Abstract
It has recently been shown that in anaerobic microorganisms the tricarboxylic acid (TCA) cycle, including the seemingly irreversible citrate synthase reaction, can be reversed and used for autotrophic fixation of carbon1,2. This reversed oxidative TCA cycle requires ferredoxin-dependent 2-oxoglutarate synthase instead of the NAD-dependent dehydrogenase as well as extremely high levels of citrate synthase (more than 7% of the proteins in the cell). In this pathway, citrate synthase replaces ATP-citrate lyase of the reductive TCA cycle, which leads to the spending of one ATP-equivalent less per one turn of the cycle. Here we show, using the thermophilic sulfur-reducing deltaproteobacterium Hippea maritima, that this route is driven by high partial pressures of CO2. These high partial pressures are especially important for the removal of the product acetyl coenzyme A (acetyl-CoA) through reductive carboxylation to pyruvate, which is catalysed by pyruvate synthase. The reversed oxidative TCA cycle may have been functioning in autotrophic CO2 fixation in a primordial atmosphere that is assumed to have been rich in CO2.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All data generated in this manuscript are included within the paper (and its Supplementary Information). The raw data are presented in the manuscript and/or available from the corresponding authors upon reasonable request. For any further inquiries about our work please contact the corresponding authors. Source data are provided with this paper.
References
Nunoura, T. et al. A primordial and reversible TCA cycle in a facultatively chemolithoautotrophic thermophile. Science 359, 559–563 (2018).
Mall, A. et al. Reversibility of citrate synthase allows autotrophic growth of a thermophilic bacterium. Science 359, 563–567 (2018).
Fuchs, G. Alternative pathways of carbon dioxide fixation: insights into the early evolution of life? Annu. Rev. Microbiol. 65, 631–658 (2011).
Berg, I. A. Ecological aspects of the distribution of different autotrophic CO2 fixation pathways. Appl. Environ. Microbiol. 77, 1925–1936 (2011).
Song, Y. et al. Functional cooperation of the glycine synthase-reductase and Wood–Ljungdahl pathways for autotrophic growth of Clostridium drakei. Proc. Natl Acad. Sci. USA 117, 7516–7523 (2020).
Könneke, M. et al. Ammonia-oxidizing archaea use the most energy-efficient aerobic pathway for CO2 fixation. Proc. Natl Acad. Sci. USA 111, 8239–8244 (2014).
Guynn, R. W., Gelberg, H. J. & Veech, R. L. Equilibrium constants of the malate dehydrogenase, citrate synthase, citrate lyase, and acetyl coenzyme A hydrolysis reactions under physiological conditions. J. Biol. Chem. 248, 6957–6965 (1973).
Bisswanger, H. Enzyme Kinetics: Principles and Methods (Wiley, 2008).
Supek, F. & Vlahovicek, K. Comparison of codon usage measures and their applicability in prediction of microbial gene expressivity. BMC Bioinformatics 6, 182 (2005).
Supek, F. & Vlahovicek, K. INCA: synonymous codon usage analysis and clustering by means of self-organizing map. Bioinformatics 20, 2329–2330 (2004).
Miroshnichenko, M. L., Rainey, F. A., Rhode, M. & Bonch-Osmolovskaya, E. A. Hippea maritima gen. nov., sp. nov., a new genus of thermophilic, sulfur-reducing bacterium from submarine hot vents. Int. J. Syst. Bacteriol. 49, 1033–1038 (1999).
Schauder, R., Widdel, F. & Fuchs, G. Carbon assimilation pathways in sulfate-reducing bacteria. II. Enzymes of a reductive citric acid cycle in the autotrophic Desulfobacter hydrogenophilus. Arch. Microbiol. 148, 218–225 (1987).
Bennett, B. D. et al. Absolute metabolite concentrations and implied enzyme active site occupancy in Escherichia coli. Nat. Chem. Biol. 5, 593–599 (2009).
Greene, A. C. in The Prokaryotes (eds Rosenberg, E. et al.) 135–142 (Springer, 2014).
Dahle, H. et al. Energy landscapes in hydrothermal chimneys shape distributions of primary producers. Front. Microbiol. 9, 1570 (2018).
Zhao, W. Diversity and Potential Geochemical Functions of Prokaryotes in Hot Springs of the Uzon Caldera, Kamchatka. PhD thesis, Univ. Georgia, Athens (2008).
