Abstract
Genetic recombination arises during meiosis through the repair of DNA double-strand breaks (DSBs) that are created by Spo11, a topoisomerase-like protein1,2. Spo11 DSBs form preferentially in nucleosome-depleted regions termed hotspots3,4, yet how Spo11 engages with its DNA substrate to catalyse DNA cleavage is poorly understood. Although most recombination events are initiated by a single Spo11 cut, here we show in Saccharomyces cerevisiae that hyperlocalized, concerted Spo11 DSBs separated by 33 to more than 100 base pairs also form, which we term ‘double cuts’. Notably, the lengths of double cuts vary with a periodicity of 10.5 base pairs, which is conserved in yeast and mice. This finding suggests a model in which the orientation of adjacent Spo11 molecules is fixed relative to the DNA helix—a proposal supported by the in vitro DNA-binding properties of the Spo11 core complex. Deep sequencing of meiotic progeny identifies recombination scars that are consistent with repair initiated from gaps generated by adjacent Spo11 DSBs. Collectively, these results revise our present understanding of the mechanics of Spo11-DSB formation and expand on the original concepts of gap repair during meiosis to include DNA gaps that are generated by Spo11 itself.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw S. cerevisiae and mouse Spo11-oligo FASTQ data were obtained from published archives GSE84696 and GSE84689, respectively. Nucleotide-resolution maps generated by paired-end Bowtie2 alignment are provided in the Supplementary Information. For mice, the maps used here were generated from the following biological samples: wild type, GSM2247728; Atm−/−, GSM2247731. FASTQ files used for mapping HR patterns in S. cerevisiae SK1 × S288c F1 hybrid octads in msh2∆; tel1∆msh2∆; and mlh1∆msh2∆, mlh3∆msh2∆ and exo1∆msh2∆ are deposited in the NCBI Sequence Read Archive (SRA) with accession numbers PRJNA479661; PRJNA480956; and PRJNA39308719, respectively. Additional data are included in the Supplementary Data. Source data are provided with this paper.
References
Keeney, S., Giroux, C. N. & Kleckner, N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88, 375–384 (1997).
Bergerat, A. et al. An atypical topoisomerase II from Archaea with implications for meiotic recombination. Nature 386, 414–417 (1997).
Pan, J. et al. A hierarchical combination of factors shapes the genome-wide topography of yeast meiotic recombination initiation. Cell 144, 719–731 (2011).
Lange, J. et al. The landscape of mouse meiotic double-strand break formation, processing, and repair. Cell 167, 695–708 (2016).
Neale, M. J., Pan, J. & Keeney, S. Endonucleolytic processing of covalent protein-linked DNA double-strand breaks. Nature 436, 1053–1057 (2005).
Garcia, V., Phelps, S. E. L., Gray, S. & Neale, M. J. Bidirectional resection of DNA double-strand breaks by Mre11 and Exo1. Nature 479, 241–244 (2011).
Fowler, K. R., Sasaki, M., Milman, N., Keeney, S. & Smith, G. R. Evolutionarily diverse determinants of meiotic DNA break and recombination landscapes across the genome. Genome Res. 24, 1650–1664 (2014).
Choi, K. et al. Nucleosomes and DNA methylation shape meiotic DSB frequency in Arabidopsis thaliana transposons and gene regulatory regions. Genome Res. 28, 532–546 (2018).
Cannavo, E. & Cejka, P. Sae2 promotes dsDNA endonuclease activity within Mre11-Rad50-Xrs2 to resect DNA breaks. Nature 514, 122–125 (2014).
Garcia, V., Gray, S., Allison, R. M., Cooper, T. J. & Neale, M. J. Tel1ATM-mediated interference suppresses clustered meiotic double-strand-break formation. Nature 520, 114–118 (2015).
Mohibullah, N. & Keeney, S. Numerical and spatial patterning of yeast meiotic DNA breaks by Tel1. Genome Res. 27, 278–288 (2017).
Zhang, L., Kim, K. P., Kleckner, N. E. & Storlazzi, A. Meiotic double-strand breaks occur once per pair of (sister) chromatids and, via Mec1/ATR and Tel1/ATM, once per quartet of chromatids. Proc. Natl Acad. Sci. USA 108, 20036–20041 (2011).
