
Ambra1 modulates the sensitivity of mantle cell lymphoma to palbociclib by regulating cyclin D1
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.