Obzhirov, A. I. Chemistry of free and dissolved gases of Matupit Bay, Rabaul Caldera, Papua New Guinea. Geo-Mar. Lett. 12, 54–59 (1992).
Zhang, T., Shi, X.-C., Ding, R., Xu, K. & Tremblay, P.-L. The hidden chemolithoautotrophic metabolism of Geobacter sulfurreducens uncovered by adaptation to formate. ISME J. 14, 2078–2089 (2020).
Spieck, E. et al. Extremophilic nitrite-oxidizing Chloroflexi from Yellowstone hot springs. ISME J. 14, 364–379 (2020).
Slobodkin, A. et al. Genomic insights into the carbon and energy metabolism of a thermophilic deep-sea bacterium Deferribacter autotrophicus revealed new metabolic traits in the phylum Deferribacteres. Genes 10, 849 (2019).
Hua, Z. S. et al. Genomic inference of the metabolism and evolution of the archaeal phylum Aigarchaeota. Nat. Commun. 9, 2832 (2018).
Berg, I. A. et al. Autotrophic carbon fixation in archaea. Nat. Rev. Microbiol. 8, 447–460 (2010).
Zahnle, K. et al. Emergence of a habitable planet. Space Sci. Rev. 129, 35–78 (2007).
German, C. R. & Von Damm, K. L. in Treatise on Geochemistry (eds Turekian, K. K. & Holland, H. D.) 18–222 (Elsevier, 2006).
Pedersen, R. B. et al. Discovery of a black smoker vent field and vent fauna at the Arctic Mid-Ocean Ridge. Nat. Commun. 1, 126 (2010).
Lupton, J. et al. Submarine venting of liquid carbon dioxide on a Mariana Arc volcano. Geochem. Geophys. Geosyst. 7, Q08007 (2006).
Ajo-Franklin, J., Voltolini, M., Molins, S. & Yang, L. in Geological Carbon Storage: Subsurface Seals and Caprock Integrity (eds Vialle, S. et al.) 187–206 (Wiley, 2019).
Probst, A. J. et al. Lipid analysis of CO2-rich subsurface aquifers suggests an autotrophy-based deep biosphere with lysolipids enriched in CPR bacteria. ISME J. 14, 1547–1560 (2020).
Martin, W. F. Older than genes: the acetyl CoA pathway and origins. Front. Microbiol. 11, 817 (2020).
Thauer, R. K., Kaster, A. K., Seedorf, H., Buckel, W. & Hedderich, R. Methanogenic archaea: ecologically relevant differences in energy conservation. Nat. Rev. Microbiol. 6, 579–591 (2008).
Schuchmann, K. & Müller, V. Autotrophy at the thermodynamic limit of life: a model for energy conservation in acetogenic bacteria. Nat. Rev. Microbiol. 12, 809–821 (2014).
Simon, E. J. & Shemin, D. The preparation of S-succinyl coenzyme A. J. Am. Chem. Soc. 75, 2520 (1953).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Eylert, E. et al. Carbon metabolism of Listeria monocytogenes growing inside macrophages. Mol. Microbiol. 69, 1008–1017 (2008).
Ahmed, Z. et al. ‘Isotopo’ a database application for facile analysis and management of mass isotopomer data. Database 2014, bau077 (2014).
Ausubel, F. M. et al. Current Protocols in Molecular Biology (Wiley, 1987).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Riddles, P. W., Blakeley, R. L. & Zerner, B. Ellman’s reagent: 5,5′-dithiobis(2-nitrobenzoic acid)—a reexamination. Anal. Biochem. 94, 75–81 (1979).
Bergmeyer, H. U. Neue Werte für die molaren Extinktions-Koeffizienten von NADH und NADPH zum Gebrauch im Routine-Laboratorium. Z. Klin. Chem. Klin. Biochem. 13, 507–508 (1975).
Hügler, M., Huber, H., Molyneaux, S. J., Vetriani, C. & Sievert, S. M. Autotrophic CO2 fixation via the reductive tricarboxylic acid cycle in different lineages within the phylum Aquificae: evidence for two ways of citrate cleavage. Environ. Microbiol. 9, 81–92 (2007).
Flint, D. H. Initial kinetic and mechanistic characterization of Escherichia coli fumarase A. Arch. Biochem. Biophys. 311, 509–516 (1994).