Joyce, E. F. et al. Drosophila ATM and ATR have distinct activities in the regulation of meiotic DNA damage and repair. J. Cell Biol. 195, 359–367 (2011).
Lange, J. et al. ATM controls meiotic double-strand-break formation. Nature 479, 237–240 (2011).
Carballo, J. A. et al. Budding yeast ATM/ATR control meiotic double-strand break (DSB) levels by down-regulating Rec114, an essential component of the DSB-machinery. PLoS Genet. 9, e1003545 (2013).
Liu, J., Wu, T. C. & Lichten, M. The location and structure of double-strand DNA breaks induced during yeast meiosis: evidence for a covalently linked DNA-protein intermediate. EMBO J. 14, 4599–4608 (1995).
Claeys Bouuaert, C. et al. Structural and functional characterization of the Spo11 core complex. Nat. Struct. Mol. Biol. 28, 92–102 (2021).
Martini, E. et al. Genome-wide analysis of heteroduplex DNA in mismatch repair-deficient yeast cells reveals novel properties of meiotic recombination pathways. PLoS Genet. 7, e1002305 (2011).
Marsolier-Kergoat, M. C., Khan, M. M., Schott, J., Zhu, X. & Llorente, B. Mechanistic view and genetic control of DNA recombination during meiosis. Mol. Cell 70, 9–20 (2018).
Kulkarni, D. S. et al. PCNA activates the MutLγ endonuclease to promote meiotic crossing over. Nature 586, 623–627 (2020).
Cannavo, E. et al. Regulation of the MLH1-MLH3 endonuclease in meiosis. Nature 586, 618–622 (2020).
Szostak, J. W., Orr-Weaver, T. L., Rothstein, R. J. & Stahl, F. W. The double-strand-break repair model for recombination. Cell 33, 25–35 (1983).
Diaz, R. L., Alcid, A. D., Berger, J. M. & Keeney, S. Identification of residues in yeast Spo11p critical for meiotic DNA double-strand break formation. Mol. Cell. Biol. 22, 1106–1115 (2002).
Prieler, S. et al. Spo11 generates gaps through concerted cuts at sites of topological stress. Nature, https://doi.org/10.1038/s41586-021-03632-x (2021).
Noll, M. Internal structure of the chromatin subunit. Nucleic Acids Res. 1, 1573–1578 (1974).
Gittens, W. H. et al. A nucleotide resolution map of Top2-linked DNA breaks in the yeast and human genome. Nat. Commun. 10, 4846 (2019).
Prieler, S., Penkner, A., Borde, V. & Klein, F. The control of Spo11’s interaction with meiotic recombination hotspots. Genes Dev. 19, 255–269 (2005).
Panizza, S. et al. Spo11-accessory proteins link double-strand break sites to the chromosome axis in early meiotic recombination. Cell 146, 372–383 (2011).
Kugou, K. et al. Rec8 guides canonical Spo11 distribution along yeast meiotic chromosomes. Mol. Biol. Cell 20, 3064–3076 (2009).
Li, J., Hooker, G. W. & Roeder, G. S. Saccharomyces cerevisiae Mer2, Mei4 and Rec114 form a complex required for meiotic double-strand break formation. Genetics 173, 1969–1981 (2006).
Kee, K., Protacio, R. U., Arora, C. & Keeney, S. Spatial organization and dynamics of the association of Rec102 and Rec104 with meiotic chromosomes. EMBO J. 23, 1815–1824 (2004).
Kumar, R., Bourbon, H. M. & de Massy, B. Functional conservation of Mei4 for meiotic DNA double-strand break formation from yeasts to mice. Genes Dev. 24, 1266–1280 (2010).
Claeys Bouuaert, C. et al. DNA-driven condensation assembles the meiotic DNA break machinery. Nature 592, 144–149 (2021).
Lukaszewicz, A., Lange, J., Keeney, S. & Jasin, M. De novo deletion mutations at recombination hotspots in mouse germlines. Preprint at https://doi.org/10.1101/2020.06.23.168138 (2020).
Crawford, M., Cooper, T. J., Marsolier-Kergoat, M.-C., Llorente, B. & Neale, M. J. Separable roles of the DNA damage response kinase Mec1ATR and its activator Rad24RAD17 within the regulation of meiotic recombination. Preprint at https://doi.org/10.1101/496182 (2018).
Johnson, D., Allison, R. M., Cannavo, E., Cejka, P. & Neale, M. Removal of Spo11 from meiotic DNA breaks in vitro but not in vivo by tyrosyl DNA phosphodiesterase 2. Preprint at https://doi.org/10.1101/527333 (2019).
Xu, L. & Kleckner, N. Sequence non-specific double-strand breaks and interhomolog interactions prior to double-strand break formation at a meiotic recombination hot spot in yeast. EMBO J. 14, 5115–5128 (1995).
Kane, S. M. & Roth, R. Carbohydrate metabolism during ascospore development in yeast. J. Bacteriol. 118, 8–14 (1974).
Cortes Ledesma, F., El Khamisy, S. F., Zuma, M. C., Osborn, K. & Caldecott, K. W. A human 5′-tyrosyl DNA phosphodiesterase that repairs topoisomerase-mediated DNA damage. Nature 461, 674–678 (2009).
Cao, L., Alani, E. & Kleckner, N. A pathway for generation and processing of double-strand breaks during meiotic recombination in S. cerevisiae. Cell 61, 1089–1101 (1990).
Alani, E., Padmore, R. & Kleckner, N. Analysis of wild-type and rad50 mutants of yeast suggests an intimate relationship between meiotic chromosome synapsis and recombination. Cell 61, 419–436 (1990).
Furuse, M. et al. Distinct roles of two separable in vitro activities of yeast Mre11 in mitotic and meiotic recombination. EMBO J. 17, 6412–6425 (1998).
Moreau, S., Ferguson, J. R. & Symington, L. S. The nuclease activity of Mre11 is required for meiosis but not for mating type switching, end joining, or telomere maintenance. Mol. Cell. Biol. 19, 556–566 (1999).
Blat, Y., Protacio, R. U., Hunter, N. & Kleckner, N. Physical and functional interactions among basic chromosome organizational features govern early steps of meiotic chiasma formation. Cell 111, 791–802 (2002).
Acquaviva, L. et al. The COMPASS subunit Spp1 links histone methylation to initiation of meiotic recombination. Science 339, 215–218 (2013).
Sommermeyer, V., Béneut, C., Chaplais, E., Serrentino, M. E. & Borde, V. Spp1, a member of the Set1 complex, promotes meiotic DSB formation in promoters by tethering histone H3K4 methylation sites to chromosome axes. Mol. Cell 49, 43–54 (2013).
Acknowledgements
We thank S. Yamada and N. Mohibullah for help accessing and analysing the mouse and yeast Spo11-oligo datasets, M.-C. Marsolier-Kergoat for sharing Python scripts, J. Carballo and M. Lichten for sharing S. cerevisiae strains containing relevant constructs (tel1∆::hphNT2 and sae2∆::kanMX6, respectively), K. Caldecott and A. Oliver for sharing recombinant TDP2 and R. Allison for critical reading of the manuscript. D.J., V.G., T.C. and M.J.N. were supported by an ERC Consolidator Grant (311336), the BBSRC (BB/M010279/1), the Wellcome Trust (200843/Z/16/Z) and a Career Development Award from the Human Frontier Science Program (CDA00060/2010). B.L. and V.G. were supported by the ANR-13-BSV6-0012-01 and ANR-16-CE12-0028-01 grants from the Agence Nationale de la Recherche and a grant from the Fondation ARC pour la Recherche sur le Cancer (PJA20181207756). Work in the S.K. laboratory was supported by the Howard Hughes Medical Institute. Memorial Sloan Kettering core facilities are supported by grant P30 CA008748 from the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
M.J.N. and V.G. conceived the project and prepared the manuscript. D.J., V.G., C.C.B. and M.J.N. performed physical analysis of Spo11-DCs. M.C. prepared and analysed whole-genome recombination maps with B.L. advising on mechanistic interpretation. T.C. and M.J.N. mapped and analysed Spo11-oligo library data. C.C.B. and S.K. contributed protein biochemistry and provided critical mechanistic interpretations. All authors helped to write the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Bernard de Massy and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Spo11-DC sizing, quantification and genetic analysis.