Dawson, R. M. C., Elliott, D. C., Elliott, W. H. & Jones, K. M. Data for Biochemical Research (Clarendon, 1986).
Distler, U., Kuharev, J., Navarro, P. & Tenzer, S. Label-free quantification in ion mobility-enhanced data-independent acquisition proteomics. Nat. Protoc. 11, 795–812 (2016).
Wildschütz, L. et al. Transcriptomic and proteomic analysis of iris tissue and aqueous humor in juvenile idiopathic arthritis-associated uveitis. J. Autoimmun. 100, 75–83 (2019).
Trüper, H. G. & Schlegel, H. G. Sulphur metabolism in Thiorhodaceae. I. Quantitative measurements on growing cells of Chromatium okenii. Antonie van Leeuwenhoek 30, 225–238 (1964).
Laemmli, U. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Noor, E. et al. Pathway thermodynamics highlights kinetic obstacles in central metabolism. PLOS Comput. Biol. 10, e1003483 (2014).
Witt, A., Pozzi, R., Diesch, S., Hädicke, O. & Grammel, H. New light on ancient enzymes – in vitro CO2 fixation by pyruvate synthase of Desulfovibrio africanus and Sulfolobus acidocaldarius. FEBS J. 286, 4494–4508 (2019).
Furdui, C. & Ragsdale, S. W. The role of pyruvate ferredoxin oxidoreductase in pyruvate synthesis during autotrophic growth by the Wood–Ljungdahl pathway. J. Biol. Chem. 275, 28494–28499 (2000).
Flamholz, A., Noor, E., Bar-Even, A. & Milo, R. eQuilibrator—the biochemical thermodynamics calculator. Nucleic Acids Res. 40, D770–D775 (2012).
Acknowledgements
We thank G. Fuchs for discussions during this work and for critical reading of the manuscript, W. Schulz for his help with cloning of citrate synthase genes, A. M. Berg and D. Ackermann for technical assistance and A. Probst for the help with bioinformatics analysis. This work was supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) Project-ID BE 4822/5-1 to I.A.B. and Project-ID 364653263–TRR 235 to W.E. as well as the Hans-Fischer Gesellschaft (Munich) to W.E.
Author information
Authors and Affiliations
Contributions
L.S. and E.P. performed growth experiments, cloning, purification and characterization of enzymes and enzyme assays. L.S performed codon usage analysis and sulfide determination. T.M.S. performed isotopologue profiling experiments and GC–MS analysis of H. maritima medium. S.K. conducted proteomics analyses. A.M. performed phylogenetic analysis and bioenergetics calculations. I.A.B. and W.E. wrote the manuscript with input from other authors. L.S., E.P., T.M.S., A.M., S.K. and W.E. prepared figures. The manuscript was reviewed and approved by all coauthors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks William Martin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Phylogenetic tree of citrate-cleaving enzymes.
Sequences were aligned using MUSCLE47, the phylogeny was calculated with IQtree48 using the maximum likelihood method (LG + R6 substitution model). The scale bar represents amino acid substitutions per site. Branch labels are SH-aLRT support/ultrafast bootstrap support (%) values from 1,000 replications. The sequences used to build the tree are available in Supplementary Table 11.
Extended Data Fig. 2 SDS–PAGE (12.5%) of recombinant citrate synthases from D. acetivorans after purification with Ni-NTA column.
M, molecular mass standard proteins; lane 1, Desace_08345 (2 μg); lane 2, Desace_06860 (2 μg); lane 3, Desace_09325 (2 μg). Their predicted molecular masses are 49 kDa (Desace_08345) and 50 kDa (Desace_06860; Desace_09325). Proteins were stained with Coomassie blue. For gel source data, see Supplementary Fig. 1. The SDS–PAGE analysis was conducted three times and all results were similar to the ones shown here.
Extended Data Fig. 3 Protein quantification using MS.
a, CLUSTAL format alignment by MAFFT (v.7.452) of sequence parts of citrate synthase isoforms to illustrate the unique peptides chosen for quantification. Desace_06, Desace_06860; Desace_08, Desace_08345; Desace_09, Desace_09325. b, Calibration curves generated for each citrate synthase using tryptic digests of the recombinant proteins isolated by one-dimensional PAGE. Injection volume of standard solution (μl, y axis) versus MS peak area in MRM experiments (x axis) as calculated by Skyline.