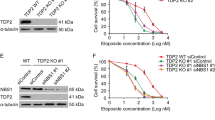
a–h, Immunoprecipitated S. cerevisiae Spo11-oligos and Spo11-DCs isolated from meiotic extracts of the indicated mutants (at 5 h, or indicated number of hours, after induction of meiosis) were radiolabelled with chain-terminating 3′-dATP using terminal transferase and separated on 19% denaturing PAGE following digestion with proteinase K. Where indicated, samples were also treated with mammalian TDP239, which removes the residual Spo11 peptide that is left after proteinase K treatment36, thereby permitting accurate estimation of the Spo11-DC oligo length. In the absence of TDP2 digestion, the residual 5′-linked Spo11 peptide retards the migration of Spo11-oligos and Spo11-DCs by the equivalent of around 6–8 nt (b–d, g). A 10-nt ladder (also radiolabelled with 3′-dATP) is included in each gel to permit accurate sizing. Spo11-DCs arose in tel1∆ cells in the presence and absence of SAE2 (a, b, h). Spo11-DCs also arose independently of single or dual mutations in the MRX complex (rad50S, mre11H125N, mre11D56N) that abrogate endonuclease activity1,5,6,40,41,42,43 (c), and were abolished by homozygous mutation of the Spo11 active site (spo11Y135F) (d). Loading normalization was not performed on samples in c, therefore differences in Spo11-DC abundance do not convey information. In c, d, Spo11-DCs were not treated with TDP2, leading to slower migration. In e, f, representative 4-h lane traces of sequencing gels shown in Fig. 1c are shown using two modes of background subtraction (top and middle), with the resulting maximum, minimum and mid Spo11-DC quantifications (bottom). Shaded areas in top panels are the area being quantified. Shaded areas in lower quantification data show the range between maximum and minimum, as indicated in the figure. Quantified average mid-values are reported in Fig. 1d (minimum–maximum range of 8–25%). Further quantification details are provided in Methods. SPO11/spo11Y135F heterozygous diploids (g) display an altered Spo11-DC oligo size distribution (biological duplicate lanes of each are presented alongside averaged intensity trace). h, Analysis of Spo11-oligo and Spo11-DC intermediates at hourly time points during meiotic prophase in the absence of Tel1.
Extended Data Fig. 2 Biased sequence composition around Spo11-DC 5′ ends.
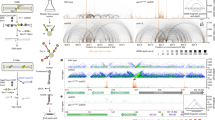
Spo11-oligos11 isolated from the indicated S. cerevisiae strains were remapped using paired-end Bowtie2 alignment. a, Size distribution of total Spo11-oligos in wild-type and tel1∆ strains. Periodic peaks in the distribution are indicated, including a subtle shoulder at 33 nt, consistent with the Spo11-DC sizes detected on gels. b, Top, cartoon depicting Spo11 dimer staggered cuts; bottom, mean nucleotide composition surrounding the Spo11 cleavage site (Methods). Population-averaged features of Spo11 cleavage sites include preferred cleavage 3′ to a C nucleotide and flanking A/T skews3,26. c–e, Nucleotide composition of Spo11-oligos of the indicated size was computed for each base position, revealing a Spo11 signature at both ends (d), or a Spo11 signature at the 5′ end plus a Mre11 signature at the 3′ end (c, e). f–i, Spo11-DCs were filtered out from total Spo11-oligo libraries based on overlapping molecules sharing 5′ and 3′ coordinates with a precise 2-nt offset (f). The theoretical dyad axis of each Spo11 dimer (at each end of the molecule) is indicated. Owing to the rotational symmetry of cleavage, the distance between such dyad axes is identical to the filtered oligo length. g, Ratio of filtered to total Spo11-oligos plotted as a function of molecule length. Because molecules of less than 30-nt length were not detected on Spo11-oligo gels in sae2∆, we infer that the retention of some molecules that are smaller than 30 nt is due to the fortuitous artefactual overlap of canonical Spo11-oligos (around 27 nt). Such filtered molecules were therefore excluded from all bioinformatic