Extended Data Fig. 4 Metabolites in H. maritima growth medium.
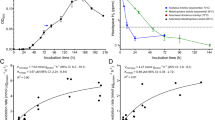
a, Sulfide production during growth under mixotrophic conditions (CO2, H2, S0 and 0.2 g l−1 yeast extract). b, Concentrations of potential fermentation products and some amino acids in the medium after inoculation and after 1, 2 and 3 days of cultivation. Data are mean ± s.e.m. of four biological replicates for sulfide determination and of three biological and three technical replicates for the potential fermentation products and amino acids in the medium.
Extended Data Fig. 5 Growth of H. maritima with different substrates.
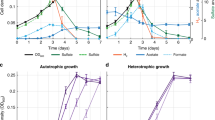
a, Growth of H. maritima in 80% H2, 20% CO2, 0.2 g l−1 yeast extract and 5 g l−1 acetate, in 80% N2, 20% CO2, 0.2 g l−1 yeast extract and 5 g l−1 acetate, and in 100% N2, 0.2 g l−1 yeast extract and 5 g l−1 acetate. b, Growth of H. maritima in 80% H2, 20% N2 and 0.2 g l−1 yeast extract, in 80% N2, 20% CO2 and 0.2 g l−1 yeast extract, in 100% N2 and 0.2 g l−1 yeast extract, in 80% H2 and 20% CO2, and in 80% H2, 20% CO2 and 0.2 g l−1 yeast extract. For all growth curves, data are presented as mean ± s.e.m. of three biological replicates.
Extended Data Fig. 6 The labelling in C1 of Ala, Asp and Glu after growth in H2, S0, 0.2 g l−1 yeast extract and 4, 10 or 20% 13CO2.
The labelling was calculated from the isotopologue composition of the corresponding amino acids and their fragments after the loss of the carboxylic atom (Supplementary Tables 4–6). Data are mean ± s.e.m. of six biological replicates for 4% 13CO2 and of three biological replicates for 10% and 20% 13CO2.
Extended Data Fig. 7 Dependency of energetic efficiencies of the rTCA cycle variants, pyruvate synthase and 2-oxoglutarate synthase reactions on CO2 concentration.
Net equations are as follows2. roTCA cycle (citrate synthase, as in Desulfurellaceae; red and orange lines): 2 CO2(total) + NADPH + 2 NADH + 2 Fdred + CoA + ATP ↔ acetyl-CoA + NADP+ + 2 NAD+ + 2 Fdox + ADP + Pi + 4 H2O. rTCA cycle in Chlorobium (ACL; dark and light blue lines): 2 CO2(total) + NADPH + 2 NADH + 2 Fdred + CoA + 2 ATP ↔ acetyl-CoA + NADP+ + 2 NAD+ + 2 Fdox + 2 ADP + 2 Pi + 3 H2O. rTCA cycle in Hydrogenobacter thermophilus (citryl-CoA synthase/citryl-CoA lyase and ATP-dependent 2-oxoglutarate carboxylase; green and olive lines): 2 CO2(total) + 3 NADH + 2 Fdred + CoA + 3 ATP ↔ acetyl-CoA + 3 NAD+ + 2 Fdox + 3 ADP + 3 Pi + 2 H2O. Fumarate reductase is assumed to be NADH-dependent2. For ferredoxin, a reduction potential E°' of −418 mV, corresponding to the hydrogen/proton couple, was assumed. Temperature is assumed 25 °C, pH 7, ionic strength 0.1 M. CO2 is the sum of CO2(aq) and its hydrated forms (H2CO3, HCO3− and CO32−). Under the D. acetivorans growth conditions (55 °C, pH 7, 2 bar pressure, 30.15 g l−1 salinity), a CO2 partial pressure of 40% corresponds to 147.6 mM CO2, a partial pressure of 1% to 3.7 mM. a, Standard conditions (1 M concentrations for all reactants except CO2). b, Assumed physiological metabolite concentrations (0.1 mM NADPH, 0.01 mM NADP+, 0.1 mM NADH, 1 mM NAD+, 5 mM ATP, 0.5 mM ADP, 10 mM phosphate49, 1.07 mM CoA, 0.013 mM acetyl-CoA2, 0.1 mM Fdred, 0.001/0.0001 mM Fdox (refs. 3,50), 0.23 mM succinyl-CoA, 0.44 mM 2-oxoglutarate13 and 0.18 mM pyruvate51). The calculations were done using eQuilibrator52.