analyses. h, The mean nucleotide composition of filtered Spo11-oligos of the indicated size (the sizes presented are peaks in the filtered size distribution) was computed for each base position and plotted relative to the inferred dyad axis of cleavage of the leftmost Spo11 DSB, revealing signature nucleotide skews characteristic of Spo11 at both the 5′ and the 3′ ends. No base skews were observed in the central regions of each molecule, arguing against a major influencer of Spo11-DC formation being localized DNA bending, which is expected to be favoured by an AT-rich base composition. i, The percentage of the total Spo11-oligo library in the filtered (Spo11-DC) fraction is plotted for the indicated size ranges. As defined, Spo11-DCs make up around 4.6% and 7.9% of the total pool of Spo11-oligos in wild-type and tel1∆ strains, respectively, consistent with the tel1∆-dependent increase measured by our physical analysis. In absolute terms, these values are presumably lower than our gel-based estimates owing to size selection during library preparation, the stringency of the filtering, and inaccuracies in quantifying Spo11-DCs on gels, which we estimate make up less than 1% of the total cellular Spo11 protein (Methods). j, In vitro DNA mobility shift assay as described17. Spo11 core complex (Spo11, Rec102, Rec104 and Ski8) was incubated with dsDNA substrate of different lengths with a 2-nt TA overhang on both ends. On the basis of previous experiments17, the robust supershift observed at around 3 nM is interpreted to indicate double-end binding by the Spo11 core complex. Quantification is provided in Fig. 2d. Although the in vitro assay involves heterotetrameric Spo11 complexes (that is, a Spo11 core-complex monomer), we assume that similar binding characteristics will take place in vivo involving octameric complexes (that is, Spo11 core-complex dimers).
Extended Data Fig. 3 Spo11-DC composition of S. cerevisiae DSB hotspots in wild-type and tel1∆ yeast.
a, Percentage of total Spo11-oligos and filtered Spo11-DCs that arise within annotated DSB hotspots. Overall, nearly all (95%) of the Spo11-DCs map within hotspots—more so than total Spo11-oligos (86%)—suggesting that Spo11-DCs are more prevalent where Spo11 activity is strongest. b, Percentage of total Spo11-oligos that are Spo11-DCs, plotted for every hotspot (n = 3,910). Although the proportion of Spo11-DCs within each hotspot varied widely from less than 0.1% to more than 10% of the Spo11-oligo signal, the majority (86%) of hotspots displayed a Spo11-DC proportion of at least 1%, and about one fifth (18%) of hotspots displayed a Spo11-DC proportion of over 5%. In tel1∆, the fraction of hotspots falling into these categories increased to 94% and 47%, respectively, consistent with the median fraction of Spo11-DCs per hotspot being about 1.7-fold greater. P value by two-tailed Kruskal–Wallis H-test. c, d, Quantitative correlation between filtered Spo11-DC frequency (c), or percentage of Spo11-DCs within each hotspot (d), and total Spo11-oligo frequency for all DSB hotspots, in wild-type (left) and tel1∆ (right) yeast. Although Spo11-DC frequency correlated positively with total Spo11-oligo counts, the relationship was nonlinear, such that Spo11-DCs were observed disproportionately more frequently within the strongest hotspots. e, Comparison between tel1∆ and wild type of the percentage of total Spo11-oligo signal within each hotspot that is classified as a Spo11-DC. These ratios are stratified on the x axis by the Spo11-DC frequency in wild-type cells, and ratios are coloured to indicate those hotspots in which the proportion of Spo11-DCs is at least twofold increased (red) or twofold decreased (blue) in tel1∆ relative to wild type. Although Spo11-DCs were globally more frequent in tel1∆ compared to wild type, this relationship was not uniform across all hotspots.
Extended Data Fig. 4 Fine-scale patterns of S. cerevisiae Spo11-DCs within DSB hotspots.
a, Spo11-DC arcs link the 5′ ends of overlapping Franklin- and Rosalind-strand filtered reads. For all bioinformatic analyses, only overlapping read pairs of lengths greater than 30 nt are considered because this is the minimum length of Spo11-DCs detected physically (Fig. 1c). We believe that enrichment of some shorter overlapping pairs arises from the artefactual overlap of canonical oligos (less than 30-nt length) within dense hotspot regions. b–e, Arc diagram depiction of Spo11-DCs mapped across example hotspots encompassing strong (b), narrow (c), and wide (d) classes, presented as in Fig. 2e, f. Top, unfiltered strand-specific Spo11-oligos (Franklin strand, red; Rosalind strand, blue). Arcs (grey-scale-frequency-weighted) link the 5′ ends of each Spo11-DC. Bottom, smoothed unfiltered strand-specific Spo11-oligos, overlaid with frequency histograms of Spo11-DC midpoints (grey). The left flanks of Spo11-DC peaks are enriched for Franklin-strand hits, whereas the right flanks are enriched for Rosalind-strand hits. This relationship was visualized most easily at narrow, low-frequency hotspots in which the patterns of Spo11-DCs were less complex. In e, wild-type and tel1∆ data are compared for the same hotspots. Although TEL1 deletion does have subtle effects on the pattern and abundance of both Spo11-oligos and Spo11-DCs, it did not alter the asymmetric pattern of Spo11-oligo strand disparity that is associated with regions of preferential Spo11-DC formation. In all plotted Spo11-oligo and Spo11-DC maps, Rosalind-strand signals are shifted by 1 bp to the left so that differences between the abundance of F- and R-mapping reads at individual cleavage sites can be more directly compared.
Extended Data Fig. 5 Global analysis of strand disparities at Spo11-DC termini in S. cerevisiae wild-type and tel1∆ cells.
a, Explanatory cartoon for the calculation of Spo11-DC strand ratio and strand-ratio differential (left/right). See Methods for further details. b–g, Strand ratios of Spo11-oligos at Spo11-DC sites in wild-type (b, c) and tel1∆ (d–g) cells. Average strand-specific Spo11-oligo signal (b, f), and strand ratio (c, g), centred on the strongest Spo11-DC midpoint within every DSB hotspot (n = 3,910). Strand ratio (Franklin/Rosalind total Spo11-oligo HPM) was computed at the left and right 5′ end of every unique Spo11-DC molecule (a), stratified by length. Strand-ratio differential (c, g) indicates the fold difference in the ratios when comparing the left and right 5′ ends of each Spo11-DC molecule. The relationships described above were unchanged after TEL1 deletion, suggesting that the Spo11-oligo patterns are an intrinsic feature of sites at which Spo11-DCs are generated, and are not subject to regulation by Tel1. h, Explanatory cartoon for strand-ratio calculation for all Spo11-oligos. i, j, Strand ratio (Franklin/Rosalind total Spo11-oligo HPM) was computed at the 5′ end of all observed (unfiltered) Franklin- or Rosalind-strand Spo11-oligos (h). Unlike at Spo11-DC sites (b, f), bulk Spo11-oligos, across all sites, display no net strand disparity in either wild-type (i) or tel1∆ (j) strains. In all plotted Spo11-oligo and Spo11-DC maps, Rosalind-strand signals are shifted by 1 bp to the left so that differences between the abundance of Franklin- and Rosalind-mapping reads at individual cleavage sites can be more directly compared. When considering all Spo11-oligo sites (h–j), some degree of strand disparity is a feature of most Spo11-oligo sites (it is a continuum of skew in both directions—some sites are skewed towards Franklin, some towards Rosalind, and some have little or no skew), but, when considered in aggregate, bulk Spo11-oligo sites have no net skew towards Franklin or Rosalind regardless of which strand the Spo11-oligo is considered. By contrast, sites at which we detect Spo11-DC formation (a–g; which is only a subset of all the sites onto which Spo11-oligos are mapped) display an asymmetric average skew within the total Spo11-oligo Franklin and Rosalind reads, with the left end of Spo11-DCs being sites at which the total Spo11-oligo pool is skewed towards Franklin and the right end sites at which the total Spo11-oligo pool is skewed towards Rosalind. Notably, this analysis uses the total Spo11-oligo pool, not just Spo11-DC molecules. Thus, the pattern of skews is a global feature of the entire Spo11-oligo pool at sites that form Spo11-DCs, and is not a feature that is observed at all Spo11-oligo sites. We interpret these observations to mean that sites at which Spo11-DCs form are different, being disproportionately sites of biased strand disparity. Owing to their low abundance at any particular site, removing Spo11-DCs from the total pool of Franklin and Rosalind reads has no effect on the strand disparity observed. Finally, consistent with these interpretations, we have found that the degree of disparity at any given site is not predictive of Spo11-DC abundance at that site, further indicating that Spo11-DC formation is not the cause of the disparity. Instead, locations of strand disparity and Spo11-DC are correlated in position, probably because they are influenced by similar properties of DSB hotspots (that is, a proposed Spo11 platform) (Fig. 4). Thus, overall, we conclude that asymmetric strand disparity is not unique to specific hotspots, nor to specific mutants, nor caused by Spo11-DCs, but is, instead, an intrinsic feature of the meiotic recombination process that informs the mechanism of Spo11-DSB formation.