Extended Data Fig. 8 Dependency of energetic efficiency of the reductive acetyl-CoA pathway in methanogens and acetogens on CO2 concentration.
Net equation of the reductive acetyl-CoA pathway in methanogens and acetogens are as follows2. Methanogens (Methanothermobacter marburgiensis; red and orange lines): 2 CO2(total) + 4 Fdred + CoA + 2 F420H2 ↔ acetyl-CoA + 4 Fdox + 2 F420 + 5 H2O. Acetogens (Acetobacterium woodii; dark and light blue lines): 2 CO2(total) + NADPH + NADH + 4 Fdred + CoA + ATP ↔ acetyl-CoA + NADP+ + NAD+ + 4 Fdox + ADP + Pi + 4 H2O. A free energy change (ΔG′) of acetogenesis and methanogenesis is usually not sufficient to drive the synthesis of 1 mol ATP per mol of product under physiological conditions, and energy generation requires chemiosmotic coupling and entails flavin-based electron bifurcation30,31. However, a high CO2 partial pressure may enable the functioning of a hybrid reductive acetyl-CoA pathway that combines CO2 reduction with ferredoxin, as in methanogens, and acetate formation from acetyl-CoA coupled with substrate phosphorylation, as in acetogens. Calculations were done as in Extended Data Fig. 7. a, Standard conditions (1 M concentrations for all reactants except CO2). b, Assumed physiological metabolite concentrations (0.1 mM NADPH, 0.01 mM NADP+, 0.1 mM NADH, 1 mM NAD+, 5 mM ATP, 0.5 mM ADP, 0.1 mM reduced coenzyme F420, 0.1 mM oxidized coenzyme F420, 10 mM phosphate49, 0.28 mM CoA, 0.0104 mM acetyl-CoA51, 0.1 mM Fdred, 0.001/0.0001 mM Fdox (refs. 3,50).
Supplementary information
Supplementary Information
This file contains Supplementary Tables 2, 3, 4, 5, 6, 7, 8, 10, a Supplementary Discussion and Supplementary References.
Supplementary Figure 1
Gel source data for Extended Data Fig. 2.
Supplementary Table 1
Proteome expression data for cell lysates of D. acetivorans grown under autotrophic (CO2 + H2 + S0) and heterotrophic (acetate + CO2 + S0) conditions. Shown is the processing output as generated by Progenesis QIP containing One Way Anova P values, Q values and power as well as further parameters such as the number of unique peptides. Entries have been coloured for improved visualisation and according to criteria such as p = 0.05 and fold values < 2. A second Excel sheet was created by deleting all those entries not matching conditions p < 0.05, fold value > 2.
Supplementary Table 9
Proteome expression data for cell lysates of H. maritima grown with 4 and 20% CO2 in the gas mixture under mixotrophic conditions (CO2 + 80% H2 + S0 + 0.2 g l-1 yeast extract). Shown is the processing output as generated by Progenesis QIP containing One Way Anova P values, Q values and power as well as further parameters such as the number of unique peptides. Selection criteria were chosen as in Supplementary Table 1.
Supplementary Table 11
Alignment for Extended Data Fig. 1.
Rights and permissions
About this article
Cite this article
Steffens, L., Pettinato, E., Steiner, T.M. et al. High CO2 levels drive the TCA cycle backwards towards autotrophy. Nature 592, 784–788 (2021). https://doi.org/10.1038/s41586-021-03456-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03456-9
This article is cited by
-
Current status of carbon monoxide dehydrogenases (CODH) and their potential for electrochemical applications
Bioresources and Bioprocessing (2023)
-
Fiber supplementation protects from antibiotic-induced gut microbiome dysbiosis by modulating gut redox potential
Nature Communications (2023)
-
Description and Genomic Analysis of the First Facultatively Lithoautotrophic, Thermophilic Bacteria of the Genus Thermaerobacter Isolated from Low-temperature Sediments of Lake Baikal
Microbial Ecology (2023)
-
Hohe CO2-Konzentrationen treiben den Citratzyklus zurück zur Autotrophie
BIOspektrum (2023)
-
Recent advances of microbial metabolism analysis: from metabolic molecules to environments
Science China Chemistry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.