Extended Data Fig. 6 Fine-scale analysis of SPO11-DCs within mouse DSB hotspots.
Mouse SPO11-oligo libraries4 were remapped using paired-end Bowtie2 alignment. a, b, SPO11-oligo length distribution of the entire library (a), or after applying the 2-bp overlap filter (b) as in upper cartoon in Fig. 2a. Filtering was less efficient than in yeast (Methods), retaining numerous molecules of less than 30-nt length that, based on our analysis in yeast, are likely to be a filtering artefact. Therefore, as for S. cerevisiae, SPO11-DCs are defined as filtered molecules of greater than 30 nt in length. Filtering retains weak peaks in Atm−/− that display an approximately 10-bp periodicity. c, Percentage of total SPO11-oligos that are SPO11-DCs on the basis of overlap filtering, plotted for every hotspot in which filtered Spo11-DCs were detected (n = 1,831 in wild-type mice; n = 8,010 in Atm−/− mice). P value by two-tailed Kruskal–Wallis H-test. d, Total Atm−/− SPO11-oligos were filtered into two size classes then aggregated around approximately 21,000 hotspot centres revealing a strand-specific disparity for SPO11-oligos greater than 30 nt. e–h, Representative arc diagrams of SPO11-DCs (grey-scale-frequency-weighted arcs) in wild-type and Atm−/− mice at four different hotspots in chromosomes 11 (e), 10 (f), 8 (g) and 5 (h) relative to total strand-specific SPO11-oligos (top, raw; bottom, smoothed; red, Franklin strand; blue, Rosalind strand). The percentage of total SPO11-oligos that are SPO11-DCs is indicated. Unlike in Atm−/− mice, SPO11-DCs in wild-type mice often did not coincide with strong SPO11-oligo signals, suggesting that some may arise from additional artefacts of the filtering.
Extended Data Fig. 7 Categorization of S. cerevisiae meiotic recombination events containing short 6:2 segments.
a, Summary of the frequency of each subclassification event type for the indicated strains. Only events containing 6:2 segments of 30 to 150 bp in length were considered. Category-A events are compatible with gap repair because flanking heteroduplex DNA patterns are in trans orientation, whereas category B events are incompatible because the flanking heteroduplex DNA patterns are in cis orientation. Category-C events lack the flanking heteroduplex DNA patterns necessary to assign the event. Events were separated into those with a single or multiple such 6:2 segments. Fractions of total events for each subtype were not calculated for multi events because they frequently contain more than one sub-type. b–g, Example event sub-classifications. Genotype calls were made at each marker (vertical line). Adjacent segments of the same genotype are joined with horizontal bars (red or blue) to aid visualization of patterns. Each horizontal bar is a sequenced haplotype from one meiotic octad. 6:2 segments are indicated in pale blue. Event limits are indicated by beige (crossover) or pink (noncrossover) bars. Orientation of 5′ and 3′ strands are indicated in instances in which it was possible to obtain phasing information from noncrossover trans events within the event, or from events elsewhere in the octad (see Methods for further details). In c, a second segment of 6:2 segregation was not considered because the minimum length is more than 1.2 kb. h, Relationship between Spo11-DC size and the probability of it overlapping at least one SNP. To estimate probable detection rates of theoretical Spo11-DCs of varying size, sliding windows of increasing size were moved across the reference genome, and the number of genetic markers within each window was recorded for each position. As examples, on average, Spo11-DCs of 30 bp and 150 bp in size will be detected only 15% and 50% of the time, respectively. Owing to the non-uniform distribution of both Spo11 DSBs and genetic markers—in particular the slightly greater density of polymorphisms within intergenic regions in which Spo11 DSBs most often arise—the probability of detection may be slightly greater than that estimated from the genome-wide polymorphism density. i, Quantification of recombination event types in tel1∆msh2∆ based on categories presented in Fig. 3b, c. Relative proportions of category A–C were unchanged when compared to wild type, but a greater number of events containing 6:2 segments of greater than 150 bp were observed. j, k, log2-transformed ratio of observed Spo11-oligo (j) or Spo11-DC (k) density within each 6:2 segment divided by the Spo11-oligo or Spo11-DC density within the entire event, in msh2∆, for each category. Although these analyses broadly agree with each other, the absence of Spo11-DCs in many of the category-B and -C segments prevented their analysis, artefactually inflating both their global ratio and decreasing the difference when compared to category A (which overlaps with both Spo11-oligos and Spo11-DCs). l–n, TEL1 deletion has no effect on the association between recombination patterns containing 6:2 segments and Spo11-DSB activity. l, Fraction of 6:2 segments overlapping hotspots in tel1∆msh2∆, as for Fig. 3f. m, log2 ratio of observed Spo11-oligo density within each 6:2 segment divided by the Spo11-oligo density within the entire event, in tel1∆msh2∆, for each category, as for j. Unlike deletion of MLH1, MLH3 and EXO1 (Fig. 3e), deletion of TEL1 had no effect on the relative frequencies of 6:2 segments within categories A–C (n). In (j, k, m), P values by two-tailed Kruskal–Wallis H-test. Individual data points are shown overlaid with box indicating median plus first and third quartiles, and whiskers indicating 1.5× the interquartile range. In l, n, whiskers indicate 95% confidence intervals. In l, P value by two-tailed Z-test of proportions. In i, l, m, 2,159 events were analysed across 10 biologically independent meiotic samples. In j, k, 1,643 events were analysed across 9 biologically independent meiotic samples.
Extended Data Fig. 8 Cartoon to explain mapped strand disparity and Tel1-insensitive Spo11-DC formation.
a, Model to account for observed strand bias. Mre11-dependent 3′→5′ exonuclease activity is shown relative to the covalent attachment of Spo11 to 5′ DNA ends. We hypothesize that Spo11-oligos and Spo11-DCs that formed within the axis-associated Spo11 platform (grey area) are protected from Mre11 nuclease activity and are therefore efficiently mapped (long Spo11-oligos), whereas efficient resection in the flanking regions leads to shorter Spo11-oligos that are not mapped. We assume that because the position and frequency of such hotspot–axis interactions will vary from one hotspot to another—and from one cell to another—this will contribute to the substantial variations in position and abundance of both Spo11-oligos and Spo11-DCs within hotspot regions. Although it is formally possible that the disparity can alternatively arise from less (rather than more) efficient resection flanking the platform area—leading to unprocessed DSB ends, and thereby a paucity of Spo11-oligo reads in these locations—we consider this unlikely because it would require there to be a high frequency of persistent unresected DSBs, something which is not observed in wild-type cells. b, In S. cerevisiae, Tel1 has no greater effect on Spo11-DC formation than on global DSB formation (Fig. 1d), yet efficiently inhibits DSB formation between adjacent hotspots10. Therefore, we propose that DSBs arise concertedly within a DSB-active hotspot region (top cartoon)—creating Spo11-DCs—before Tel1 can act to inhibit their formation. Such hotspot activation is likely to arise through the proposed tethering of nucleosome-depleted hotspot DNA to the pro-DSB axis components28,44,45,46 (bottom). These interactions will enable the formation of both single Spo11 DSBs and/or Spo11-DCs at the tethered locus, either of which will cause Tel1 activation. Once activated, Tel1 inhibits DSB formation at hotspots within the rest of the tethered loop10, and at any axis-associated DSB hotspots within adjacent loop regions. Such inhibition may arise through direct inhibition and/or destabilization of Spo11 and/or other pro-DSB axis components such as Rec114–Mer2–Mei4 (RMM), and/or inhibition of hotspot–axis interactions10,11,15.
Supplementary information
Rights and permissions
About this article
Cite this article
Johnson, D., Crawford, M., Cooper, T. et al. Concerted cutting by Spo11 illuminates meiotic DNA break mechanics. Nature 594, 572–576 (2021). https://doi.org/10.1038/s41586-021-03389-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03389-3
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.